Jianpeng Zhang
Domain and Content Adaptive Convolution for Domain Generalization in Medical Image Segmentation
Sep 13, 2021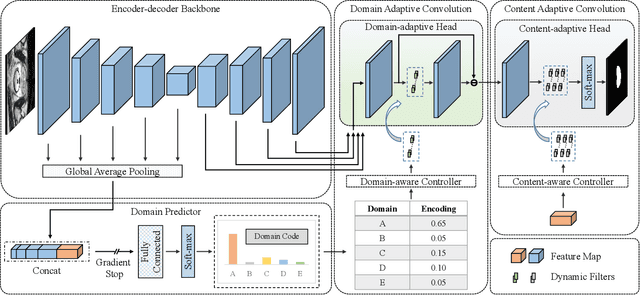

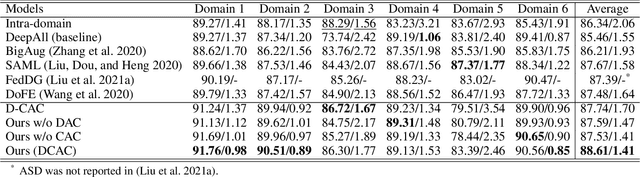
Abstract:The domain gap caused mainly by variable medical image quality renders a major obstacle on the path between training a segmentation model in the lab and applying the trained model to unseen clinical data. To address this issue, domain generalization methods have been proposed, which however usually use static convolutions and are less flexible. In this paper, we propose a multi-source domain generalization model, namely domain and content adaptive convolution (DCAC), for medical image segmentation. Specifically, we design the domain adaptive convolution (DAC) module and content adaptive convolution (CAC) module and incorporate both into an encoder-decoder backbone. In the DAC module, a dynamic convolutional head is conditioned on the predicted domain code of the input to make our model adapt to the unseen target domain. In the CAC module, a dynamic convolutional head is conditioned on the global image features to make our model adapt to the test image. We evaluated the DCAC model against the baseline and four state-of-the-art domain generalization methods on the prostate segmentation, COVID-19 lesion segmentation, and optic cup/optic disc segmentation tasks. Our results indicate that the proposed DCAC model outperforms all competing methods on each segmentation task, and also demonstrate the effectiveness of the DAC and CAC modules.
CoTr: Efficiently Bridging CNN and Transformer for 3D Medical Image Segmentation
Mar 04, 2021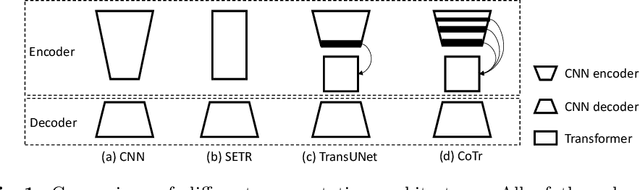
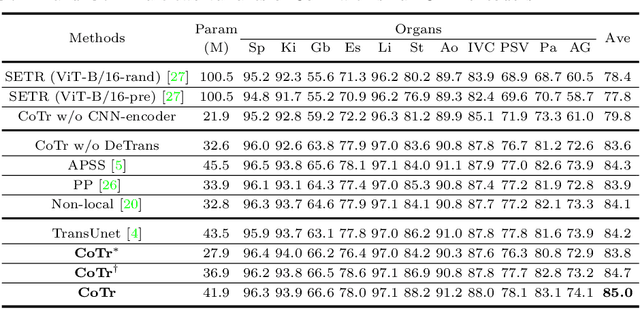
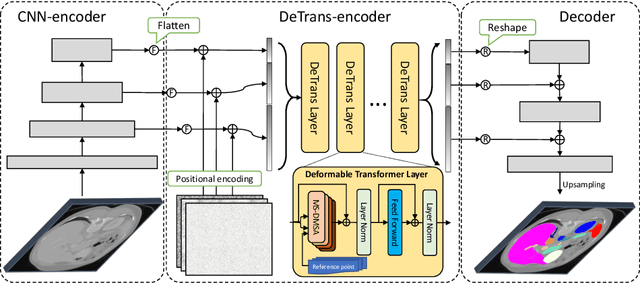
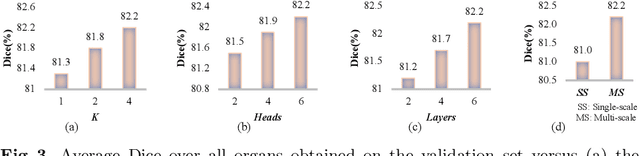
Abstract:Convolutional neural networks (CNNs) have been the de facto standard for nowadays 3D medical image segmentation. The convolutional operations used in these networks, however, inevitably have limitations in modeling the long-range dependency due to their inductive bias of locality and weight sharing. Although Transformer was born to address this issue, it suffers from extreme computational and spatial complexities in processing high-resolution 3D feature maps. In this paper, we propose a novel framework that efficiently bridges a {\bf Co}nvolutional neural network and a {\bf Tr}ansformer {\bf (CoTr)} for accurate 3D medical image segmentation. Under this framework, the CNN is constructed to extract feature representations and an efficient deformable Transformer (DeTrans) is built to model the long-range dependency on the extracted feature maps. Different from the vanilla Transformer which treats all image positions equally, our DeTrans pays attention only to a small set of key positions by introducing the deformable self-attention mechanism. Thus, the computational and spatial complexities of DeTrans have been greatly reduced, making it possible to process the multi-scale and high-resolution feature maps, which are usually of paramount importance for image segmentation. We conduct an extensive evaluation on the Multi-Atlas Labeling Beyond the Cranial Vault (BCV) dataset that covers 11 major human organs. The results indicate that our CoTr leads to a substantial performance improvement over other CNN-based, transformer-based, and hybrid methods on the 3D multi-organ segmentation task. Code is available at \def\UrlFont{\rm\small\ttfamily} \url{https://github.com/YtongXie/CoTr}
Inter-slice Context Residual Learning for 3D Medical Image Segmentation
Nov 28, 2020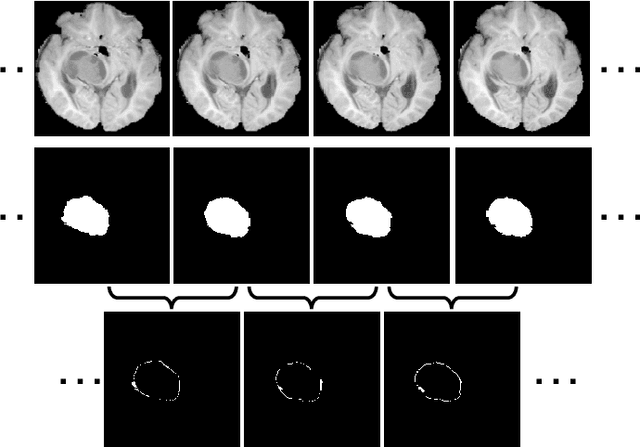
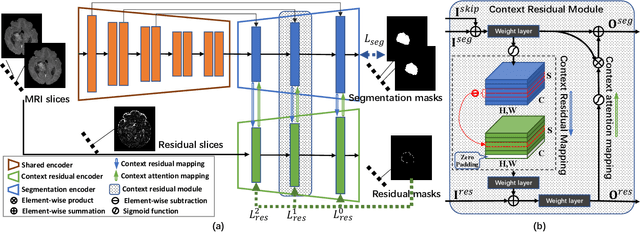

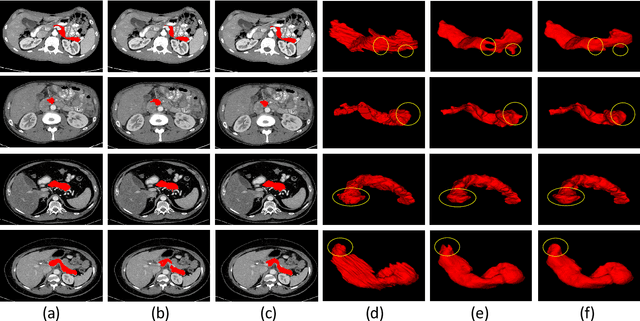
Abstract:Automated and accurate 3D medical image segmentation plays an essential role in assisting medical professionals to evaluate disease progresses and make fast therapeutic schedules. Although deep convolutional neural networks (DCNNs) have widely applied to this task, the accuracy of these models still need to be further improved mainly due to their limited ability to 3D context perception. In this paper, we propose the 3D context residual network (ConResNet) for the accurate segmentation of 3D medical images. This model consists of an encoder, a segmentation decoder, and a context residual decoder. We design the context residual module and use it to bridge both decoders at each scale. Each context residual module contains both context residual mapping and context attention mapping, the formal aims to explicitly learn the inter-slice context information and the latter uses such context as a kind of attention to boost the segmentation accuracy. We evaluated this model on the MICCAI 2018 Brain Tumor Segmentation (BraTS) dataset and NIH Pancreas Segmentation (Pancreas-CT) dataset. Our results not only demonstrate the effectiveness of the proposed 3D context residual learning scheme but also indicate that the proposed ConResNet is more accurate than six top-ranking methods in brain tumor segmentation and seven top-ranking methods in pancreas segmentation. Code is available at https://git.io/ConResNet
* Accpeted by IEEE-TMI
PGL: Prior-Guided Local Self-supervised Learning for 3D Medical Image Segmentation
Nov 25, 2020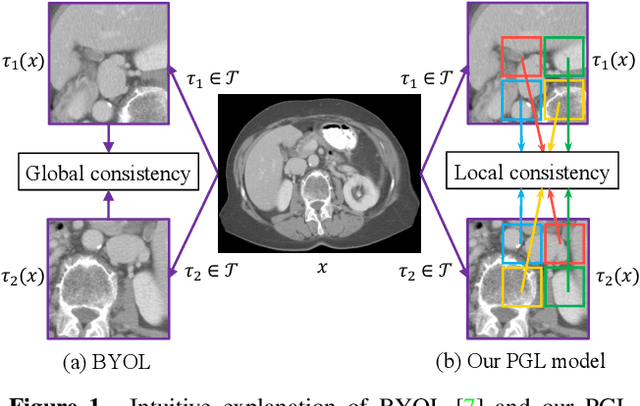
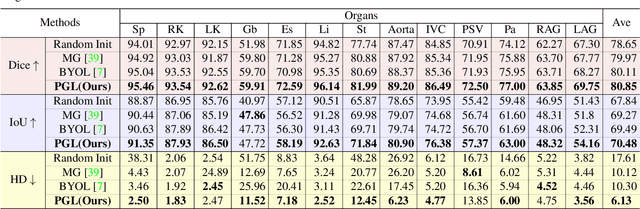
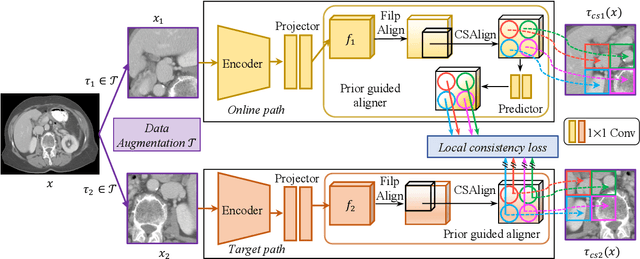
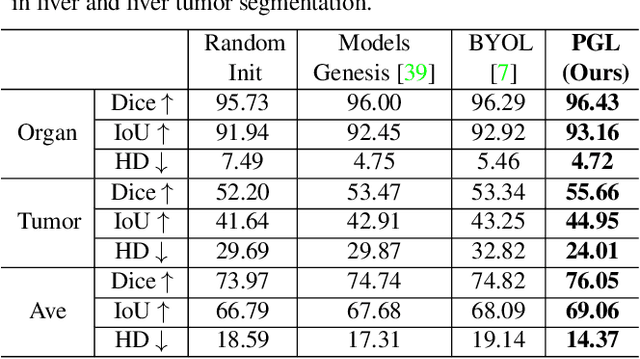
Abstract:It has been widely recognized that the success of deep learning in image segmentation relies overwhelmingly on a myriad amount of densely annotated training data, which, however, are difficult to obtain due to the tremendous labor and expertise required, particularly for annotating 3D medical images. Although self-supervised learning (SSL) has shown great potential to address this issue, most SSL approaches focus only on image-level global consistency, but ignore the local consistency which plays a pivotal role in capturing structural information for dense prediction tasks such as segmentation. In this paper, we propose a PriorGuided Local (PGL) self-supervised model that learns the region-wise local consistency in the latent feature space. Specifically, we use the spatial transformations, which produce different augmented views of the same image, as a prior to deduce the location relation between two views, which is then used to align the feature maps of the same local region but being extracted on two views. Next, we construct a local consistency loss to minimize the voxel-wise discrepancy between the aligned feature maps. Thus, our PGL model learns the distinctive representations of local regions, and hence is able to retain structural information. This ability is conducive to downstream segmentation tasks. We conducted an extensive evaluation on four public computerized tomography (CT) datasets that cover 11 kinds of major human organs and two tumors. The results indicate that using pre-trained PGL model to initialize a downstream network leads to a substantial performance improvement over both random initialization and the initialization with global consistency-based models. Code and pre-trained weights will be made available at: https://git.io/PGL.
DoDNet: Learning to segment multi-organ and tumors from multiple partially labeled datasets
Nov 20, 2020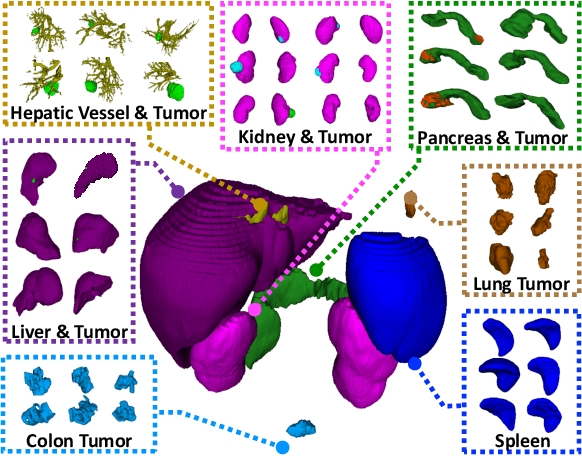
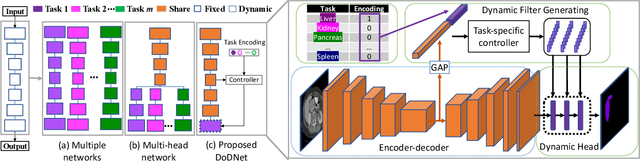
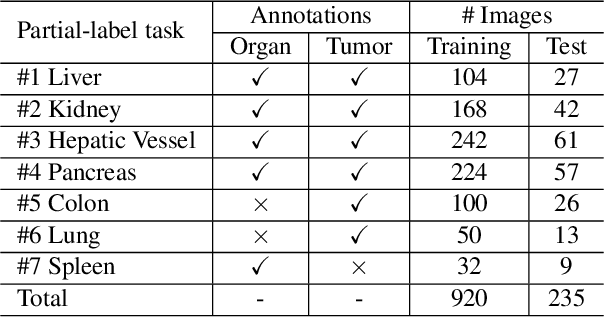
Abstract:Due to the intensive cost of labor and expertise in annotating 3D medical images at a voxel level, most benchmark datasets are equipped with the annotations of only one type of organs and/or tumors, resulting in the so-called partially labeling issue. To address this, we propose a dynamic on-demand network (DoDNet) that learns to segment multiple organs and tumors on partially labeled datasets. DoDNet consists of a shared encoder-decoder architecture, a task encoding module, a controller for generating dynamic convolution filters, and a single but dynamic segmentation head. The information of the current segmentation task is encoded as a task-aware prior to tell the model what the task is expected to solve. Different from existing approaches which fix kernels after training, the kernels in dynamic head are generated adaptively by the controller, conditioned on both input image and assigned task. Thus, DoDNet is able to segment multiple organs and tumors, as done by multiple networks or a multi-head network, in a much efficient and flexible manner. We have created a large-scale partially labeled dataset, termed MOTS, and demonstrated the superior performance of our DoDNet over other competitors on seven organ and tumor segmentation tasks. We also transferred the weights pre-trained on MOTS to a downstream multi-organ segmentation task and achieved state-of-the-art performance. This study provides a general 3D medical image segmentation model that has been pre-trained on a large-scale partially labelled dataset and can be extended (after fine-tuning) to downstream volumetric medical data segmentation tasks. The dataset and code areavailableat: https://git.io/DoDNet
Pairwise Relation Learning for Semi-supervised Gland Segmentation
Aug 06, 2020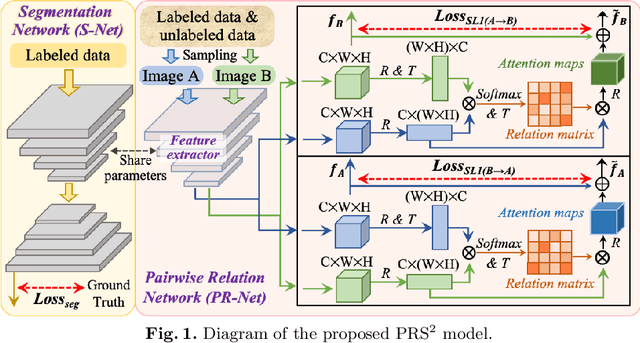
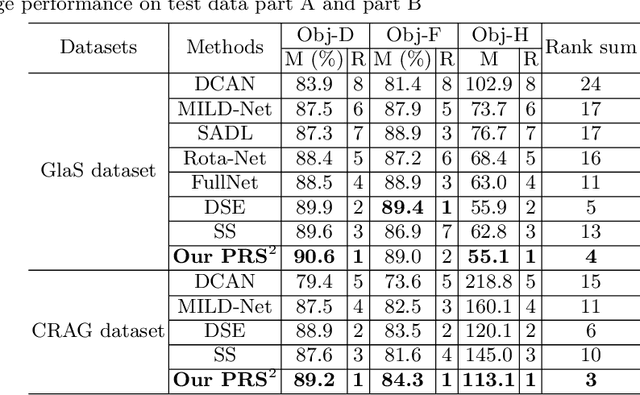
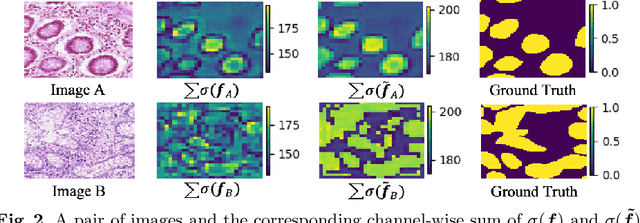
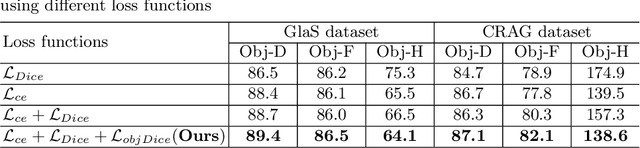
Abstract:Accurate and automated gland segmentation on histology tissue images is an essential but challenging task in the computer-aided diagnosis of adenocarcinoma. Despite their prevalence, deep learning models always require a myriad number of densely annotated training images, which are difficult to obtain due to extensive labor and associated expert costs related to histology image annotations. In this paper, we propose the pairwise relation-based semi-supervised (PRS^2) model for gland segmentation on histology images. This model consists of a segmentation network (S-Net) and a pairwise relation network (PR-Net). The S-Net is trained on labeled data for segmentation, and PR-Net is trained on both labeled and unlabeled data in an unsupervised way to enhance its image representation ability via exploiting the semantic consistency between each pair of images in the feature space. Since both networks share their encoders, the image representation ability learned by PR-Net can be transferred to S-Net to improve its segmentation performance. We also design the object-level Dice loss to address the issues caused by touching glands and combine it with other two loss functions for S-Net. We evaluated our model against five recent methods on the GlaS dataset and three recent methods on the CRAG dataset. Our results not only demonstrate the effectiveness of the proposed PR-Net and object-level Dice loss, but also indicate that our PRS^2 model achieves the state-of-the-art gland segmentation performance on both benchmarks.
COVID-19 Screening on Chest X-ray Images Using Deep Learning based Anomaly Detection
Mar 27, 2020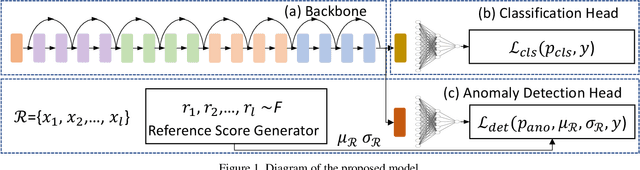
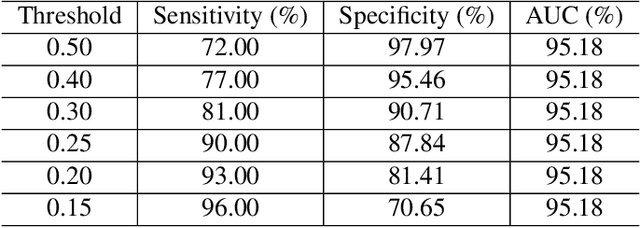
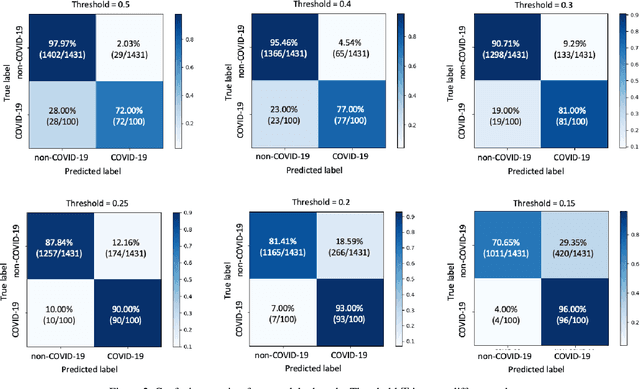
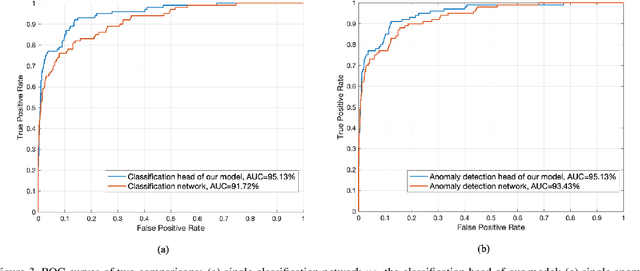
Abstract:Coronaviruses are important human and animal pathogens. To date the novel COVID-19 coronavirus is rapidly spreading worldwide and subsequently threatening health of billions of humans. Clinical studies have shown that most COVID-19 patients suffer from the lung infection. Although chest CT has been shown to be an effective imaging technique for lung-related disease diagnosis, chest Xray is more widely available due to its faster imaging time and considerably lower cost than CT. Deep learning, one of the most successful AI techniques, is an effective means to assist radiologists to analyze the vast amount of chest X-ray images, which can be critical for efficient and reliable COVID-19 screening. In this work, we aim to develop a new deep anomaly detection model for fast, reliable screening. To evaluate the model performance, we have collected 100 chest X-ray images of 70 patients confirmed with COVID-19 from the Github repository. To facilitate deep learning, more data are needed. Thus, we have also collected 1431 additional chest X-ray images confirmed as other pneumonia of 1008 patients from the public ChestX-ray14 dataset. Our initial experimental results show that the model developed here can reliably detect 96.00% COVID-19 cases (sensitivity being 96.00%) and 70.65% non-COVID-19 cases (specificity being 70.65%) when evaluated on 1531 Xray images with two splits of the dataset.
Natural-Logarithm-Rectified Activation Function in Convolutional Neural Networks
Aug 25, 2019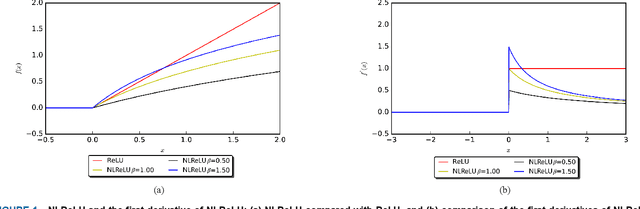
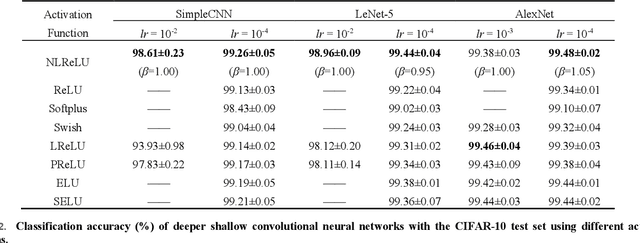
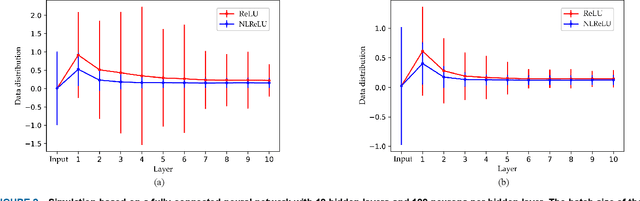
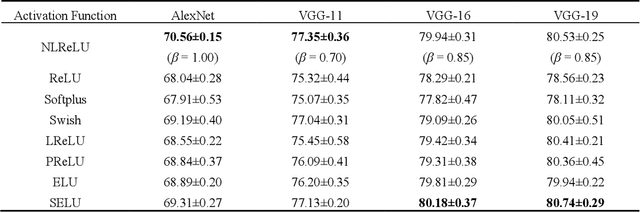
Abstract:Activation functions play a key role in providing remarkable performance in deep neural networks, and the rectified linear unit (ReLU) is one of the most widely used activation functions. Various new activation functions and improvements on ReLU have been proposed, but each carry performance drawbacks. In this paper, we propose an improved activation function, which we name the natural-logarithm-rectified linear unit (NLReLU). This activation function uses the parametric natural logarithmic transform to improve ReLU and is simply defined as. NLReLU not only retains the sparse activation characteristic of ReLU, but it also alleviates the "dying ReLU" and vanishing gradient problems to some extent. It also reduces the bias shift effect and heteroscedasticity of neuron data distributions among network layers in order to accelerate the learning process. The proposed method was verified across ten convolutional neural networks with different depths for two essential datasets. Experiments illustrate that convolutional neural networks with NLReLU exhibit higher accuracy than those with ReLU, and that NLReLU is comparable to other well-known activation functions. NLReLU provides 0.16% and 2.04% higher classification accuracy on average compared to ReLU when used in shallow convolutional neural networks with the MNIST and CIFAR-10 datasets, respectively. The average accuracy of deep convolutional neural networks with NLReLU is 1.35% higher on average with the CIFAR-10 dataset.
A Sensitivity Analysis of Attention-Gated Convolutional Neural Networks for Sentence Classification
Aug 25, 2019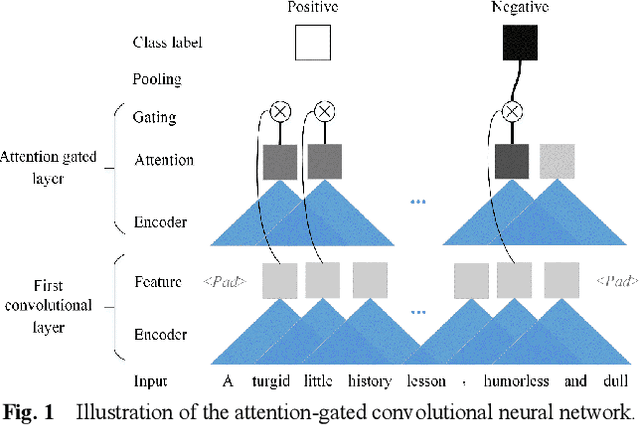
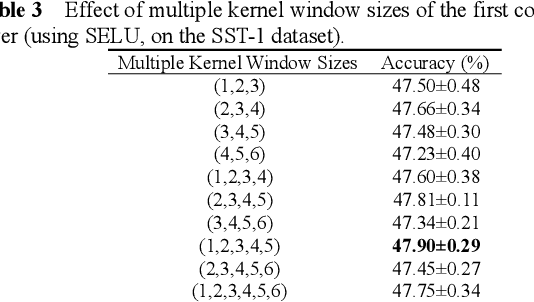
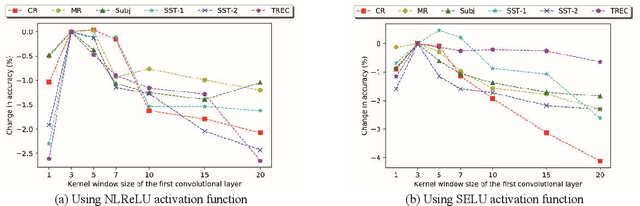

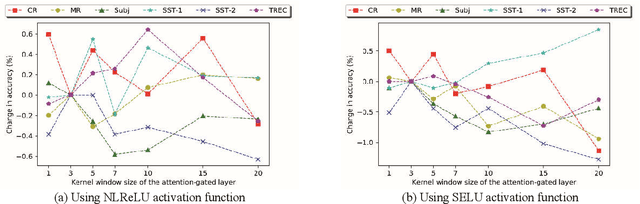
Abstract:Recently, Attention-Gated Convolutional Neural Networks (AGCNNs) perform well on several essential sentence classification tasks and show robust performance in practical applications. However, AGCNNs are required to set many hyperparameters, and it is not known how sensitive the model's performance changes with them. In this paper, we conduct a sensitivity analysis on the effect of different hyperparameters s of AGCNNs, e.g., the kernel window size and the number of feature maps. Also, we investigate the effect of different combinations of hyperparameters settings on the model's performance to analyze to what extent different parameters settings contribute to AGCNNs' performance. Meanwhile, we draw practical advice from a wide range of empirical results. Through the sensitivity analysis experiment, we improve the hyperparameters settings of AGCNNs. Experiments show that our proposals achieve an average of 0.81% and 0.67% improvements on AGCNN-NLReLU-rand and AGCNN-SELU-rand, respectively; and an average of 0.47% and 0.45% improvements on AGCNN-NLReLU-static and AGCNN-SELU-static, respectively.
Semi- and Weakly Supervised Directional Bootstrapping Model for Automated Skin Lesion Segmentation
Mar 12, 2019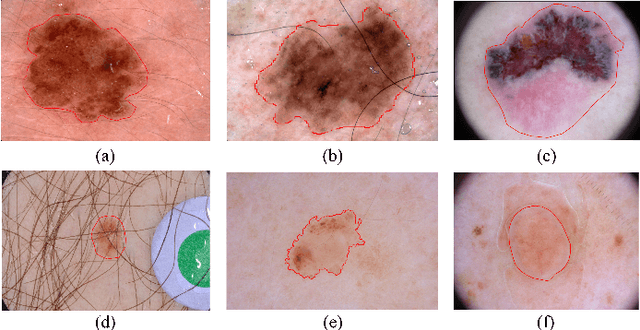
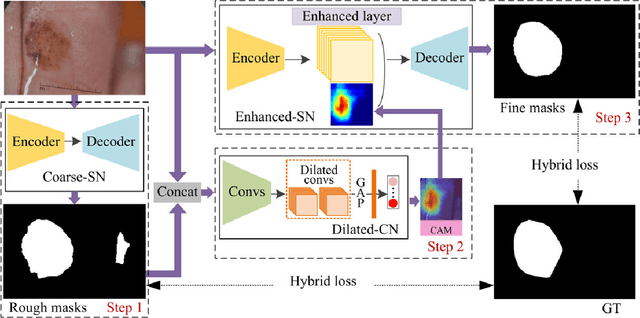
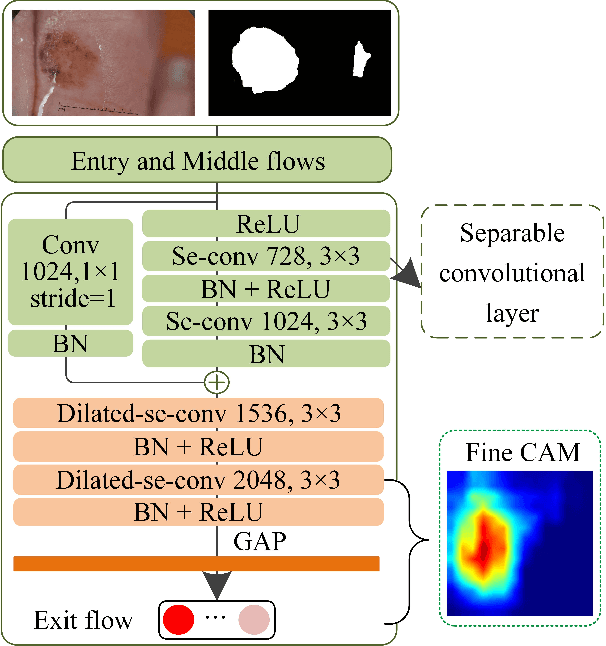

Abstract:Automated skin lesion segmentation on dermoscopy images is an essential and challenging task in the computer-aided diagnosis of skin cancer. Despite their prevalence and relatively good performance, deep learning based segmentation methods require a myriad number of training images with pixel-level dense annotation, which is hard to obtain due to the efforts and costs related to dermoscopy images acquisition and annotation. In this paper, we propose the semi- and weakly supervised directional bootstrapping (SWSDB) model for skin lesion segmentation, which consists of three deep convolutional neural networks: a coarse segmentation network (coarse-SN), a dilated classification network (dilated-CN) and an enhanced segmentation network (enhanced-SN). Both the coarse-SN and enhanced-SN are trained using the images with pixel-level annotation, and the dilated-CN is trained using the images with image-level class labels. The coarse-SN generates rough segmentation masks that provide a prior bootstrapping for the dilated-CN and help it produce accurate lesion localization maps. The maps are then fed into the enhanced-SN to transfer the localization information learned from image-level labels to the enhanced-SN to generate segmentation results. Furthermore, we introduce a hybrid loss that is the weighted sum of a dice loss and a rank loss to the coarse-SN and enhanced-SN, ensuring both networks' good compatibility for the data with imbalanced classes and imbalanced hard-easy pixels. We evaluated the proposed SWSDB model on the ISIC-2017 challenge dataset and PH2 dataset and achieved a Jaccard index of 80.4% and 89.4%, respectively, setting a new record in skin lesion segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge