Jeroen Bertels
A Robust Ensemble Algorithm for Ischemic Stroke Lesion Segmentation: Generalizability and Clinical Utility Beyond the ISLES Challenge
Apr 03, 2024



Abstract:Diffusion-weighted MRI (DWI) is essential for stroke diagnosis, treatment decisions, and prognosis. However, image and disease variability hinder the development of generalizable AI algorithms with clinical value. We address this gap by presenting a novel ensemble algorithm derived from the 2022 Ischemic Stroke Lesion Segmentation (ISLES) challenge. ISLES'22 provided 400 patient scans with ischemic stroke from various medical centers, facilitating the development of a wide range of cutting-edge segmentation algorithms by the research community. Through collaboration with leading teams, we combined top-performing algorithms into an ensemble model that overcomes the limitations of individual solutions. Our ensemble model achieved superior ischemic lesion detection and segmentation accuracy on our internal test set compared to individual algorithms. This accuracy generalized well across diverse image and disease variables. Furthermore, the model excelled in extracting clinical biomarkers. Notably, in a Turing-like test, neuroradiologists consistently preferred the algorithm's segmentations over manual expert efforts, highlighting increased comprehensiveness and precision. Validation using a real-world external dataset (N=1686) confirmed the model's generalizability. The algorithm's outputs also demonstrated strong correlations with clinical scores (admission NIHSS and 90-day mRS) on par with or exceeding expert-derived results, underlining its clinical relevance. This study offers two key findings. First, we present an ensemble algorithm (https://github.com/Tabrisrei/ISLES22_Ensemble) that detects and segments ischemic stroke lesions on DWI across diverse scenarios on par with expert (neuro)radiologists. Second, we show the potential for biomedical challenge outputs to extend beyond the challenge's initial objectives, demonstrating their real-world clinical applicability.
Dice Semimetric Losses: Optimizing the Dice Score with Soft Labels
Apr 01, 2023



Abstract:The soft Dice loss (SDL) has taken a pivotal role in many automated segmentation pipelines in the medical imaging community. Over the last years, some reasons behind its superior functioning have been uncovered and further optimizations have been explored. However, there is currently no implementation that supports its direct use in settings with soft labels. Hence, a synergy between the use of SDL and research leveraging the use of soft labels, also in the context of model calibration, is still missing. In this work, we introduce Dice semimetric losses (DMLs), which (i) are by design identical to SDL in a standard setting with hard labels, but (ii) can be used in settings with soft labels. Our experiments on the public QUBIQ, LiTS and KiTS benchmarks confirm the potential synergy of DMLs with soft labels (e.g. averaging, label smoothing, and knowledge distillation) over hard labels (e.g. majority voting and random selection). As a result, we obtain superior Dice scores and model calibration, which supports the wider adoption of DMLs in practice. Code is available at \href{https://github.com/zifuwanggg/JDTLosses}{https://github.com/zifuwanggg/JDTLosses}.
Convolutional neural networks for medical image segmentation
Nov 17, 2022



Abstract:In this article, we look into some essential aspects of convolutional neural networks (CNNs) with the focus on medical image segmentation. First, we discuss the CNN architecture, thereby highlighting the spatial origin of the data, voxel-wise classification and the receptive field. Second, we discuss the sampling of input-output pairs, thereby highlighting the interaction between voxel-wise classification, patch size and the receptive field. Finally, we give a historical overview of crucial changes to CNN architectures for classification and segmentation, giving insights in the relation between three pivotal CNN architectures: FCN, U-Net and DeepMedic.
DeepVoxNet2: Yet another CNN framework
Nov 17, 2022Abstract:We know that both the CNN mapping function and the sampling scheme are of paramount importance for CNN-based image analysis. It is clear that both functions operate in the same space, with an image axis $\mathcal{I}$ and a feature axis $\mathcal{F}$. Remarkably, we found that no frameworks existed that unified the two and kept track of the spatial origin of the data automatically. Based on our own practical experience, we found the latter to often result in complex coding and pipelines that are difficult to exchange. This article introduces our framework for 1, 2 or 3D image classification or segmentation: DeepVoxNet2 (DVN2). This article serves as an interactive tutorial, and a pre-compiled version, including the outputs of the code blocks, can be found online in the public DVN2 repository. This tutorial uses data from the multimodal Brain Tumor Image Segmentation Benchmark (BRATS) of 2018 to show an example of a 3D segmentation pipeline.
Final infarct prediction in acute ischemic stroke
Nov 09, 2022Abstract:This article focuses on the control center of each human body: the brain. We will point out the pivotal role of the cerebral vasculature and how its complex mechanisms may vary between subjects. We then emphasize a specific acute pathological state, i.e., acute ischemic stroke, and show how medical imaging and its analysis can be used to define the treatment. We show how the core-penumbra concept is used in practice using mismatch criteria and how machine learning can be used to make predictions of the final infarct, either via deconvolution or convolutional neural networks.
Theoretical analysis and experimental validation of volume bias of soft Dice optimized segmentation maps in the context of inherent uncertainty
Nov 08, 2022



Abstract:The clinical interest is often to measure the volume of a structure, which is typically derived from a segmentation. In order to evaluate and compare segmentation methods, the similarity between a segmentation and a predefined ground truth is measured using popular discrete metrics, such as the Dice score. Recent segmentation methods use a differentiable surrogate metric, such as soft Dice, as part of the loss function during the learning phase. In this work, we first briefly describe how to derive volume estimates from a segmentation that is, potentially, inherently uncertain or ambiguous. This is followed by a theoretical analysis and an experimental validation linking the inherent uncertainty to common loss functions for training CNNs, namely cross-entropy and soft Dice. We find that, even though soft Dice optimization leads to an improved performance with respect to the Dice score and other measures, it may introduce a volume bias for tasks with high inherent uncertainty. These findings indicate some of the method's clinical limitations and suggest doing a closer ad-hoc volume analysis with an optional re-calibration step.
* 18 pages, 7 figures, 3 tables, published in Elsevier Medical Image Analysis (2021)
Evaluation of Medical Image Segmentation Models for Uncertain, Small or Empty Reference Annotations
Sep 30, 2022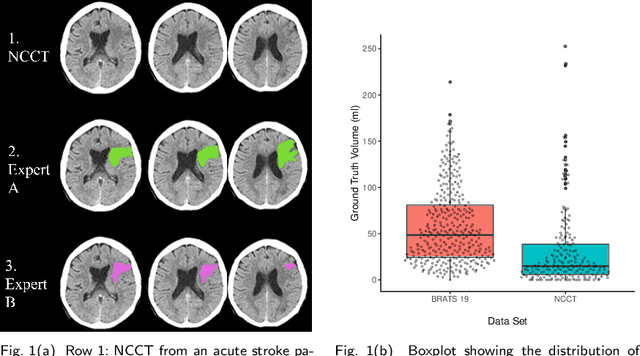
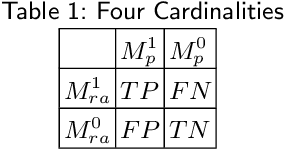


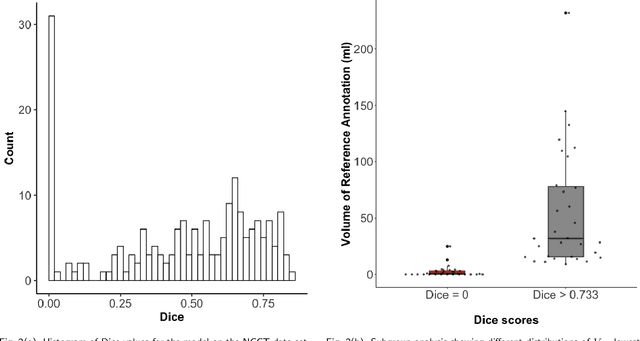
Abstract:Performance metrics for medical image segmentation models are used to measure agreement between the reference annotation and the prediction. A common set of metrics is used in the development of such models to make results more comparable. However, there is a mismatch between the distributions in public data sets and cases encountered in clinical practice. Many common metrics fail to measure the impact of this mismatch, especially for clinical data sets containing uncertain, small or empty reference annotation. Thus, models may not be validated for clinically meaningful agreement by such metrics. Dimensions of evaluating clinical value include independence from reference annotation volume size, consideration of uncertainty of reference annotations, reward of volumetric and/or location agreement and reward of correct classification of empty reference annotations. Unlike common public data sets, our in-house data set is more representative. It contains uncertain, small or empty reference annotations. We examine publicly available metrics on the predictions of a deep learning framework in order to identify for which settings common metrics provide clinical meaningful results. We compare to a public benchmark data set without uncertain, small or empty reference annotations. https://github.com/SophieOstmeier/UncertainSmallEmpty
The Dice loss in the context of missing or empty labels: Introducing $Φ$ and $ε$
Jul 19, 2022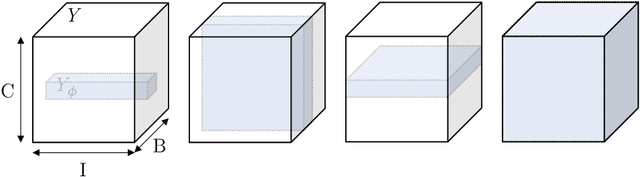
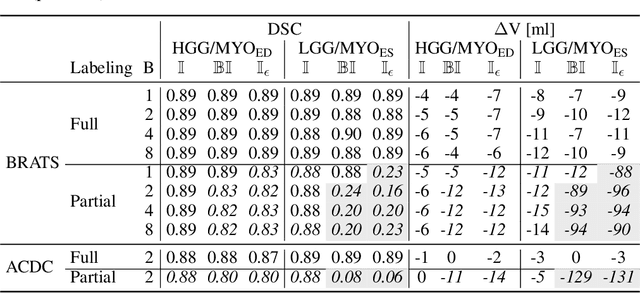
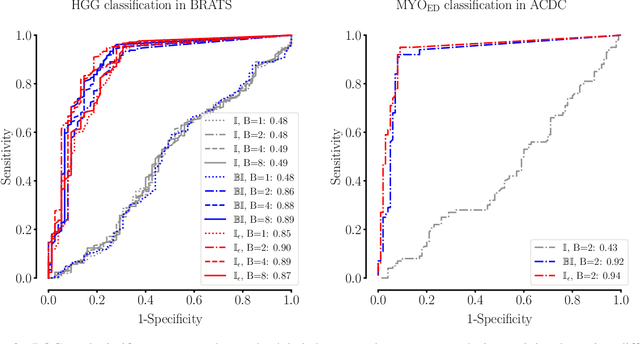
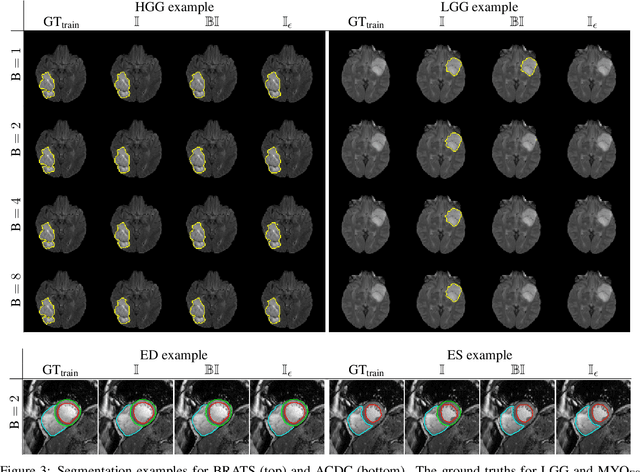
Abstract:Albeit the Dice loss is one of the dominant loss functions in medical image segmentation, most research omits a closer look at its derivative, i.e. the real motor of the optimization when using gradient descent. In this paper, we highlight the peculiar action of the Dice loss in the presence of missing or empty labels. First, we formulate a theoretical basis that gives a general description of the Dice loss and its derivative. It turns out that the choice of the reduction dimensions $\Phi$ and the smoothing term $\epsilon$ is non-trivial and greatly influences its behavior. We find and propose heuristic combinations of $\Phi$ and $\epsilon$ that work in a segmentation setting with either missing or empty labels. Second, we empirically validate these findings in a binary and multiclass segmentation setting using two publicly available datasets. We confirm that the choice of $\Phi$ and $\epsilon$ is indeed pivotal. With $\Phi$ chosen such that the reductions happen over a single batch (and class) element and with a negligible $\epsilon$, the Dice loss deals with missing labels naturally and performs similarly compared to recent adaptations specific for missing labels. With $\Phi$ chosen such that the reductions happen over multiple batch elements or with a heuristic value for $\epsilon$, the Dice loss handles empty labels correctly. We believe that this work highlights some essential perspectives and hope that it encourages researchers to better describe their exact implementation of the Dice loss in future work.
On the relationship between calibrated predictors and unbiased volume estimation
Dec 23, 2021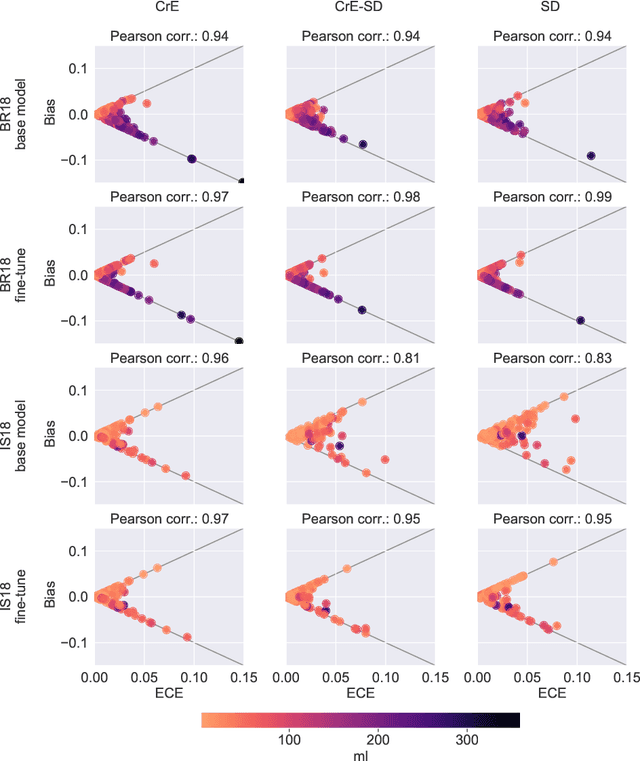
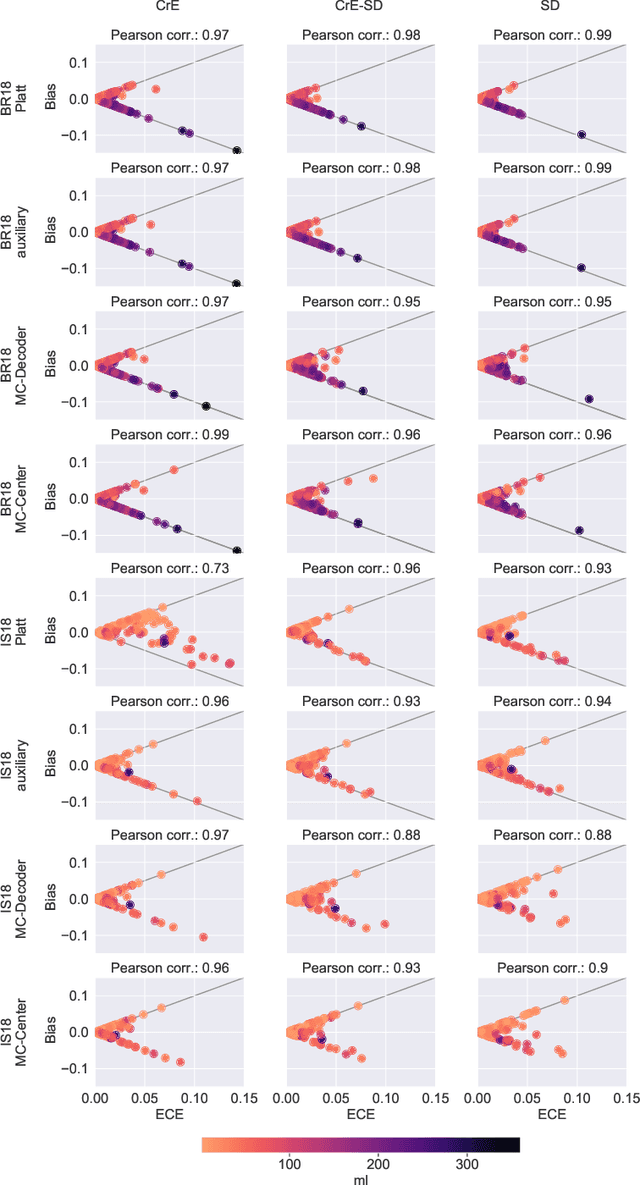
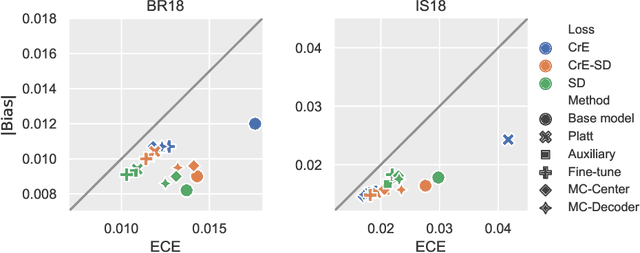
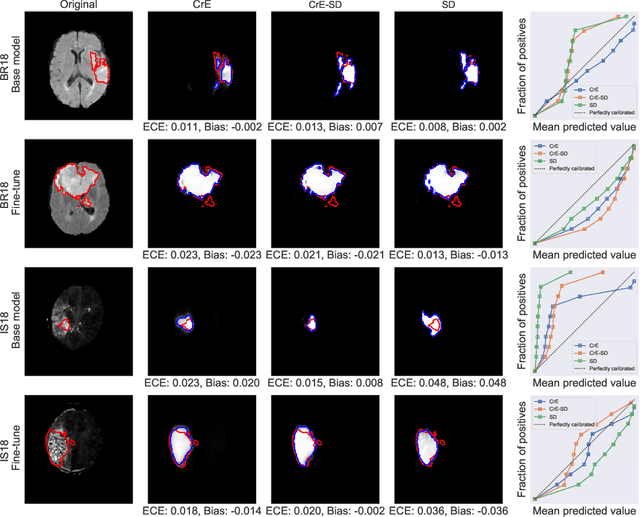
Abstract:Machine learning driven medical image segmentation has become standard in medical image analysis. However, deep learning models are prone to overconfident predictions. This has led to a renewed focus on calibrated predictions in the medical imaging and broader machine learning communities. Calibrated predictions are estimates of the probability of a label that correspond to the true expected value of the label conditioned on the confidence. Such calibrated predictions have utility in a range of medical imaging applications, including surgical planning under uncertainty and active learning systems. At the same time it is often an accurate volume measurement that is of real importance for many medical applications. This work investigates the relationship between model calibration and volume estimation. We demonstrate both mathematically and empirically that if the predictor is calibrated per image, we can obtain the correct volume by taking an expectation of the probability scores per pixel/voxel of the image. Furthermore, we show that convex combinations of calibrated classifiers preserve volume estimation, but do not preserve calibration. Therefore, we conclude that having a calibrated predictor is a sufficient, but not necessary condition for obtaining an unbiased estimate of the volume. We validate our theoretical findings empirically on a collection of 18 different (calibrated) training strategies on the tasks of glioma volume estimation on BraTS 2018, and ischemic stroke lesion volume estimation on ISLES 2018 datasets.
Explainable-by-design Semi-Supervised Representation Learning for COVID-19 Diagnosis from CT Imaging
Dec 02, 2020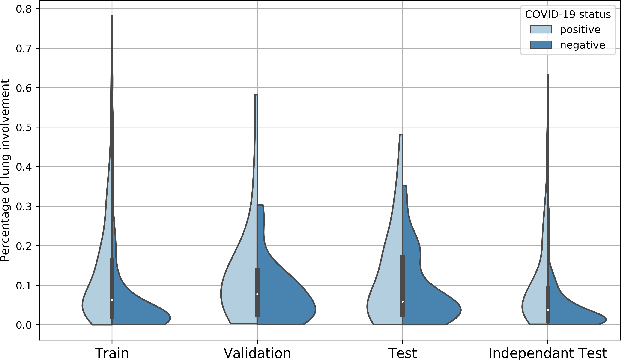
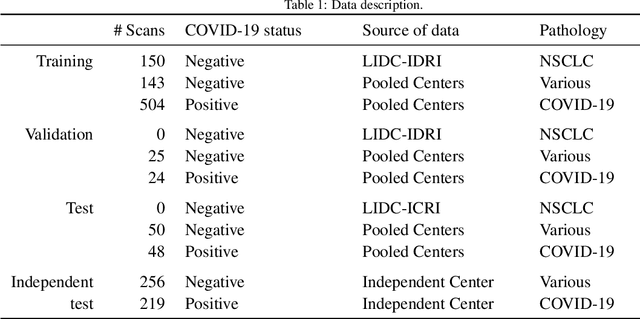
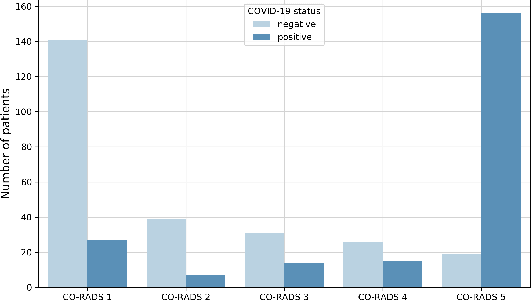
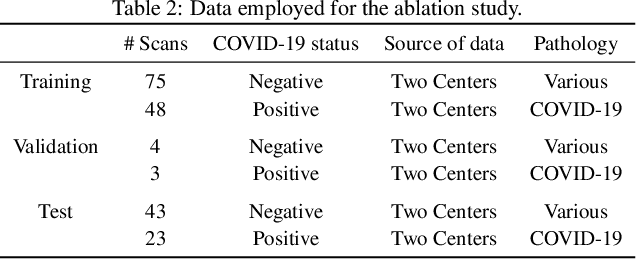
Abstract:Our motivating application is a real-world problem: COVID-19 classification from CT imaging, for which we present an explainable Deep Learning approach based on a semi-supervised classification pipeline that employs variational autoencoders to extract efficient feature embedding. We have optimized the architecture of two different networks for CT images: (i) a novel conditional variational autoencoder (CVAE) with a specific architecture that integrates the class labels inside the encoder layers and uses side information with shared attention layers for the encoder, which make the most of the contextual clues for representation learning, and (ii) a downstream convolutional neural network for supervised classification using the encoder structure of the CVAE. With the explainable classification results, the proposed diagnosis system is very effective for COVID-19 classification. Based on the promising results obtained qualitatively and quantitatively, we envisage a wide deployment of our developed technique in large-scale clinical studies.Code is available at https://git.etrovub.be/AVSP/ct-based-covid-19-diagnostic-tool.git.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge