Paul Suetens
Theoretical analysis and experimental validation of volume bias of soft Dice optimized segmentation maps in the context of inherent uncertainty
Nov 08, 2022



Abstract:The clinical interest is often to measure the volume of a structure, which is typically derived from a segmentation. In order to evaluate and compare segmentation methods, the similarity between a segmentation and a predefined ground truth is measured using popular discrete metrics, such as the Dice score. Recent segmentation methods use a differentiable surrogate metric, such as soft Dice, as part of the loss function during the learning phase. In this work, we first briefly describe how to derive volume estimates from a segmentation that is, potentially, inherently uncertain or ambiguous. This is followed by a theoretical analysis and an experimental validation linking the inherent uncertainty to common loss functions for training CNNs, namely cross-entropy and soft Dice. We find that, even though soft Dice optimization leads to an improved performance with respect to the Dice score and other measures, it may introduce a volume bias for tasks with high inherent uncertainty. These findings indicate some of the method's clinical limitations and suggest doing a closer ad-hoc volume analysis with an optional re-calibration step.
* 18 pages, 7 figures, 3 tables, published in Elsevier Medical Image Analysis (2021)
Explainable-by-design Semi-Supervised Representation Learning for COVID-19 Diagnosis from CT Imaging
Dec 02, 2020
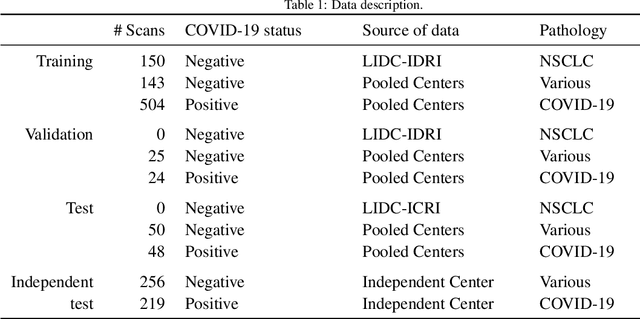
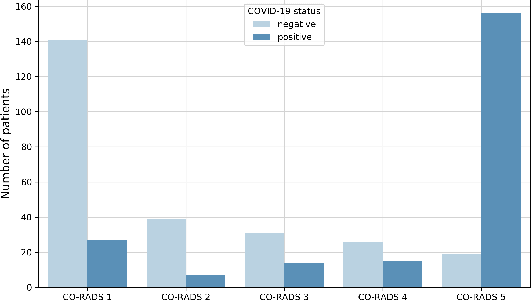
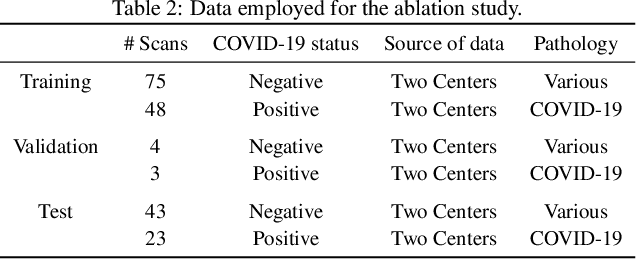
Abstract:Our motivating application is a real-world problem: COVID-19 classification from CT imaging, for which we present an explainable Deep Learning approach based on a semi-supervised classification pipeline that employs variational autoencoders to extract efficient feature embedding. We have optimized the architecture of two different networks for CT images: (i) a novel conditional variational autoencoder (CVAE) with a specific architecture that integrates the class labels inside the encoder layers and uses side information with shared attention layers for the encoder, which make the most of the contextual clues for representation learning, and (ii) a downstream convolutional neural network for supervised classification using the encoder structure of the CVAE. With the explainable classification results, the proposed diagnosis system is very effective for COVID-19 classification. Based on the promising results obtained qualitatively and quantitatively, we envisage a wide deployment of our developed technique in large-scale clinical studies.Code is available at https://git.etrovub.be/AVSP/ct-based-covid-19-diagnostic-tool.git.
Comparative study of deep learning methods for the automatic segmentation of lung, lesion and lesion type in CT scans of COVID-19 patients
Aug 21, 2020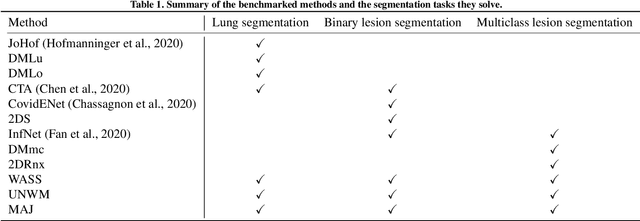
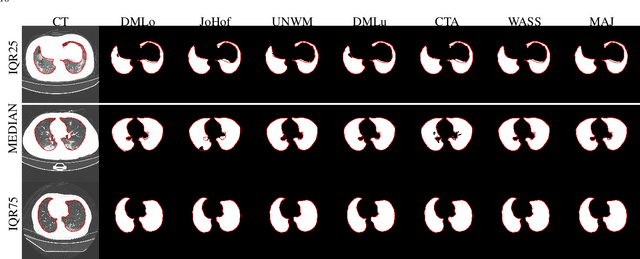
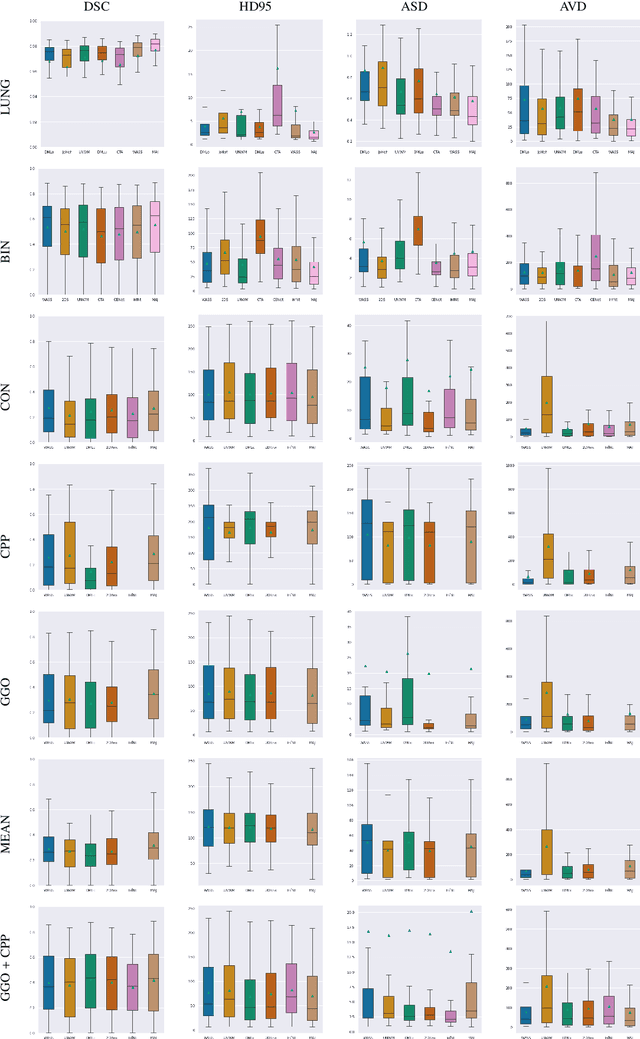
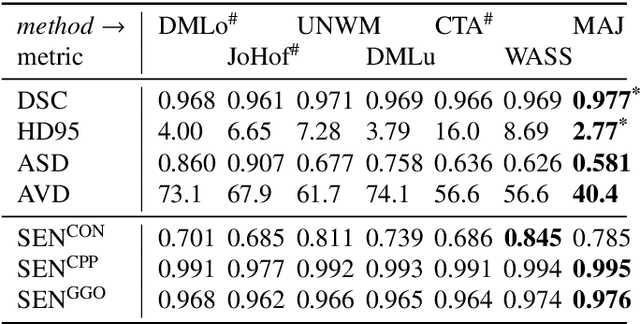
Abstract:Recent research on COVID-19 suggests that CT imaging provides useful information to assess disease progression and assist diagnosis, in addition to help understanding the disease. There is an increasing number of studies that propose to use deep learning to provide fast and accurate quantification of COVID-19 using chest CT scans. The main tasks of interest are the automatic segmentation of lung and lung lesions in chest CT scans of confirmed or suspected COVID-19 patients. In this study, we compare twelve deep learning algorithms using a multi-center dataset, including both open-source and in-house developed algorithms. Results show that ensembling different methods can boost the overall test set performance for lung segmentation, binary lesion segmentation and multiclass lesion segmentation, resulting in mean Dice scores of 0.982, 0.724 and 0.469, respectively. The resulting binary lesions were segmented with a mean absolute volume error of 91.3 ml. In general, the task of distinguishing different lesion types was more difficult, with a mean absolute volume difference of 152 ml and mean Dice scores of 0.369 and 0.523 for consolidation and ground glass opacity, respectively. All methods perform binary lesion segmentation with an average volume error that is better than visual assessment by human raters, suggesting these methods are mature enough for a large-scale evaluation for use in clinical practice.
Optimization with soft Dice can lead to a volumetric bias
Nov 06, 2019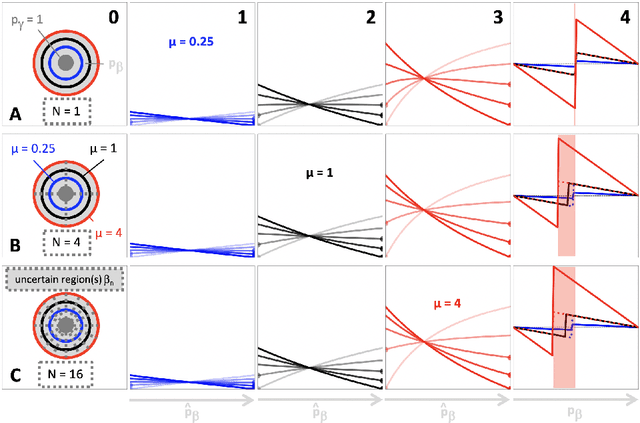

Abstract:Segmentation is a fundamental task in medical image analysis. The clinical interest is often to measure the volume of a structure. To evaluate and compare segmentation methods, the similarity between a segmentation and a predefined ground truth is measured using metrics such as the Dice score. Recent segmentation methods based on convolutional neural networks use a differentiable surrogate of the Dice score, such as soft Dice, explicitly as the loss function during the learning phase. Even though this approach leads to improved Dice scores, we find that, both theoretically and empirically on four medical tasks, it can introduce a volumetric bias for tasks with high inherent uncertainty. As such, this may limit the method's clinical applicability.
Prediction of final infarct volume from native CT perfusion and treatment parameters using deep learning
Dec 06, 2018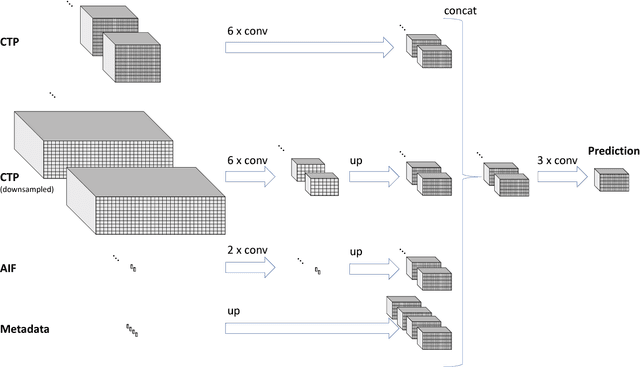
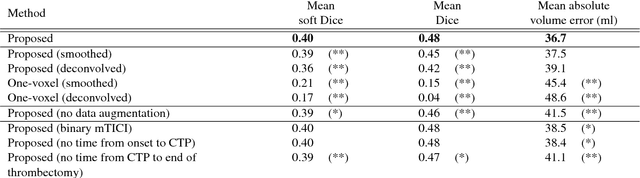
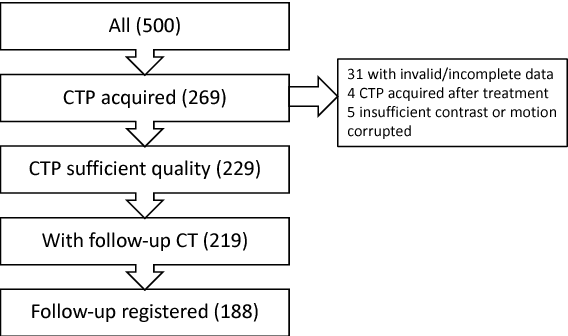
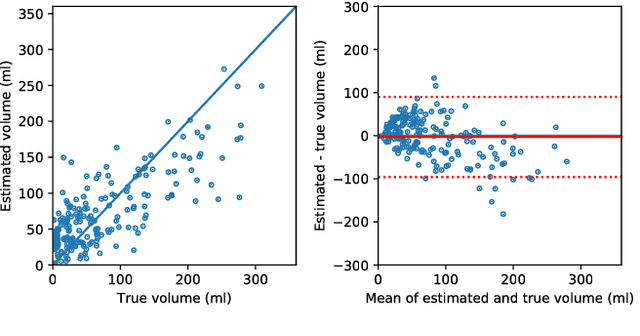
Abstract:CT Perfusion (CTP) imaging has gained importance in the diagnosis of acute stroke. Conventional perfusion analysis performs a deconvolution of the measurements and thresholds the perfusion parameters to determine the tissue status. We pursue a data-driven and deconvolution-free approach, where a deep neural network learns to predict the final infarct volume directly from the native CTP images and metadata such as the time parameters and treatment. This would allow clinicians to simulate various treatments and gain insight into predicted tissue status over time. We demonstrate on a multicenter dataset that our approach is able to predict the final infarct and effectively uses the metadata. An ablation study shows that using the native CTP measurements instead of the deconvolved measurements improves the prediction.
Perfusion parameter estimation using neural networks and data augmentation
Oct 11, 2018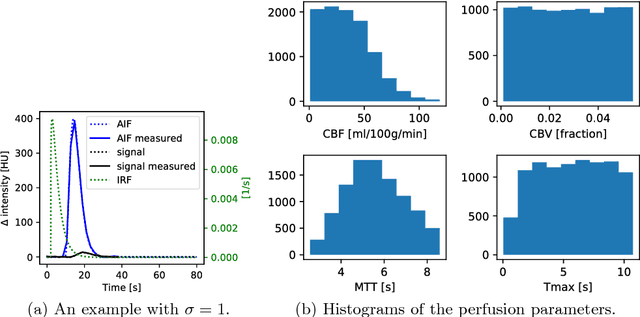
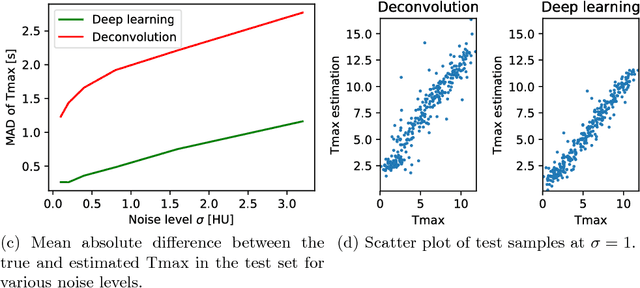
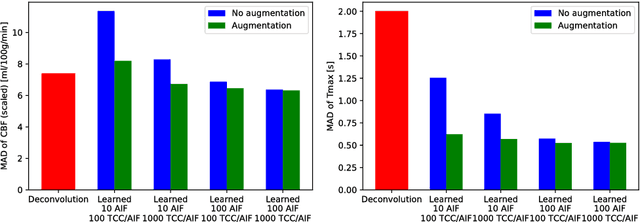
Abstract:Perfusion imaging plays a crucial role in acute stroke diagnosis and treatment decision making. Current perfusion analysis relies on deconvolution of the measured signals, an operation that is mathematically ill-conditioned and requires strong regularization. We propose a neural network and a data augmentation approach to predict perfusion parameters directly from the native measurements. A comparison on simulated CT Perfusion data shows that the neural network provides better estimations for both CBF and Tmax than a state of the art deconvolution method, and this over a wide range of noise levels. The proposed data augmentation enables to achieve these results with less than 100 datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge