Jeroen van der Laak
and on behalf of the UNICORN consortium
Automated risk classification of colon biopsies based on semantic segmentation of histopathology images
Sep 16, 2021
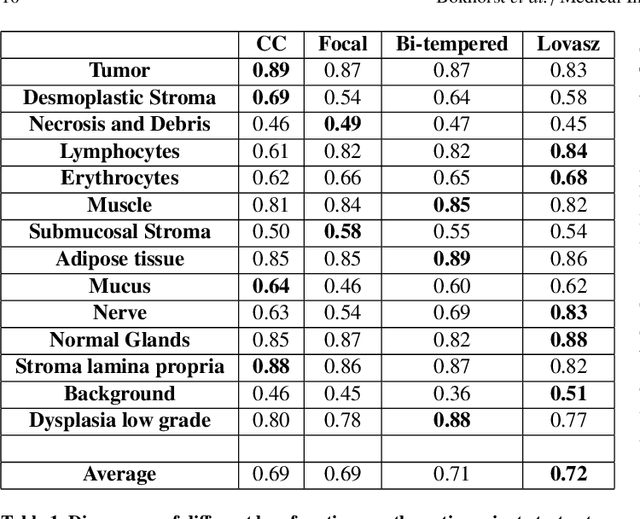
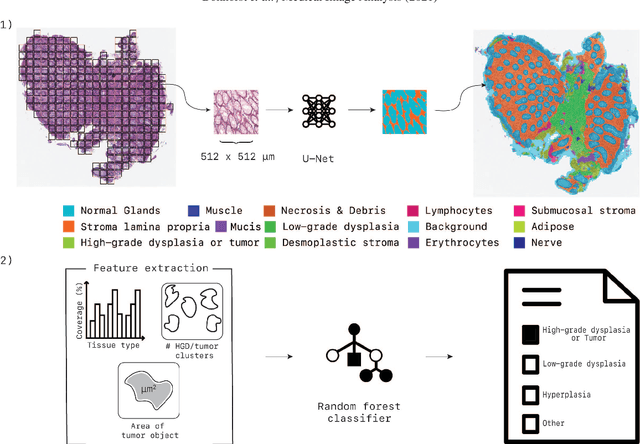
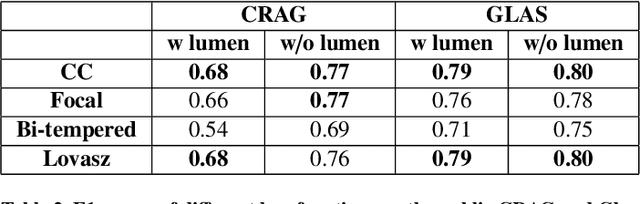
Abstract:Artificial Intelligence (AI) can potentially support histopathologists in the diagnosis of a broad spectrum of cancer types. In colorectal cancer (CRC), AI can alleviate the laborious task of characterization and reporting on resected biopsies, including polyps, the numbers of which are increasing as a result of CRC population screening programs, ongoing in many countries all around the globe. Here, we present an approach to address two major challenges in automated assessment of CRC histopathology whole-slide images. First, we present an AI-based method to segment multiple tissue compartments in the H\&E-stained whole-slide image, which provides a different, more perceptible picture of tissue morphology and composition. We test and compare a panel of state-of-the-art loss functions available for segmentation models, and provide indications about their use in histopathology image segmentation, based on the analysis of a) a multi-centric cohort of CRC cases from five medical centers in the Netherlands and Germany, and b) two publicly available datasets on segmentation in CRC. Second, we use the best performing AI model as the basis for a computer-aided diagnosis system (CAD) that classifies colon biopsies into four main categories that are relevant pathologically. We report the performance of this system on an independent cohort of more than 1,000 patients. The results show the potential of such an AI-based system to assist pathologists in diagnosis of CRC in the context of population screening. We have made the segmentation model available for research use on https://grand-challenge.org/algorithms/colon-tissue-segmentation/.
High-resolution Image Registration of Consecutive and Re-stained Sections in Histopathology
Jun 24, 2021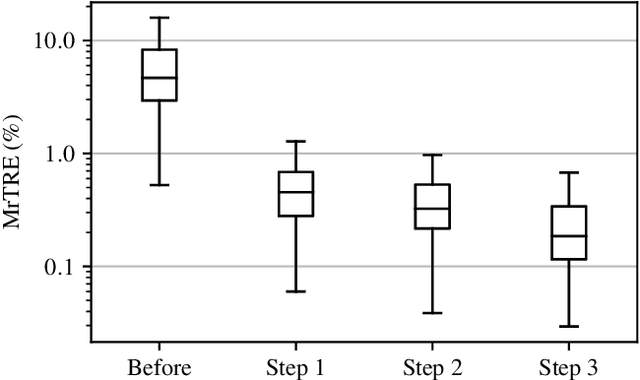
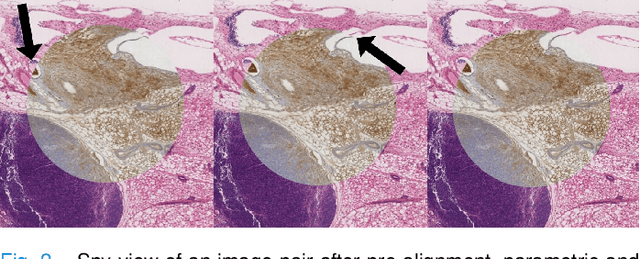
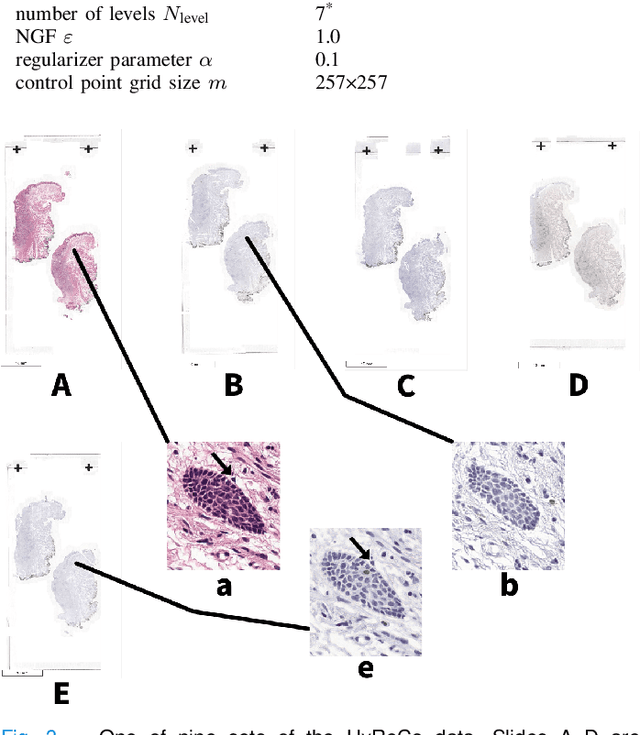
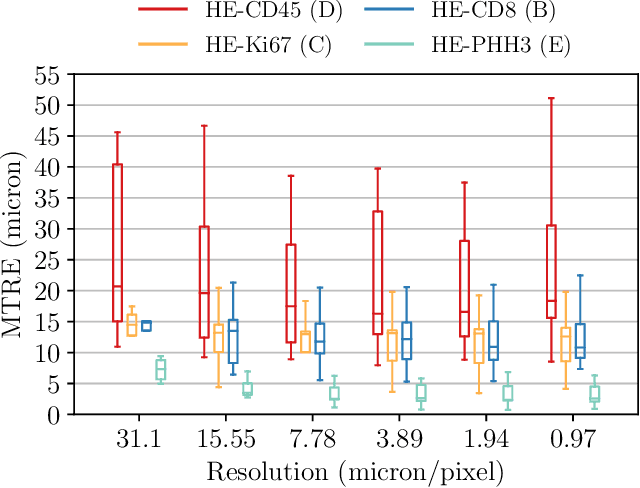
Abstract:We compare variational image registration in consectutive and re-stained sections from histopathology. We present a fully-automatic algorithm for non-parametric (nonlinear) image registration and apply it to a previously existing dataset from the ANHIR challenge (230 slide pairs, consecutive sections) and a new dataset (hybrid re-stained and consecutive, 81 slide pairs, ca. 3000 landmarks) which is made publicly available. Registration hyperparameters are obtained in the ANHIR dataset and applied to the new dataset without modification. In the new dataset, landmark errors after registration range from 13.2 micrometers for consecutive sections to 1 micrometer for re-stained sections. We observe that non-parametric registration leads to lower landmark errors in both cases, even though the effect is smaller in re-stained sections. The nucleus-level alignment after non-parametric registration of re-stained sections provides a valuable tool to generate automatic ground-truth for machine learning applications in histopathology.
Automated Scoring of Nuclear Pleomorphism Spectrum with Pathologist-level Performance in Breast Cancer
Dec 24, 2020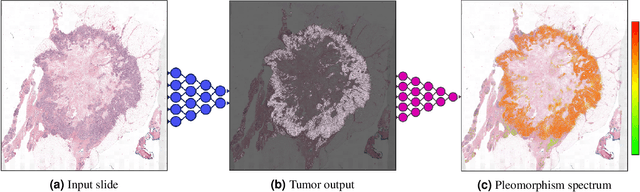
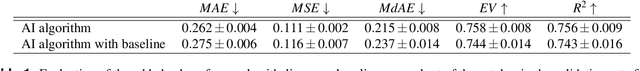
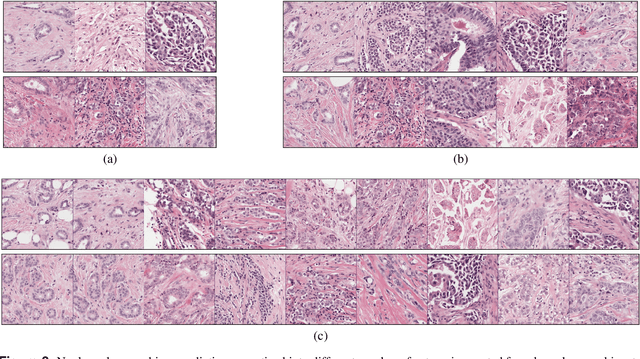
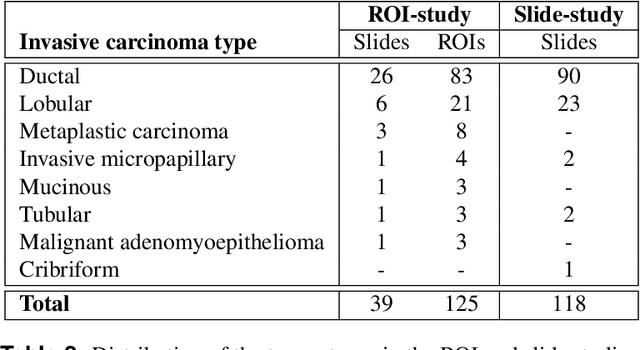
Abstract:Nuclear pleomorphism, defined herein as the extent of abnormalities in the overall appearance of tumor nuclei, is one of the components of the three-tiered breast cancer grading. Given that nuclear pleomorphism reflects a continuous spectrum of variation, we trained a deep neural network on a large variety of tumor regions from the collective knowledge of several pathologists, without constraining the network to the traditional three-category classification. We also motivate an additional approach in which we discuss the additional benefit of normal epithelium as baseline, following the routine clinical practice where pathologists are trained to score nuclear pleomorphism in tumor, having the normal breast epithelium for comparison. In multiple experiments, our fully-automated approach could achieve top pathologist-level performance in select regions of interest as well as at whole slide images, compared to ten and four pathologists, respectively.
HookNet: multi-resolution convolutional neural networks for semantic segmentation in histopathology whole-slide images
Jun 22, 2020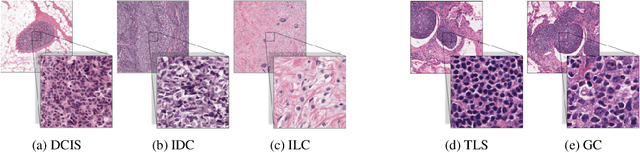
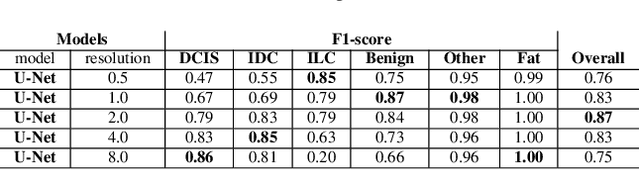
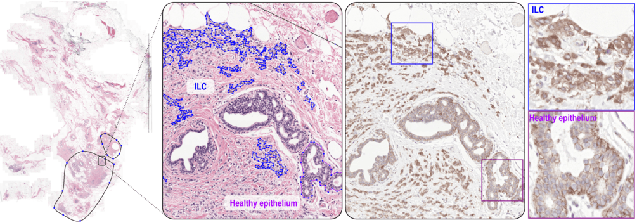
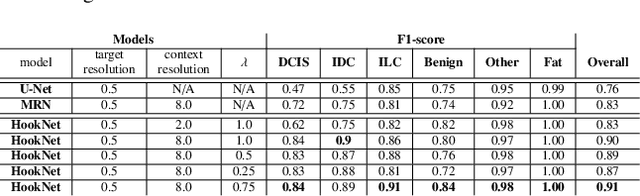
Abstract:We propose HookNet, a semantic segmentation model for histopathology whole-slide images, which combines context and details via multiple branches of encoder-decoder convolutional neural networks. Concentricpatches at multiple resolutions with different fields of view are used to feed different branches of HookNet, and intermediate representations are combined via a hooking mechanism. We describe a framework to design and train HookNet for achieving high-resolution semantic segmentation and introduce constraints to guarantee pixel-wise alignment in feature maps during hooking. We show the advantages of using HookNet in two histopathology image segmentation tasks where tissue type prediction accuracy strongly depends on contextual information, namely (1) multi-class tissue segmentation in breast cancer and, (2) segmentation of tertiary lymphoid structures and germinal centers in lung cancer. Weshow the superiority of HookNet when compared with single-resolution U-Net models working at different resolutions as well as with a recently published multi-resolution model for histopathology image segmentation
Detection of prostate cancer in whole-slide images through end-to-end training with image-level labels
Jun 05, 2020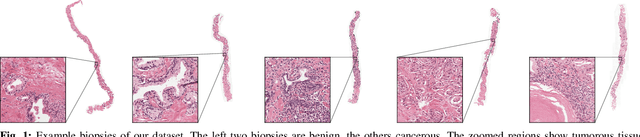
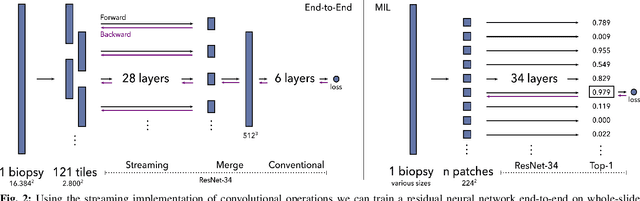
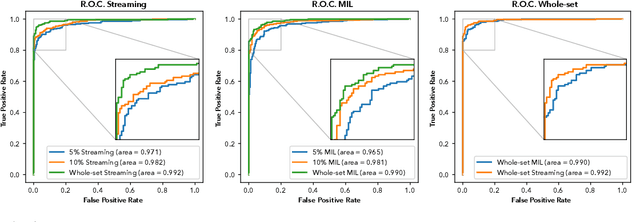

Abstract:Prostate cancer is the most prevalent cancer among men in Western countries, with 1.1 million new diagnoses every year. The gold standard for the diagnosis of prostate cancer is a pathologists' evaluation of prostate tissue. To potentially assist pathologists deep-learning-based cancer detection systems have been developed. Many of the state-of-the-art models are patch-based convolutional neural networks, as the use of entire scanned slides is hampered by memory limitations on accelerator cards. Patch-based systems typically require detailed, pixel-level annotations for effective training. However, such annotations are seldom readily available, in contrast to the clinical reports of pathologists, which contain slide-level labels. As such, developing algorithms which do not require manual pixel-wise annotations, but can learn using only the clinical report would be a significant advancement for the field. In this paper, we propose to use a streaming implementation of convolutional layers, to train a modern CNN (ResNet-34) with 21 million parameters end-to-end on 4712 prostate biopsies. The method enables the use of entire biopsy images at high-resolution directly by reducing the GPU memory requirements by 2.4 TB. We show that modern CNNs, trained using our streaming approach, can extract meaningful features from high-resolution images without additional heuristics, reaching similar performance as state-of-the-art patch-based and multiple-instance learning methods. By circumventing the need for manual annotations, this approach can function as a blueprint for other tasks in histopathological diagnosis. The source code to reproduce the streaming models is available at https://github.com/DIAGNijmegen/pathology-streaming-pipeline .
Extending Unsupervised Neural Image Compression With Supervised Multitask Learning
Apr 15, 2020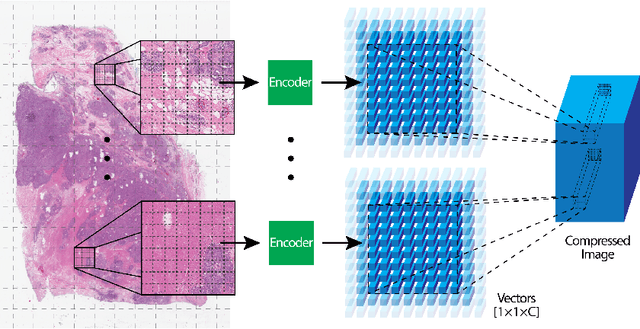
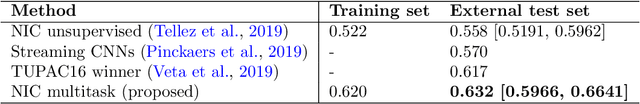
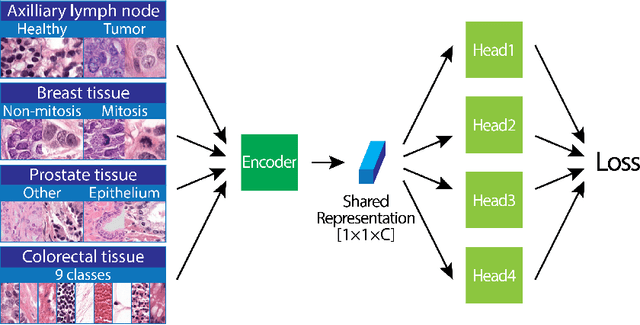
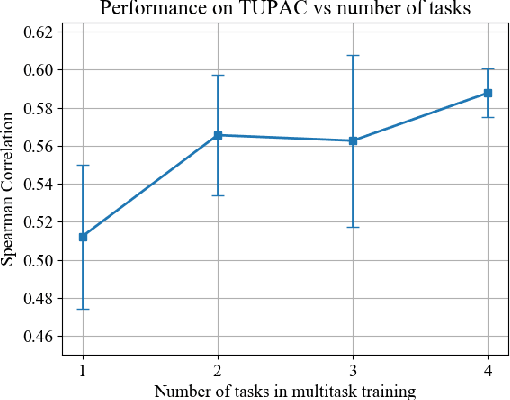
Abstract:We focus on the problem of training convolutional neural networks on gigapixel histopathology images to predict image-level targets. For this purpose, we extend Neural Image Compression (NIC), an image compression framework that reduces the dimensionality of these images using an encoder network trained unsupervisedly. We propose to train this encoder using supervised multitask learning (MTL) instead. We applied the proposed MTL NIC to two histopathology datasets and three tasks. First, we obtained state-of-the-art results in the Tumor Proliferation Assessment Challenge of 2016 (TUPAC16). Second, we successfully classified histopathological growth patterns in images with colorectal liver metastasis (CLM). Third, we predicted patient risk of death by learning directly from overall survival in the same CLM data. Our experimental results suggest that the representations learned by the MTL objective are: (1) highly specific, due to the supervised training signal, and (2) transferable, since the same features perform well across different tasks. Additionally, we trained multiple encoders with different training objectives, e.g. unsupervised and variants of MTL, and observed a positive correlation between the number of tasks in MTL and the system performance on the TUPAC16 dataset.
Artificial Intelligence Assistance Significantly Improves Gleason Grading of Prostate Biopsies by Pathologists
Feb 11, 2020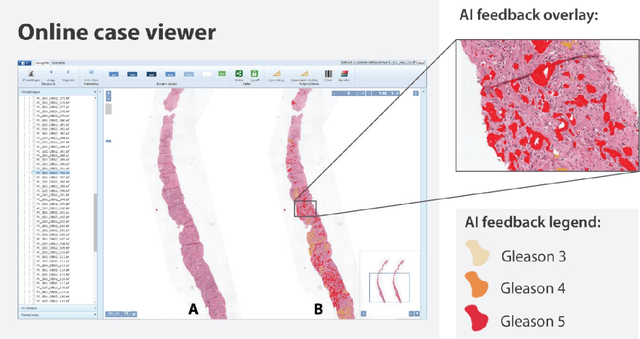
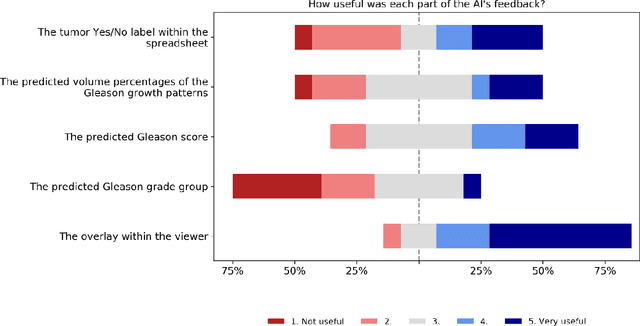
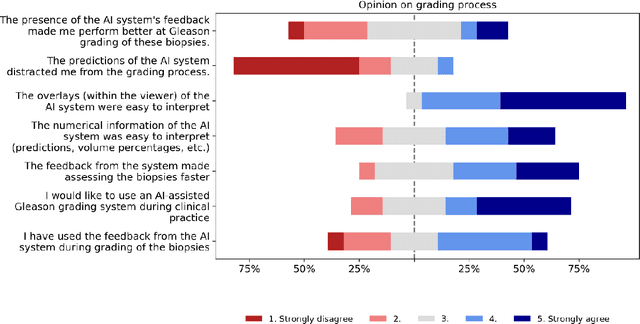
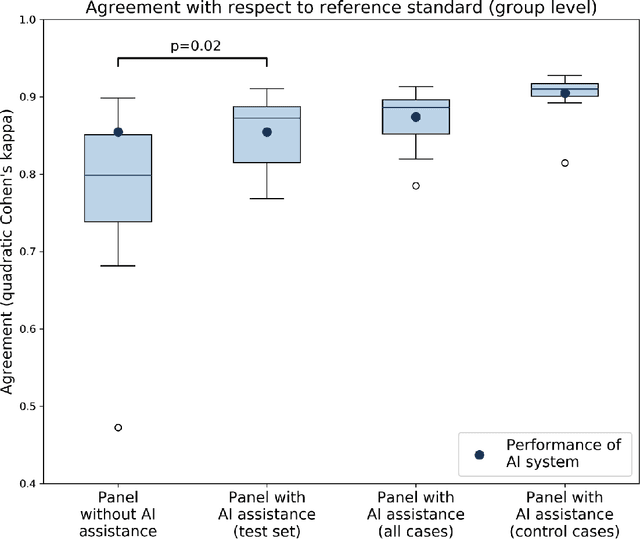
Abstract:While the Gleason score is the most important prognostic marker for prostate cancer patients, it suffers from significant observer variability. Artificial Intelligence (AI) systems, based on deep learning, have proven to achieve pathologist-level performance at Gleason grading. However, the performance of such systems can degrade in the presence of artifacts, foreign tissue, or other anomalies. Pathologists integrating their expertise with feedback from an AI system could result in a synergy that outperforms both the individual pathologist and the system. Despite the hype around AI assistance, existing literature on this topic within the pathology domain is limited. We investigated the value of AI assistance for grading prostate biopsies. A panel of fourteen observers graded 160 biopsies with and without AI assistance. Using AI, the agreement of the panel with an expert reference standard significantly increased (quadratically weighted Cohen's kappa, 0.799 vs 0.872; p=0.018). Our results show the added value of AI systems for Gleason grading, but more importantly, show the benefits of pathologist-AI synergy.
Automated Gleason Grading of Prostate Biopsies using Deep Learning
Jul 18, 2019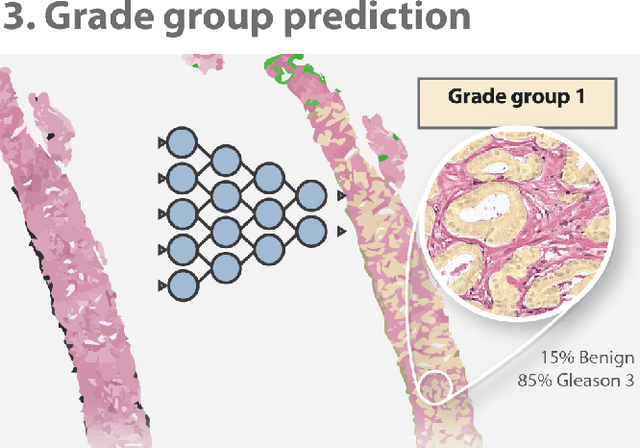
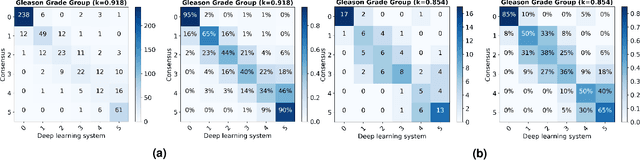
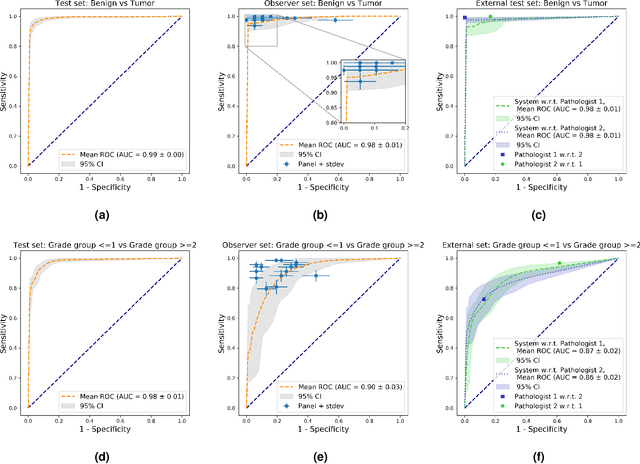
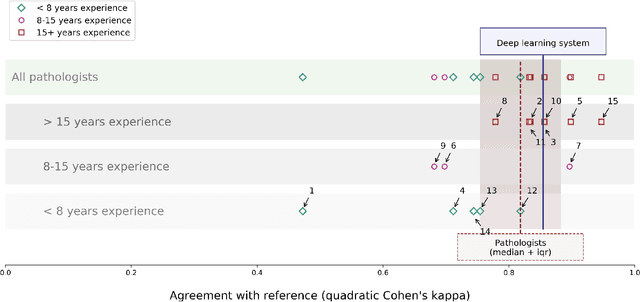
Abstract:The Gleason score is the most important prognostic marker for prostate cancer patients but suffers from significant inter-observer variability. We developed a fully automated deep learning system to grade prostate biopsies. The system was developed using 5834 biopsies from 1243 patients. A semi-automatic labeling technique was used to circumvent the need for full manual annotation by pathologists. The developed system achieved a high agreement with the reference standard. In a separate observer experiment, the deep learning system outperformed 10 out of 15 pathologists. The system has the potential to improve prostate cancer prognostics by acting as a first or second reader.
Quantifying the effects of data augmentation and stain color normalization in convolutional neural networks for computational pathology
Feb 18, 2019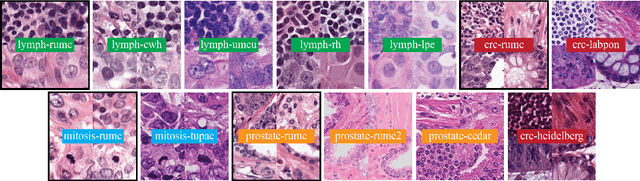
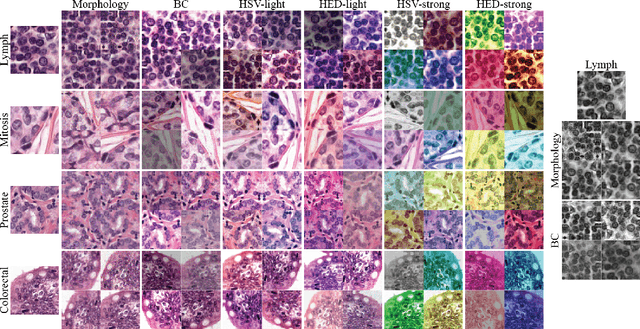
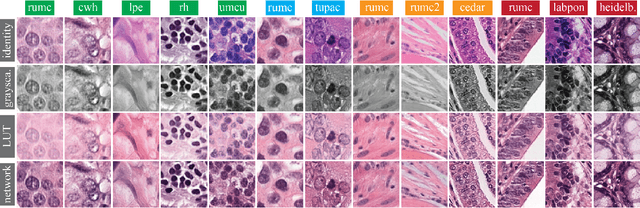
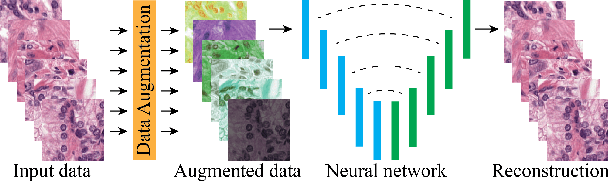
Abstract:Stain variation is a phenomenon observed when distinct pathology laboratories stain tissue slides that exhibit similar but not identical color appearance. Due to this color shift between laboratories, convolutional neural networks (CNNs) trained with images from one lab often underperform on unseen images from the other lab. Several techniques have been proposed to reduce the generalization error, mainly grouped into two categories: stain color augmentation and stain color normalization. The former simulates a wide variety of realistic stain variations during training, producing stain-invariant CNNs. The latter aims to match training and test color distributions in order to reduce stain variation. For the first time, we compared some of these techniques and quantified their effect on CNN classification performance using a heterogeneous dataset of hematoxylin and eosin histopathology images from 4 organs and 9 pathology laboratories. Additionally, we propose a novel unsupervised method to perform stain color normalization using a neural network. Based on our experimental results, we provide practical guidelines on how to use stain color augmentation and stain color normalization in future computational pathology applications.
Neural Image Compression for Gigapixel Histopathology Image Analysis
Nov 07, 2018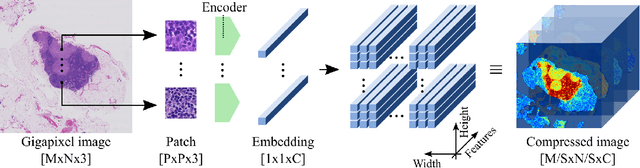
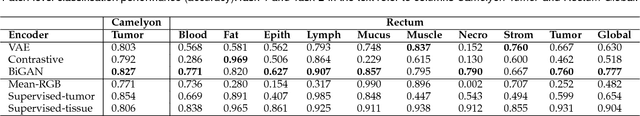
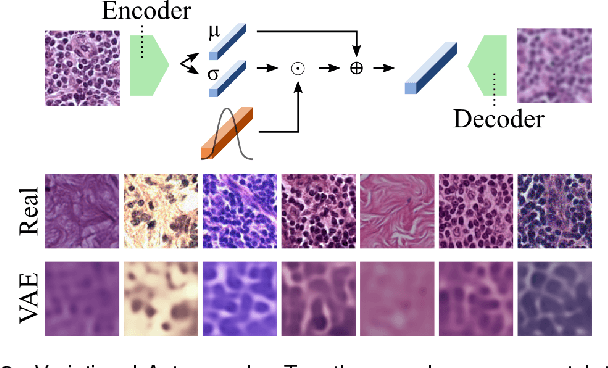
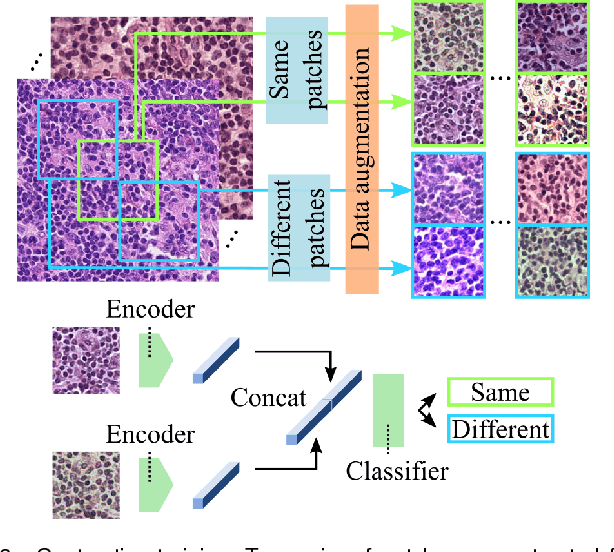
Abstract:We present Neural Image Compression (NIC), a method to reduce the size of gigapixel images by mapping them to a compact latent space using neural networks. We show that this compression allows us to train convolutional neural networks on histopathology whole-slide images end-to-end using weak image-level labels.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge