Jeremy Lynch
A Position Statement on Endovascular Models and Effectiveness Metrics for Mechanical Thrombectomy Navigation, on behalf of the Stakeholder Taskforce for AI-assisted Robotic Thrombectomy (START)
Mar 30, 2026Abstract:While we are making progress in overcoming infectious diseases and cancer; one of the major medical challenges of the mid-21st century will be the rising prevalence of stroke. Large vessels occlusions are especially debilitating, yet effective treatment (needed within hours to achieve best outcomes) remains limited due to geography. One solution for improving timely access to mechanical thrombectomy in geographically diverse populations is the deployment of robotic surgical systems. Artificial intelligence (AI) assistance may enable the upskilling of operators in this emerging therapeutic delivery approach. Our aim was to establish consensus frameworks for developing and validating AI-assisted robots for thrombectomy. Objectives included standardizing effectiveness metrics and defining reference testbeds across in silico, in vitro, ex vivo, and in vivo environments. To achieve this, we convened experts in neurointervention, robotics, data science, health economics, policy, statistics, and patient advocacy. Consensus was built through an incubator day, a Delphi process, and a final Position Statement. We identified that the four essential testbed environments each had distinct validation roles. Realism requirements vary: simpler testbeds should include realistic vessel anatomy compatible with guidewire and catheter use, while standard testbeds should incorporate deformable vessels. More advanced testbeds should include blood flow, pulsatility, and disease features. There are two macro-classes of effectiveness metrics: one for in silico, in vitro, and ex vivo stages focusing on technical navigation, and another for in vivo stages, focused on clinical outcomes. Patient safety is central to this technology's development. One requisite patient safety task needed now is to correlate in vitro measurements to in vivo complications.
* Published in Journal of the American Heart Association
Automated triaging of head MRI examinations using convolutional neural networks
Jun 15, 2021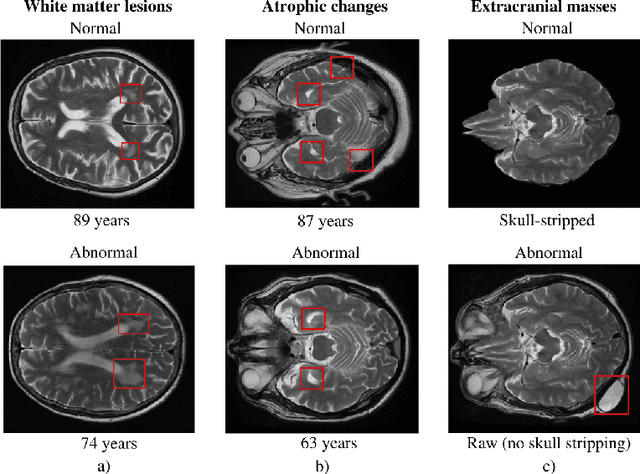
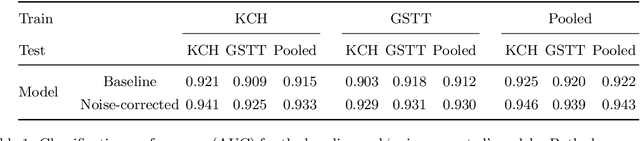
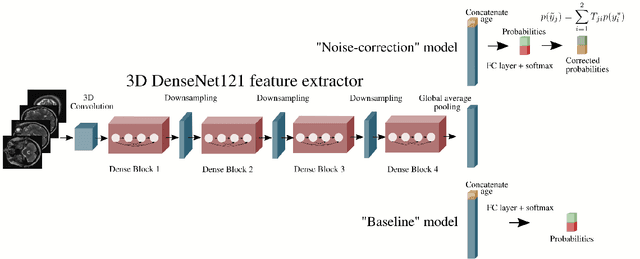
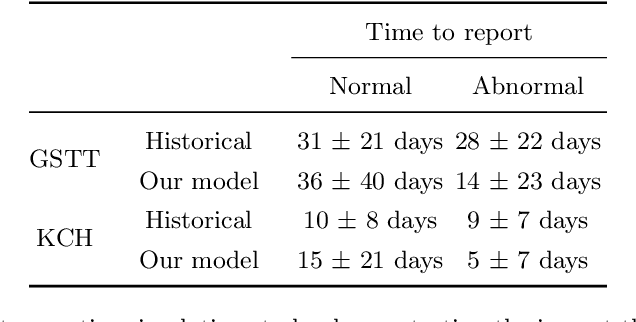
Abstract:The growing demand for head magnetic resonance imaging (MRI) examinations, along with a global shortage of radiologists, has led to an increase in the time taken to report head MRI scans around the world. For many neurological conditions, this delay can result in increased morbidity and mortality. An automated triaging tool could reduce reporting times for abnormal examinations by identifying abnormalities at the time of imaging and prioritizing the reporting of these scans. In this work, we present a convolutional neural network for detecting clinically-relevant abnormalities in $\text{T}_2$-weighted head MRI scans. Using a validated neuroradiology report classifier, we generated a labelled dataset of 43,754 scans from two large UK hospitals for model training, and demonstrate accurate classification (area under the receiver operating curve (AUC) = 0.943) on a test set of 800 scans labelled by a team of neuroradiologists. Importantly, when trained on scans from only a single hospital the model generalized to scans from the other hospital ($\Delta$AUC $\leq$ 0.02). A simulation study demonstrated that our model would reduce the mean reporting time for abnormal examinations from 28 days to 14 days and from 9 days to 5 days at the two hospitals, demonstrating feasibility for use in a clinical triage environment.
Labelling imaging datasets on the basis of neuroradiology reports: a validation study
Jul 08, 2020
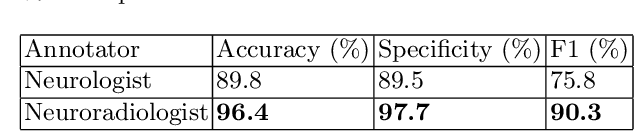

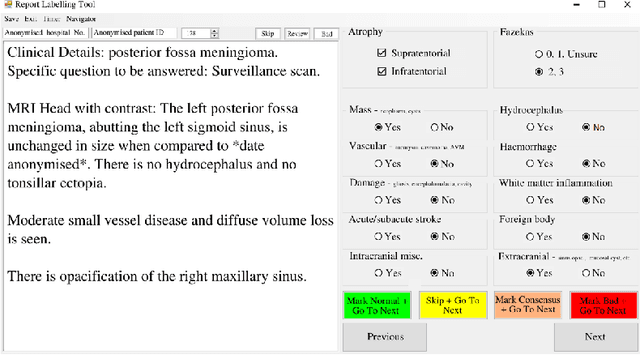

Abstract:Natural language processing (NLP) shows promise as a means to automate the labelling of hospital-scale neuroradiology magnetic resonance imaging (MRI) datasets for computer vision applications. To date, however, there has been no thorough investigation into the validity of this approach, including determining the accuracy of report labels compared to image labels as well as examining the performance of non-specialist labellers. In this work, we draw on the experience of a team of neuroradiologists who labelled over 5000 MRI neuroradiology reports as part of a project to build a dedicated deep learning-based neuroradiology report classifier. We show that, in our experience, assigning binary labels (i.e. normal vs abnormal) to images from reports alone is highly accurate. In contrast to the binary labels, however, the accuracy of more granular labelling is dependent on the category, and we highlight reasons for this discrepancy. We also show that downstream model performance is reduced when labelling of training reports is performed by a non-specialist. To allow other researchers to accelerate their research, we make our refined abnormality definitions and labelling rules available, as well as our easy-to-use radiology report labelling app which helps streamline this process.
Automated Labelling using an Attention model for Radiology reports of MRI scans (ALARM)
Feb 16, 2020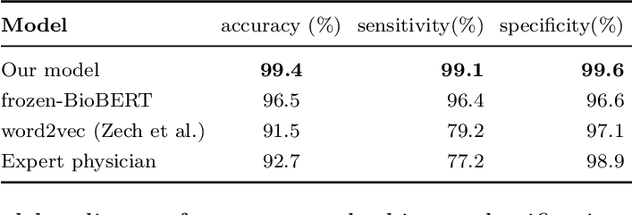
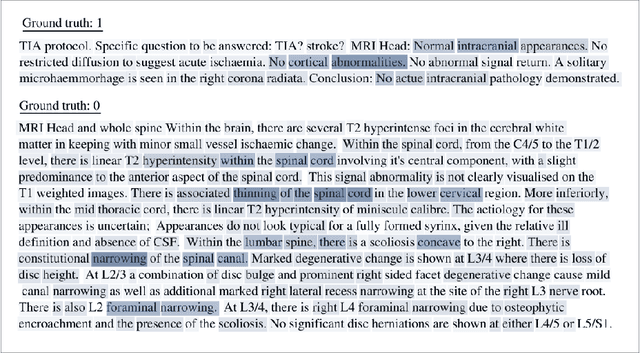
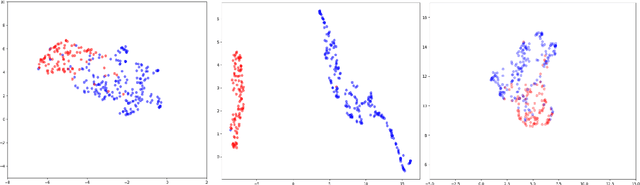
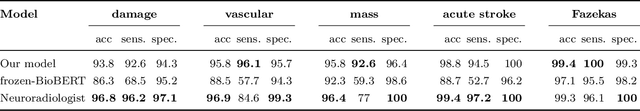

Abstract:Labelling large datasets for training high-capacity neural networks is a major obstacle to the development of deep learning-based medical imaging applications. Here we present a transformer-based network for magnetic resonance imaging (MRI) radiology report classification which automates this task by assigning image labels on the basis of free-text expert radiology reports. Our model's performance is comparable to that of an expert radiologist, and better than that of an expert physician, demonstrating the feasibility of this approach. We make code available online for researchers to label their own MRI datasets for medical imaging applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge