Jennifer Dy
Spectral Non-Convex Optimization for Dimension Reduction with Hilbert-Schmidt Independence Criterion
Sep 06, 2019
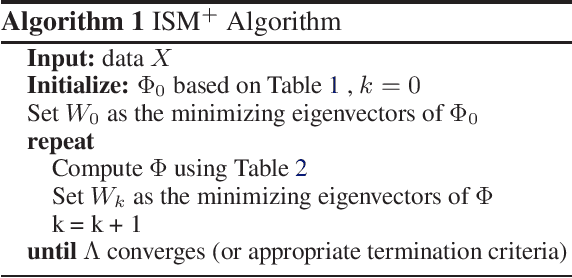
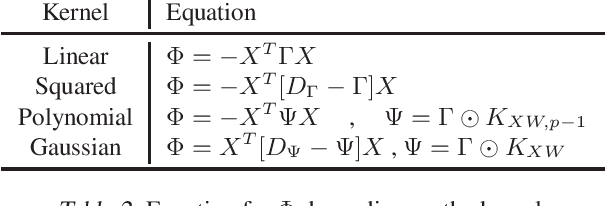


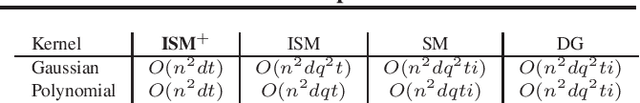
Abstract:The Hilbert Schmidt Independence Criterion (HSIC) is a kernel dependence measure that has applications in various aspects of machine learning. Conveniently, the objectives of different dimensionality reduction applications using HSIC often reduce to the same optimization problem. However, the nonconvexity of the objective function arising from non-linear kernels poses a serious challenge to optimization efficiency and limits the potential of HSIC-based formulations. As a result, only linear kernels have been computationally tractable in practice. This paper proposes a spectral-based optimization algorithm that extends beyond the linear kernel. The algorithm identifies a family of suitable kernels and provides the first and second-order local guarantees when a fixed point is reached. Furthermore, we propose a principled initialization strategy, thereby removing the need to repeat the algorithm at random initialization points. Compared to state-of-the-art optimization algorithms, our empirical results on real data show a run-time improvement by as much as a factor of $10^5$ while consistently achieving lower cost and classification/clustering errors. The implementation source code is publicly available on https://github.com/endsley.
Deep Kernel Learning for Clustering
Aug 09, 2019
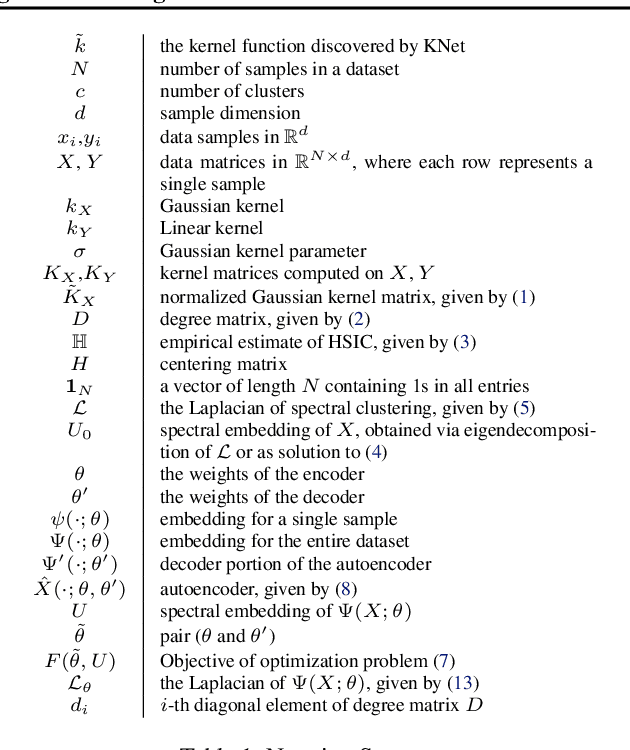

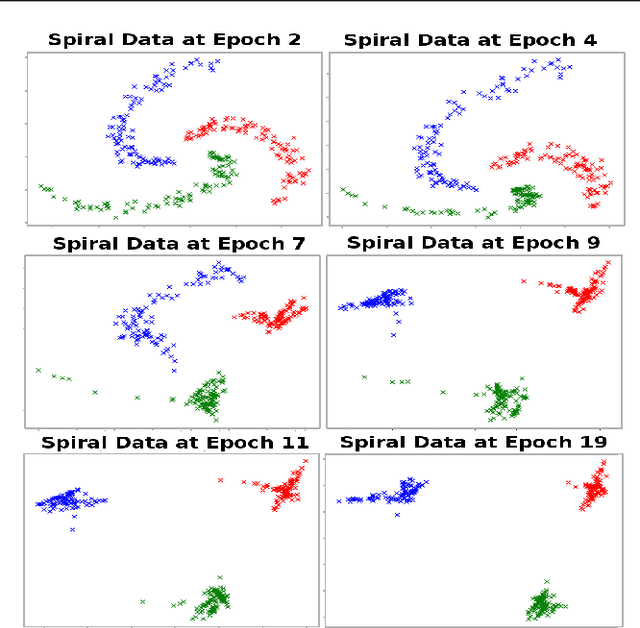
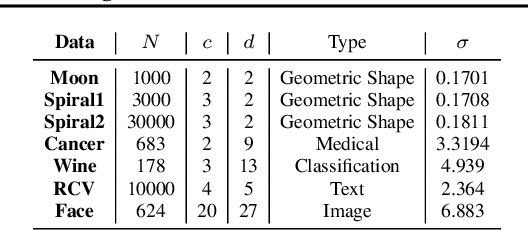
Abstract:We propose a deep learning approach for discovering kernels tailored to identifying clusters over sample data. Our neural network produces sample embeddings that are motivated by--and are at least as expressive as--spectral clustering. Our training objective, based on the Hilbert Schmidt Information Criterion, can be optimized via gradient adaptations on the Stiefel manifold, leading to significant acceleration over spectral methods relying on eigendecompositions. Finally, our trained embedding can be directly applied to out-of-sample data. We show experimentally that our approach outperforms several state-of-the-art deep clustering methods, as well as traditional approaches such as $k$-means and spectral clustering over a broad array of real-life and synthetic datasets.
Neural Topographic Factor Analysis for fMRI Data
Jun 21, 2019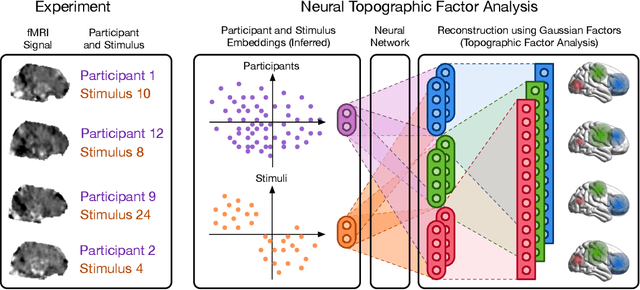
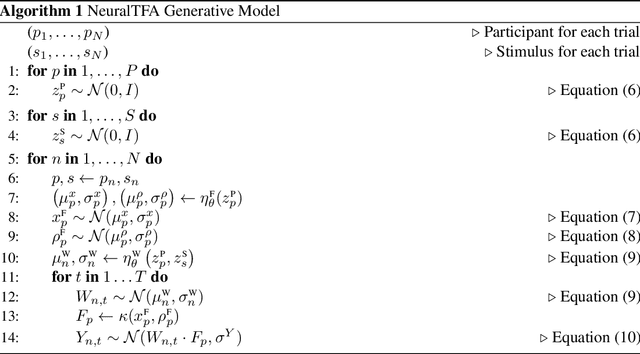

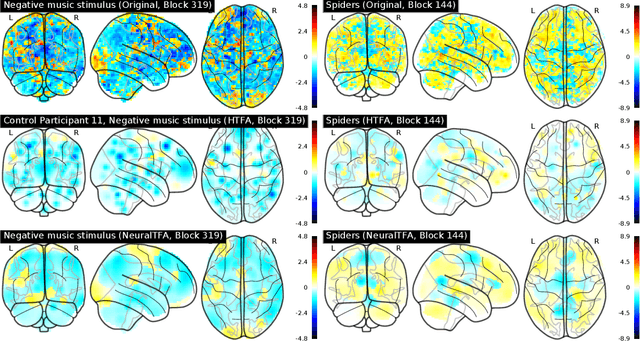
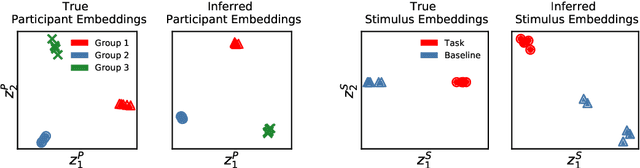
Abstract:Neuroimaging experiments produce a large volume (gigabytes) of high-dimensional spatio-temporal data for a small number of sampled participants and stimuli. Analyses of this data commonly compute averages over all trials, ignoring variation within groups of participants and stimuli. To enable the analysis of fMRI data without this implicit assumption of uniformity, we propose Neural Topographic Factor Analysis (NTFA), a deep generative model that parameterizes factors as functions of embeddings for participants and stimuli. We evaluate NTFA on a synthetically generated dataset as well as on three datasets from fMRI experiments. Our results demonstrate that NTFA yields more accurate reconstructions than a state-of-the-art method with fewer parameters. Moreover, learned embeddings uncover latent categories of participants and stimuli, which suggests that NTFA takes a first step towards reasoning about individual variation in fMRI experiments.
Accelerated Experimental Design for Pairwise Comparisons
Jan 18, 2019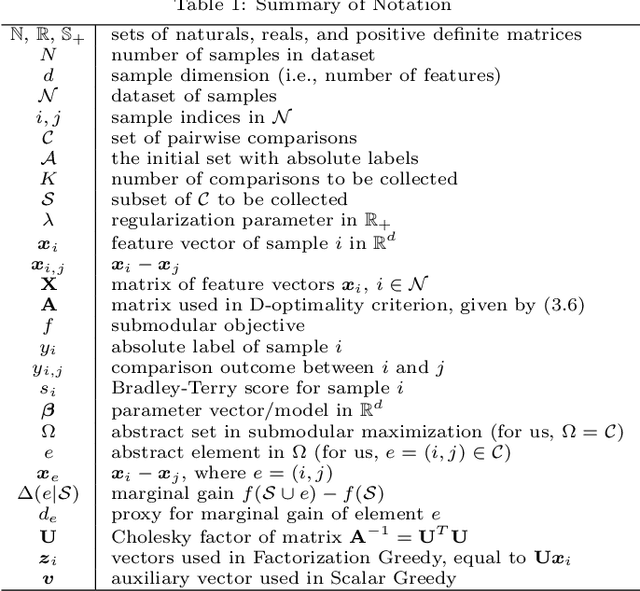
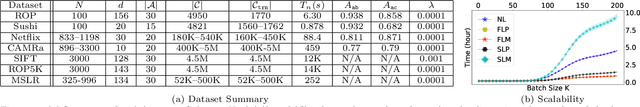
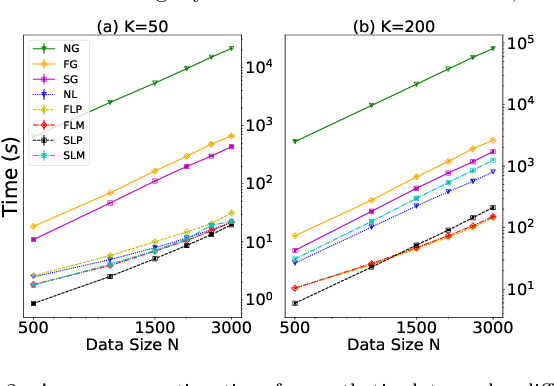
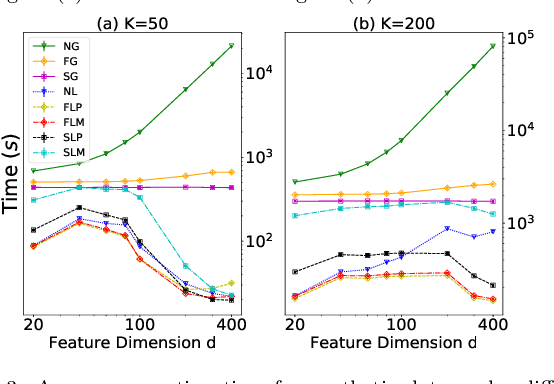
Abstract:Pairwise comparison labels are more informative and less variable than class labels, but generating them poses a challenge: their number grows quadratically in the dataset size. We study a natural experimental design objective, namely, D-optimality, that can be used to identify which $K$ pairwise comparisons to generate. This objective is known to perform well in practice, and is submodular, making the selection approximable via the greedy algorithm. A na\"ive greedy implementation has $O(N^2d^2K)$ complexity, where $N$ is the dataset size, $d$ is the feature space dimension, and $K$ is the number of generated comparisons. We show that, by exploiting the inherent geometry of the dataset--namely, that it consists of pairwise comparisons--the greedy algorithm's complexity can be reduced to $O(N^2(K+d)+N(dK+d^2) +d^2K).$ We apply the same acceleration also to the so-called lazy greedy algorithm. When combined, the above improvements lead to an execution time of less than 1 hour for a dataset with $10^8$ comparisons; the na\"ive greedy algorithm on the same dataset would require more than 10 days to terminate.
Deep feature transfer between localization and segmentation tasks
Nov 10, 2018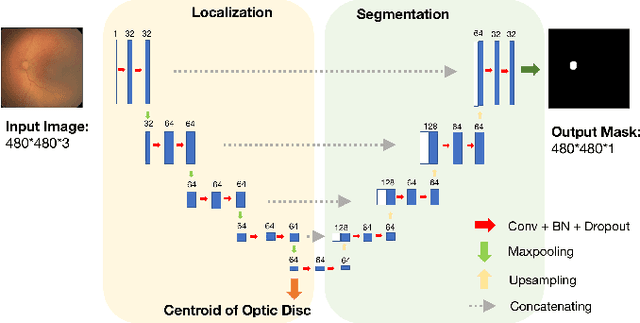
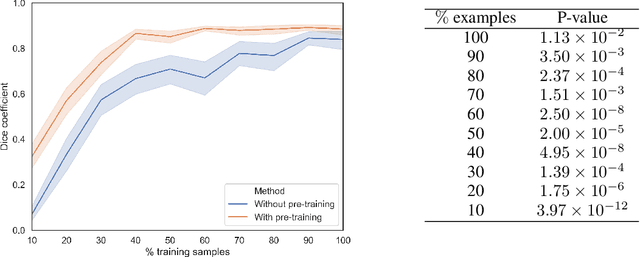
Abstract:In this paper, we propose a new pre-training scheme for U-net based image segmentation. We first train the encoding arm as a localization network to predict the center of the target, before extending it into a U-net architecture for segmentation. We apply our proposed method to the problem of segmenting the optic disc from fundus photographs. Our work shows that the features learned by encoding arm can be transferred to the segmentation network to reduce the annotation burden. We propose that an approach could have broad utility for medical image segmentation, and alleviate the burden of delineating complex structures by pre-training on annotations that are much easier to acquire.
Structured Disentangled Representations
May 29, 2018



Abstract:Deep latent-variable models learn representations of high-dimensional data in an unsupervised manner. A number of recent efforts have focused on learning representations that disentangle statistically independent axes of variation by introducing modifications to the standard objective function. These approaches generally assume a simple diagonal Gaussian prior and as a result are not able to reliably disentangle discrete factors of variation. We propose a two-level hierarchical objective to control relative degree of statistical independence between blocks of variables and individual variables within blocks. We derive this objective as a generalization of the evidence lower bound, which allows us to explicitly represent the trade-offs between mutual information between data and representation, KL divergence between representation and prior, and coverage of the support of the empirical data distribution. Experiments on a variety of datasets demonstrate that our objective can not only disentangle discrete variables, but that doing so also improves disentanglement of other variables and, importantly, generalization even to unseen combinations of factors.
Quantifying Uncertainty in Discrete-Continuous and Skewed Data with Bayesian Deep Learning
May 24, 2018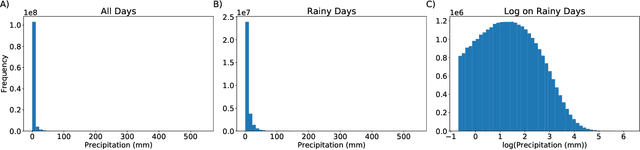
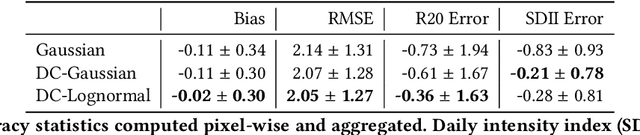
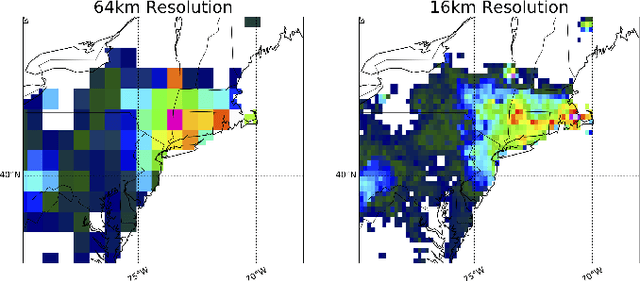
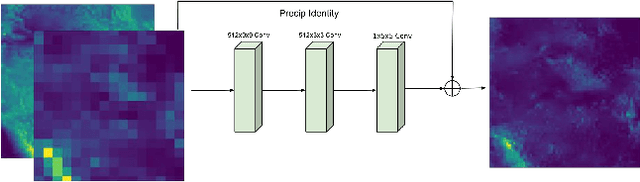
Abstract:Deep Learning (DL) methods have been transforming computer vision with innovative adaptations to other domains including climate change. For DL to pervade Science and Engineering (S&E) applications where risk management is a core component, well-characterized uncertainty estimates must accompany predictions. However, S&E observations and model-simulations often follow heavily skewed distributions and are not well modeled with DL approaches, since they usually optimize a Gaussian, or Euclidean, likelihood loss. Recent developments in Bayesian Deep Learning (BDL), which attempts to capture uncertainties from noisy observations, aleatoric, and from unknown model parameters, epistemic, provide us a foundation. Here we present a discrete-continuous BDL model with Gaussian and lognormal likelihoods for uncertainty quantification (UQ). We demonstrate the approach by developing UQ estimates on `DeepSD', a super-resolution based DL model for Statistical Downscaling (SD) in climate applied to precipitation, which follows an extremely skewed distribution. We find that the discrete-continuous models outperform a basic Gaussian distribution in terms of predictive accuracy and uncertainty calibration. Furthermore, we find that the lognormal distribution, which can handle skewed distributions, produces quality uncertainty estimates at the extremes. Such results may be important across S&E, as well as other domains such as finance and economics, where extremes are often of significant interest. Furthermore, to our knowledge, this is the first UQ model in SD where both aleatoric and epistemic uncertainties are characterized.
* 10 Pages
Evaluating Crowdsourcing Participants in the Absence of Ground-Truth
May 30, 2016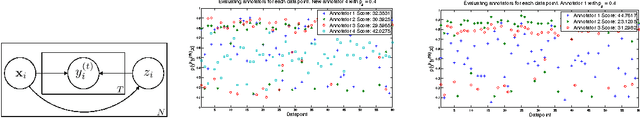
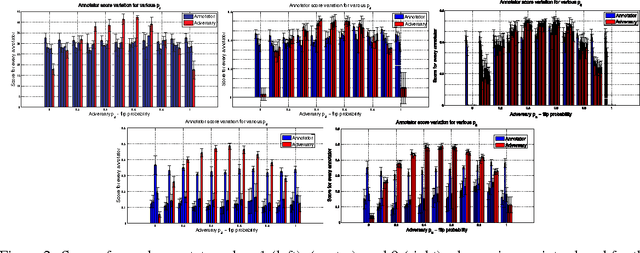
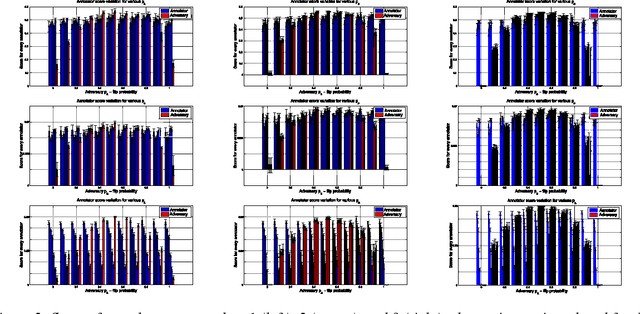
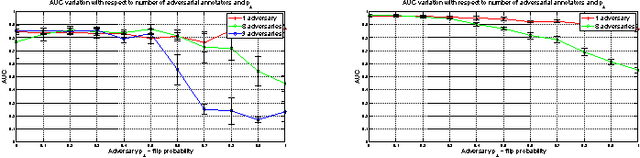
Abstract:Given a supervised/semi-supervised learning scenario where multiple annotators are available, we consider the problem of identification of adversarial or unreliable annotators.
Modeling Multiple Annotator Expertise in the Semi-Supervised Learning Scenario
Mar 15, 2012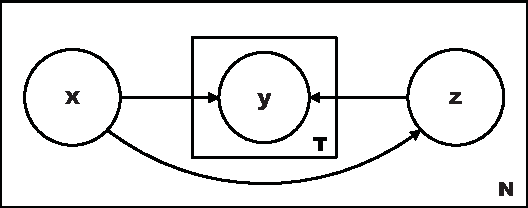
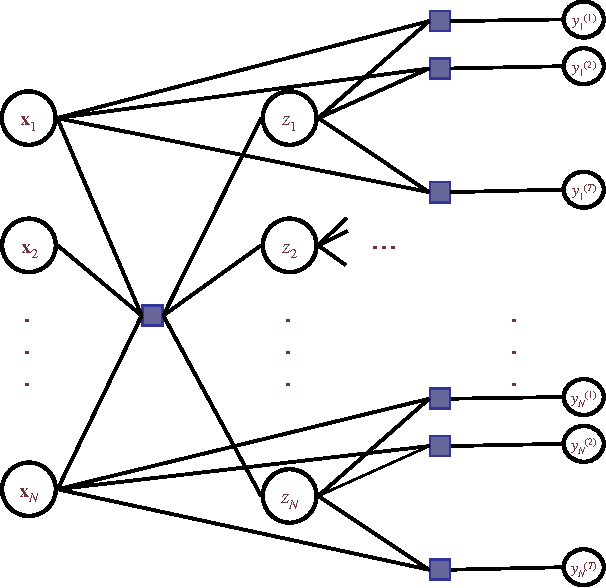
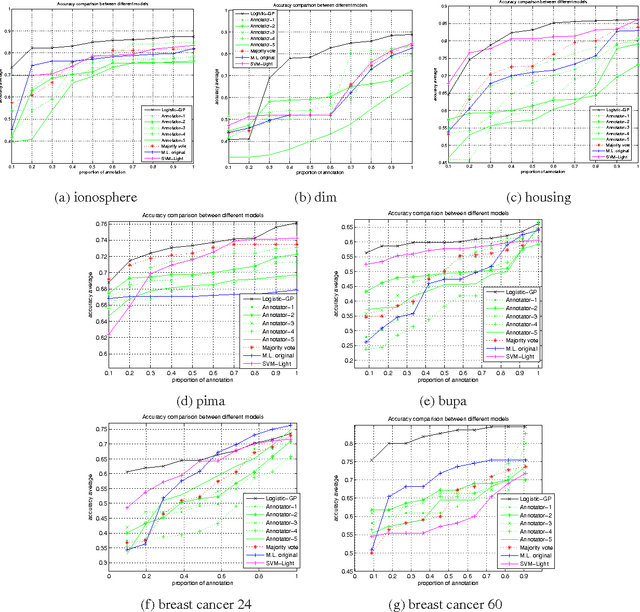
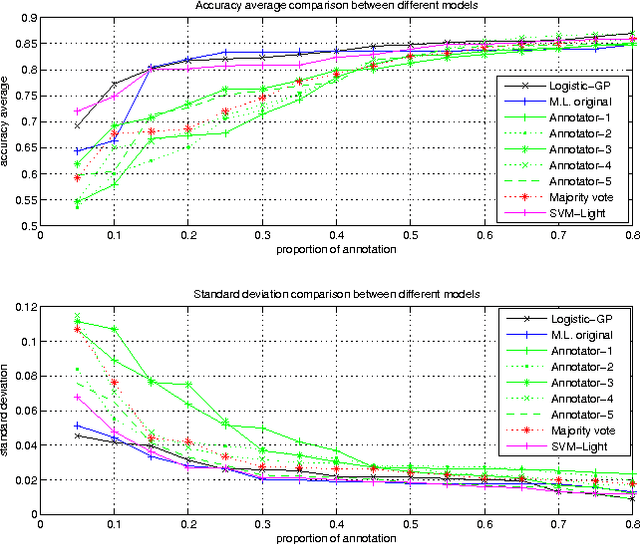
Abstract:Learning algorithms normally assume that there is at most one annotation or label per data point. However, in some scenarios, such as medical diagnosis and on-line collaboration,multiple annotations may be available. In either case, obtaining labels for data points can be expensive and time-consuming (in some circumstances ground-truth may not exist). Semi-supervised learning approaches have shown that utilizing the unlabeled data is often beneficial in these cases. This paper presents a probabilistic semi-supervised model and algorithm that allows for learning from both unlabeled and labeled data in the presence of multiple annotators. We assume that it is known what annotator labeled which data points. The proposed approach produces annotator models that allow us to provide (1) estimates of the true label and (2) annotator variable expertise for both labeled and unlabeled data. We provide numerical comparisons under various scenarios and with respect to standard semi-supervised learning. Experiments showed that the presented approach provides clear advantages over multi-annotator methods that do not use the unlabeled data and over methods that do not use multi-labeler information.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge