Jan Witowski
A Multi-Modal AI System for Screening Mammography: Integrating 2D and 3D Imaging to Improve Breast Cancer Detection in a Prospective Clinical Study
Apr 08, 2025



Abstract:Although digital breast tomosynthesis (DBT) improves diagnostic performance over full-field digital mammography (FFDM), false-positive recalls remain a concern in breast cancer screening. We developed a multi-modal artificial intelligence system integrating FFDM, synthetic mammography, and DBT to provide breast-level predictions and bounding-box localizations of suspicious findings. Our AI system, trained on approximately 500,000 mammography exams, achieved 0.945 AUROC on an internal test set. It demonstrated capacity to reduce recalls by 31.7% and radiologist workload by 43.8% while maintaining 100% sensitivity, underscoring its potential to improve clinical workflows. External validation confirmed strong generalizability, reducing the gap to a perfect AUROC by 35.31%-69.14% relative to strong baselines. In prospective deployment across 18 sites, the system reduced recall rates for low-risk cases. An improved version, trained on over 750,000 exams with additional labels, further reduced the gap by 18.86%-56.62% across large external datasets. Overall, these results underscore the importance of utilizing all available imaging modalities, demonstrate the potential for clinical impact, and indicate feasibility of further reduction of the test error with increased training set when using large-capacity neural networks.
Multi-modal AI for comprehensive breast cancer prognostication
Oct 28, 2024


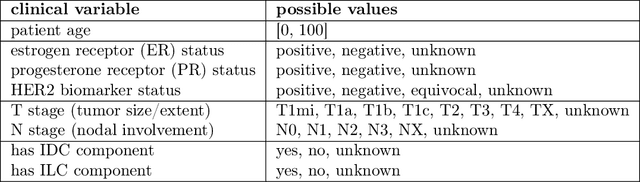
Abstract:Treatment selection in breast cancer is guided by molecular subtypes and clinical characteristics. Recurrence risk assessment plays a crucial role in personalizing treatment. Current methods, including genomic assays, have limited accuracy and clinical utility, leading to suboptimal decisions for many patients. We developed a test for breast cancer patient stratification based on digital pathology and clinical characteristics using novel AI methods. Specifically, we utilized a vision transformer-based pan-cancer foundation model trained with self-supervised learning to extract features from digitized H&E-stained slides. These features were integrated with clinical data to form a multi-modal AI test predicting cancer recurrence and death. The test was developed and evaluated using data from a total of 8,161 breast cancer patients across 15 cohorts originating from seven countries. Of these, 3,502 patients from five cohorts were used exclusively for evaluation, while the remaining patients were used for training. Our test accurately predicted our primary endpoint, disease-free interval, in the five external cohorts (C-index: 0.71 [0.68-0.75], HR: 3.63 [3.02-4.37, p<0.01]). In a direct comparison (N=858), the AI test was more accurate than Oncotype DX, the standard-of-care 21-gene assay, with a C-index of 0.67 [0.61-0.74] versus 0.61 [0.49-0.73], respectively. Additionally, the AI test added independent information to Oncotype DX in a multivariate analysis (HR: 3.11 [1.91-5.09, p<0.01)]). The test demonstrated robust accuracy across all major breast cancer subtypes, including TNBC (C-index: 0.71 [0.62-0.81], HR: 3.81 [2.35-6.17, p=0.02]), where no diagnostic tools are currently recommended by clinical guidelines. These results suggest that our AI test can improve accuracy, extend applicability to a wider range of patients, and enhance access to treatment selection tools.
3D-GMIC: an efficient deep neural network to find small objects in large 3D images
Oct 16, 2022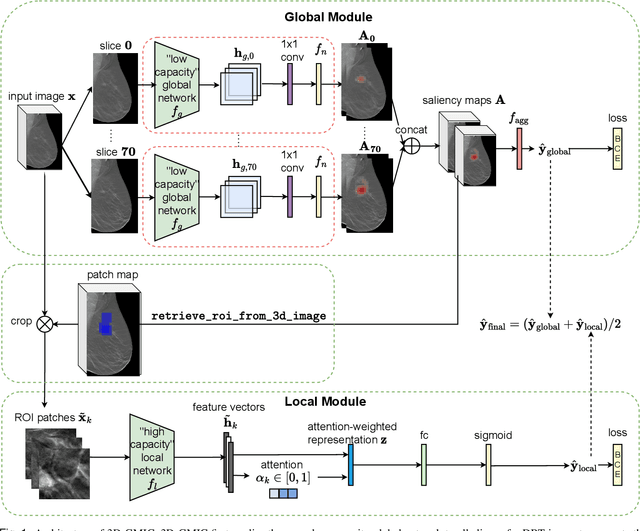
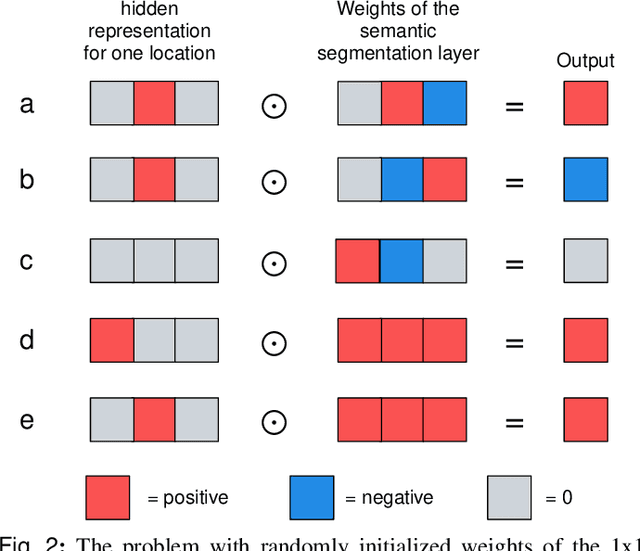
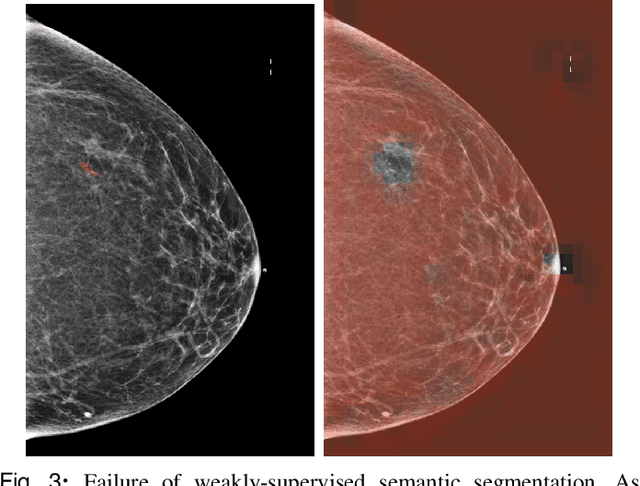
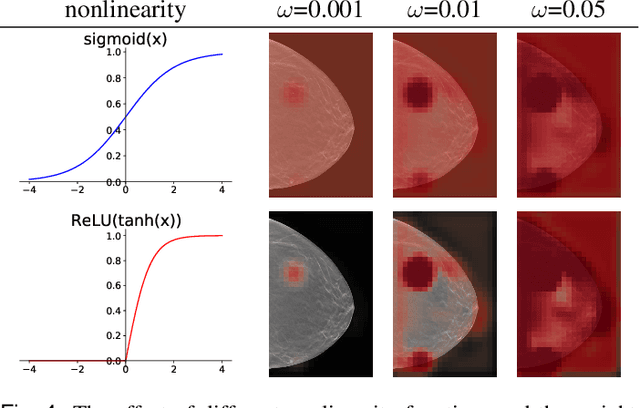
Abstract:3D imaging enables a more accurate diagnosis by providing spatial information about organ anatomy. However, using 3D images to train AI models is computationally challenging because they consist of tens or hundreds of times more pixels than their 2D counterparts. To train with high-resolution 3D images, convolutional neural networks typically resort to downsampling them or projecting them to two dimensions. In this work, we propose an effective alternative, a novel neural network architecture that enables computationally efficient classification of 3D medical images in their full resolution. Compared to off-the-shelf convolutional neural networks, 3D-GMIC uses 77.98%-90.05% less GPU memory and 91.23%-96.02% less computation. While our network is trained only with image-level labels, without segmentation labels, it explains its classification predictions by providing pixel-level saliency maps. On a dataset collected at NYU Langone Health, including 85,526 patients with full-field 2D mammography (FFDM), synthetic 2D mammography, and 3D mammography (DBT), our model, the 3D Globally-Aware Multiple Instance Classifier (3D-GMIC), achieves a breast-wise AUC of 0.831 (95% CI: 0.769-0.887) in classifying breasts with malignant findings using DBT images. As DBT and 2D mammography capture different information, averaging predictions on 2D and 3D mammography together leads to a diverse ensemble with an improved breast-wise AUC of 0.841 (95% CI: 0.768-0.895). Our model generalizes well to an external dataset from Duke University Hospital, achieving an image-wise AUC of 0.848 (95% CI: 0.798-0.896) in classifying DBT images with malignant findings.
Meta-repository of screening mammography classifiers
Aug 10, 2021
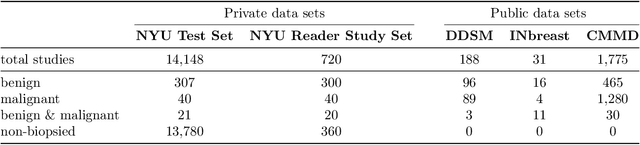
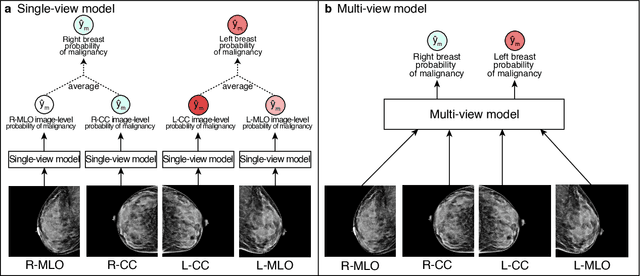
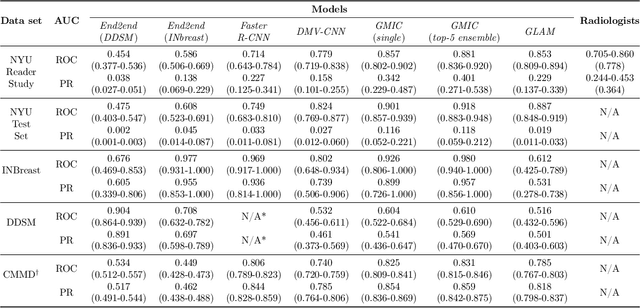
Abstract:Artificial intelligence (AI) is transforming medicine and showing promise in improving clinical diagnosis. In breast cancer screening, several recent studies show that AI has the potential to improve radiologists' accuracy, subsequently helping in early cancer diagnosis and reducing unnecessary workup. As the number of proposed models and their complexity grows, it is becoming increasingly difficult to re-implement them in order to reproduce the results and to compare different approaches. To enable reproducibility of research in this application area and to enable comparison between different methods, we release a meta-repository containing deep learning models for classification of screening mammograms. This meta-repository creates a framework that enables the evaluation of machine learning models on any private or public screening mammography data set. At its inception, our meta-repository contains five state-of-the-art models with open-source implementations and cross-platform compatibility. We compare their performance on five international data sets: two private New York University breast cancer screening data sets as well as three public (DDSM, INbreast and Chinese Mammography Database) data sets. Our framework has a flexible design that can be generalized to other medical image analysis tasks. The meta-repository is available at https://www.github.com/nyukat/mammography_metarepository.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge