J. Mason Earles
Does Your VFM Speak Plant? The Botanical Grammar of Vision Foundation Models for Object Detection
Apr 10, 2026Abstract:Vision foundation models (VFMs) offer the promise of zero-shot object detection without task-specific training data, yet their performance in complex agricultural scenes remains highly sensitive to text prompt construction. We present a systematic prompt optimization framework evaluating four open-vocabulary detectors -- YOLO World, SAM3, Grounding DINO, and OWLv2 -- for cowpea flower and pod detection across synthetic and real field imagery. We decompose prompts into eight axes and conduct one-factor-at-a-time analysis followed by combinatorial optimization, revealing that models respond divergently to prompt structure: conditions that optimize one architecture can collapse another. Applying model-specific combinatorial prompts yields substantial gains over a naive species-name baseline, including +0.357 mAP@0.5 for YOLO World and +0.362 mAP@0.5 for OWLv2 on synthetic cowpea flower data. To evaluate cross-task generalization, we use an LLM to translate the discovered axis structure to a morphologically distinct target -- cowpea pods -- and compare against prompting using the discovered optimal structures from synthetic flower data. Crucially, prompt structures optimized exclusively on synthetic data transfer effectively to real-world fields: synthetic-pipeline prompts match or exceed those discovered on labeled real data for the majority of model-object combinations (flower: 0.374 vs. 0.353 for YOLO World; pod: 0.429 vs. 0.371 for SAM3). Our findings demonstrate that prompt engineering can substantially close the gap between zero-shot VFMs and supervised detectors without requiring manual annotation, and that optimal prompts are model-specific, non-obvious, and transferable across domains.
A Vision Language Model for Generating Procedural Plant Architecture Representations from Simulated Images
Mar 23, 2026Abstract:Three-dimensional (3D) procedural plant architecture models have emerged as an important tool for simulation-based studies of plant structure and function, extracting plant architectural parameters from field measurements, and for generating realistic plants in computer graphics. However, measuring the architectural parameters and nested structures for these models at the field scales remains prohibitively labor-intensive. We present a novel algorithm that generates a 3D plant architecture from an image, creating a functional structural plant model that reflects organ-level geometric and topological parameters and provides a more comprehensive representation of the plant's architecture. Instead of using 3D sensors or processing multi-view images with computer vision to obtain the 3D structure of plants, we proposed a method that generates token sequences that encode a procedural definition of plant architecture. This work used only synthetic images for training and testing, with exact architectural parameters known, allowing testing of the hypothesis that organ-level architectural parameters could be extracted from image data using a vision-language model (VLM). A synthetic dataset of cowpea plant images was generated using the Helios 3D plant simulator, with the detailed plant architecture encoded in XML files. We developed a plant architecture tokenizer for the XML file defining plant architecture, converting it into a token sequence that a language model can predict. The model achieved a token F1 score of 0.73 during teacher-forced training. Evaluation of the model was performed through autoregressive generation, achieving a BLEU-4 score of 94.00% and a ROUGE-L score of 0.5182. This led to the conclusion that such plant architecture model generation and parameter extraction were possible from synthetic images; thus, future work will extend the approach to real imagery data.
Using Vision Language Foundation Models to Generate Plant Simulation Configurations via In-Context Learning
Mar 09, 2026Abstract:This paper introduces a synthetic benchmark to evaluate the performance of vision language models (VLMs) in generating plant simulation configurations for digital twins. While functional-structural plant models (FSPMs) are useful tools for simulating biophysical processes in agricultural environments, their high complexity and low throughput create bottlenecks for deployment at scale. We propose a novel approach that leverages state-of-the-art open-source VLMs -- Gemma 3 and Qwen3-VL -- to directly generate simulation parameters in JSON format from drone-based remote sensing images. Using a synthetic cowpea plot dataset generated via the Helios 3D procedural plant generation library, we tested five in-context learning methods and evaluated the models across three categories: JSON integrity, geometric evaluations, and biophysical evaluations. Our results show that while VLMs can interpret structural metadata and estimate parameters like plant count and sun azimuth, they often exhibit performance degradation due to contextual bias or rely on dataset means when visual cues are insufficient. Validation on a real-world drone orthophoto dataset and an ablation study using a blind baseline further characterize the models' reasoning capabilities versus their reliance on contextual priors. To the best of our knowledge, this is the first study to utilize VLMs to generate structural JSON configurations for plant simulations, providing a scalable framework for reconstruction 3D plots for digital twin in agriculture.
AGILE: A Diffusion-Based Attention-Guided Image and Label Translation for Efficient Cross-Domain Plant Trait Identification
Mar 27, 2025Abstract:Semantically consistent cross-domain image translation facilitates the generation of training data by transferring labels across different domains, making it particularly useful for plant trait identification in agriculture. However, existing generative models struggle to maintain object-level accuracy when translating images between domains, especially when domain gaps are significant. In this work, we introduce AGILE (Attention-Guided Image and Label Translation for Efficient Cross-Domain Plant Trait Identification), a diffusion-based framework that leverages optimized text embeddings and attention guidance to semantically constrain image translation. AGILE utilizes pretrained diffusion models and publicly available agricultural datasets to improve the fidelity of translated images while preserving critical object semantics. Our approach optimizes text embeddings to strengthen the correspondence between source and target images and guides attention maps during the denoising process to control object placement. We evaluate AGILE on cross-domain plant datasets and demonstrate its effectiveness in generating semantically accurate translated images. Quantitative experiments show that AGILE enhances object detection performance in the target domain while maintaining realism and consistency. Compared to prior image translation methods, AGILE achieves superior semantic alignment, particularly in challenging cases where objects vary significantly or domain gaps are substantial.
VisTA-SR: Improving the Accuracy and Resolution of Low-Cost Thermal Imaging Cameras for Agriculture
May 29, 2024Abstract:Thermal cameras are an important tool for agricultural research because they allow for non-invasive measurement of plant temperature, which relates to important photochemical, hydraulic, and agronomic traits. Utilizing low-cost thermal cameras can lower the barrier to introducing thermal imaging in agricultural research and production. This paper presents an approach to improve the temperature accuracy and image quality of low-cost thermal imaging cameras for agricultural applications. Leveraging advancements in computer vision techniques, particularly deep learning networks, we propose a method, called $\textbf{VisTA-SR}$ ($\textbf{Vis}$ual \& $\textbf{T}$hermal $\textbf{A}$lignment and $\textbf{S}$uper-$\textbf{R}$esolution Enhancement) that combines RGB and thermal images to enhance the capabilities of low-resolution thermal cameras. The research includes calibration and validation of temperature measurements, acquisition of paired image datasets, and the development of a deep learning network tailored for agricultural thermal imaging. Our study addresses the challenges of image enhancement in the agricultural domain and explores the potential of low-cost thermal cameras to replace high-resolution industrial cameras. Experimental results demonstrate the effectiveness of our approach in enhancing temperature accuracy and image sharpness, paving the way for more accessible and efficient thermal imaging solutions in agriculture.
End-to-end deep learning for directly estimating grape yield from ground-based imagery
Aug 04, 2022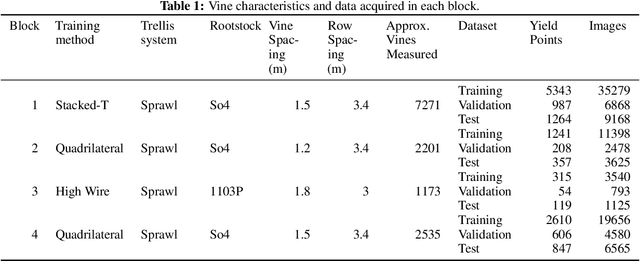
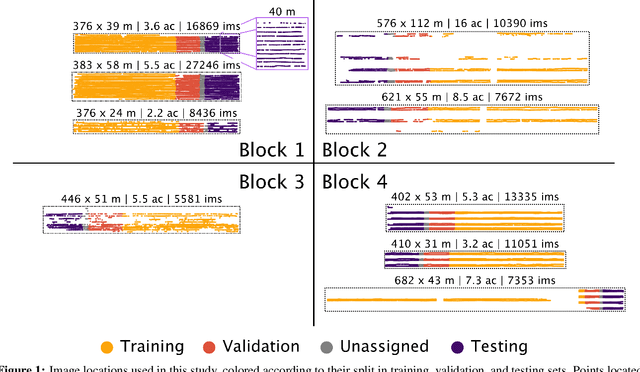
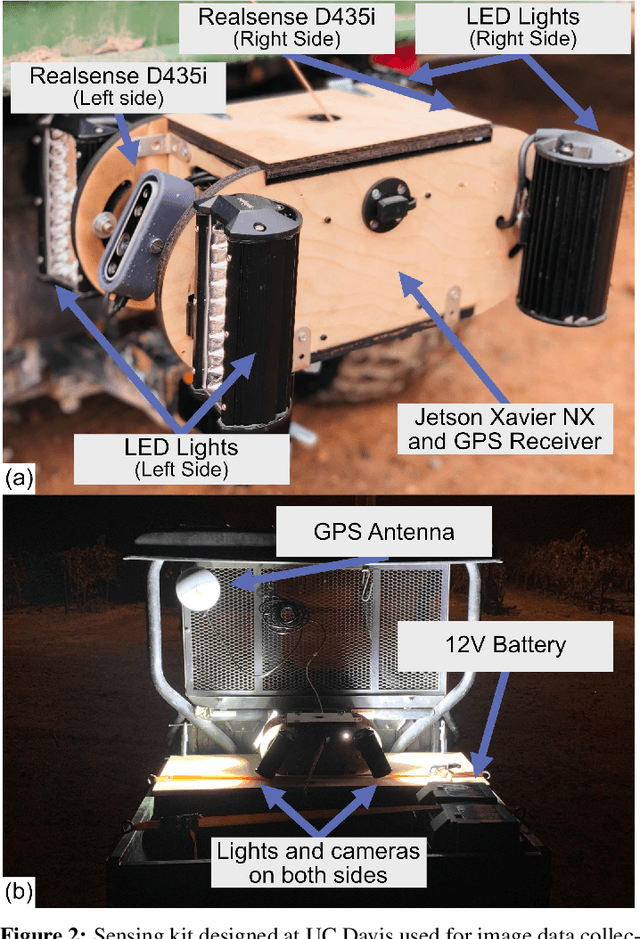
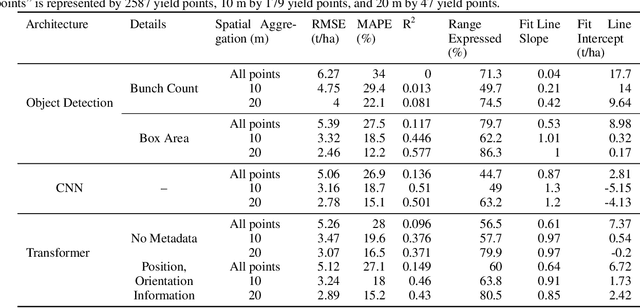
Abstract:Yield estimation is a powerful tool in vineyard management, as it allows growers to fine-tune practices to optimize yield and quality. However, yield estimation is currently performed using manual sampling, which is time-consuming and imprecise. This study demonstrates the application of proximal imaging combined with deep learning for yield estimation in vineyards. Continuous data collection using a vehicle-mounted sensing kit combined with collection of ground truth yield data at harvest using a commercial yield monitor allowed for the generation of a large dataset of 23,581 yield points and 107,933 images. Moreover, this study was conducted in a mechanically managed commercial vineyard, representing a challenging environment for image analysis but a common set of conditions in the California Central Valley. Three model architectures were tested: object detection, CNN regression, and transformer models. The object detection model was trained on hand-labeled images to localize grape bunches, and either bunch count or pixel area was summed to correlate with grape yield. Conversely, regression models were trained end-to-end to predict grape yield from image data without the need for hand labeling. Results demonstrated that both a transformer as well as the object detection model with pixel area processing performed comparably, with a mean absolute percent error of 18% and 18.5%, respectively on a representative holdout dataset. Saliency mapping was used to demonstrate the attention of the CNN model was localized near the predicted location of grape bunches, as well as on the top of the grapevine canopy. Overall, the study showed the applicability of proximal imaging and deep learning for prediction of grapevine yield on a large scale. Additionally, the end-to-end modeling approach was able to perform comparably to the object detection approach while eliminating the need for hand-labeling.
A workflow for segmenting soil and plant X-ray CT images with deep learning in Googles Colaboratory
Mar 18, 2022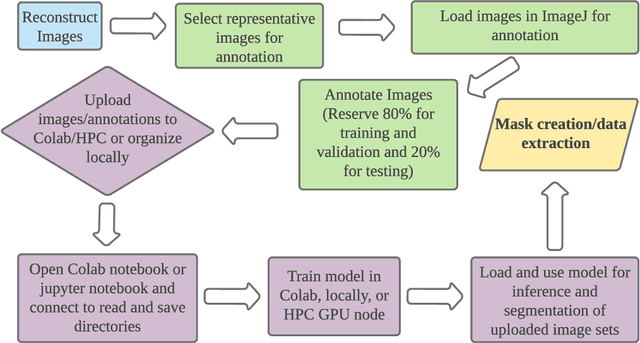
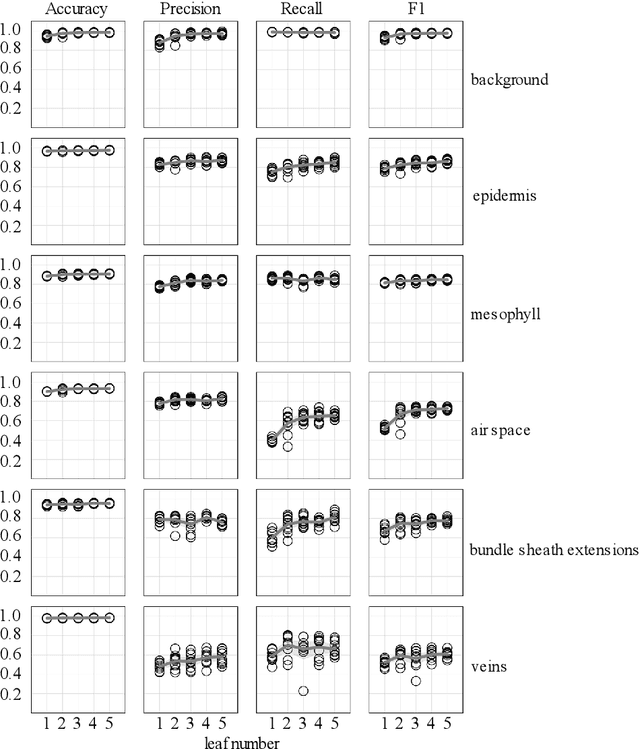
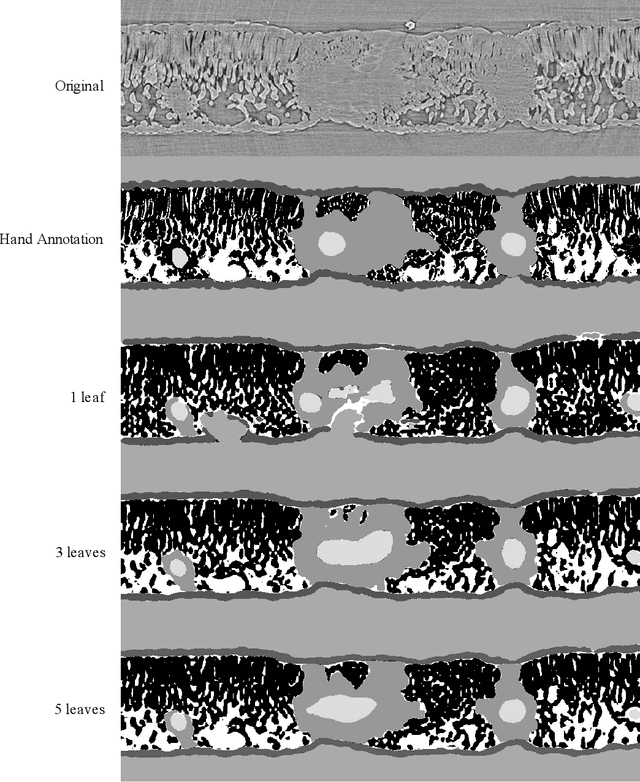
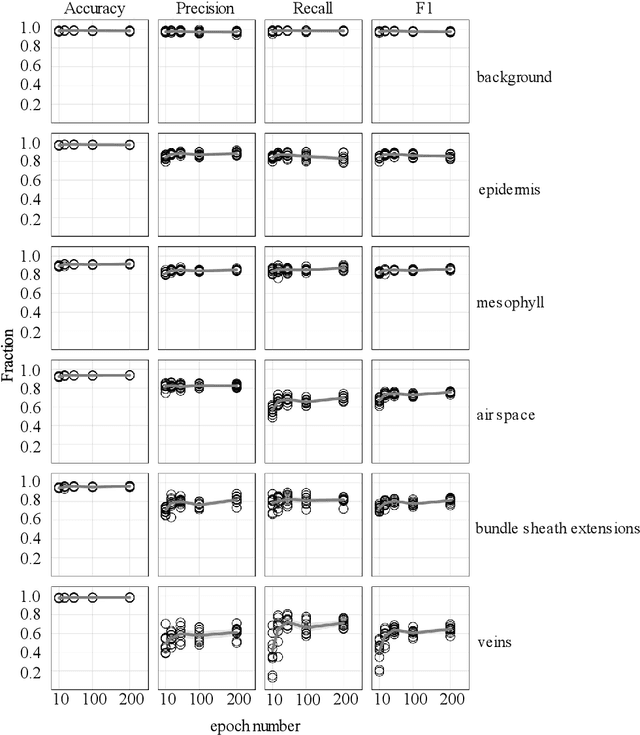
Abstract:X-ray micro-computed tomography (X-ray microCT) has enabled the characterization of the properties and processes that take place in plants and soils at the micron scale. Despite the widespread use of this advanced technique, major limitations in both hardware and software limit the speed and accuracy of image processing and data analysis. Recent advances in machine learning, specifically the application of convolutional neural networks to image analysis, have enabled rapid and accurate segmentation of image data. Yet, challenges remain in applying convolutional neural networks to the analysis of environmentally and agriculturally relevant images. Specifically, there is a disconnect between the computer scientists and engineers, who build these AI/ML tools, and the potential end users in agricultural research, who may be unsure of how to apply these tools in their work. Additionally, the computing resources required for training and applying deep learning models are unique, more common to computer gaming systems or graphics design work, than to traditional computational systems. To navigate these challenges, we developed a modular workflow for applying convolutional neural networks to X-ray microCT images, using low-cost resources in Googles Colaboratory web application. Here we present the results of the workflow, illustrating how parameters can be optimized to achieve best results using example scans from walnut leaves, almond flower buds, and a soil aggregate. We expect that this framework will accelerate the adoption and use of emerging deep learning techniques within the plant and soil sciences.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge