Pranav Raja
ToolComp: A Multi-Tool Reasoning & Process Supervision Benchmark
Jan 02, 2025



Abstract:Despite recent advances in AI, the development of systems capable of executing complex, multi-step reasoning tasks involving multiple tools remains a significant challenge. Current benchmarks fall short in capturing the real-world complexity of tool-use reasoning, where verifying the correctness of not only the final answer but also the intermediate steps is important for evaluation, development, and identifying failures during inference time. To bridge this gap, we introduce ToolComp, a comprehensive benchmark designed to evaluate multi-step tool-use reasoning. ToolComp is developed through a collaboration between models and human annotators, featuring human-edited/verified prompts, final answers, and process supervision labels, allowing for the evaluation of both final outcomes and intermediate reasoning. Evaluation across six different model families demonstrates the challenging nature of our dataset, with the majority of models achieving less than 50% accuracy. Additionally, we generate synthetic training data to compare the performance of outcome-supervised reward models (ORMs) with process-supervised reward models (PRMs) to assess their ability to improve complex tool-use reasoning as evaluated by ToolComp. Our results show that PRMs generalize significantly better than ORMs, achieving a 19% and 11% improvement in rank@1 accuracy for ranking base and fine-tuned model trajectories, respectively. These findings highlight the critical role of process supervision in both the evaluation and training of AI models, paving the way for more robust and capable systems in complex, multi-step tool-use tasks.
A Careful Examination of Large Language Model Performance on Grade School Arithmetic
May 02, 2024



Abstract:Large language models (LLMs) have achieved impressive success on many benchmarks for mathematical reasoning. However, there is growing concern that some of this performance actually reflects dataset contamination, where data closely resembling benchmark questions leaks into the training data, instead of true reasoning ability. To investigate this claim rigorously, we commission Grade School Math 1000 (GSM1k). GSM1k is designed to mirror the style and complexity of the established GSM8k benchmark, the gold standard for measuring elementary mathematical reasoning. We ensure that the two benchmarks are comparable across important metrics such as human solve rates, number of steps in solution, answer magnitude, and more. When evaluating leading open- and closed-source LLMs on GSM1k, we observe accuracy drops of up to 13%, with several families of models (e.g., Phi and Mistral) showing evidence of systematic overfitting across almost all model sizes. At the same time, many models, especially those on the frontier, (e.g., Gemini/GPT/Claude) show minimal signs of overfitting. Further analysis suggests a positive relationship (Spearman's r^2=0.32) between a model's probability of generating an example from GSM8k and its performance gap between GSM8k and GSM1k, suggesting that many models may have partially memorized GSM8k.
A workflow for segmenting soil and plant X-ray CT images with deep learning in Googles Colaboratory
Mar 18, 2022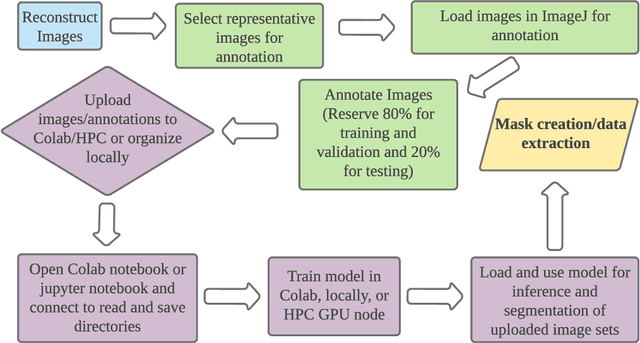
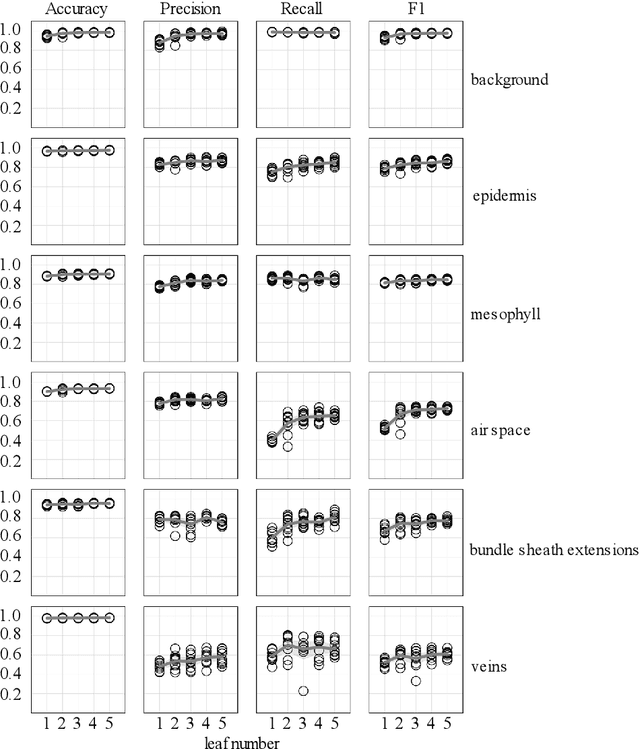
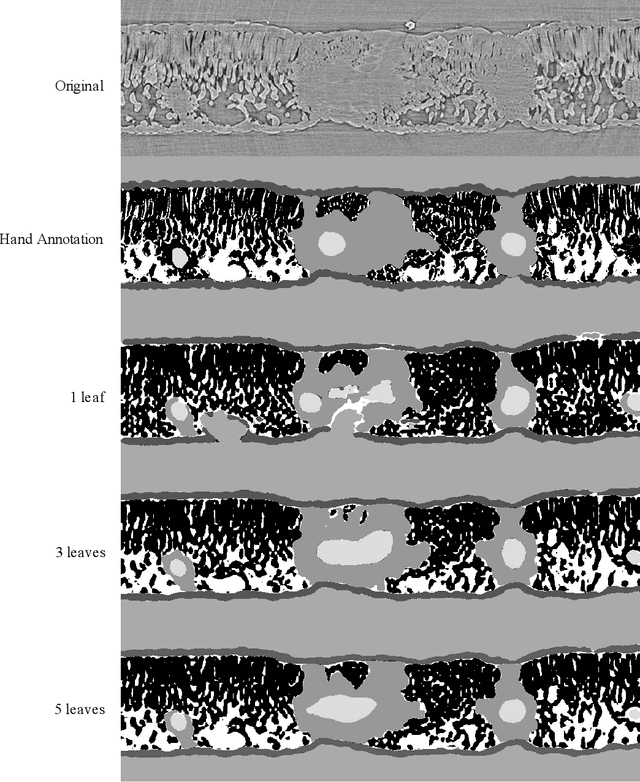
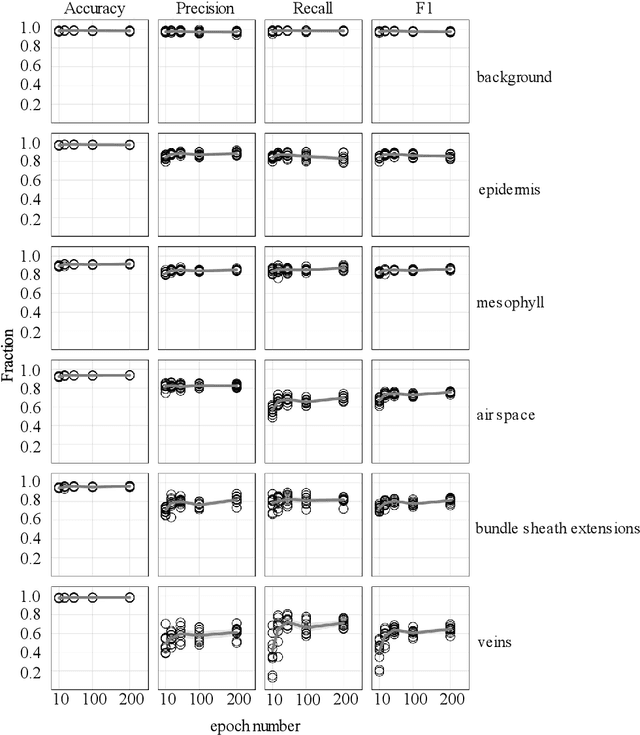
Abstract:X-ray micro-computed tomography (X-ray microCT) has enabled the characterization of the properties and processes that take place in plants and soils at the micron scale. Despite the widespread use of this advanced technique, major limitations in both hardware and software limit the speed and accuracy of image processing and data analysis. Recent advances in machine learning, specifically the application of convolutional neural networks to image analysis, have enabled rapid and accurate segmentation of image data. Yet, challenges remain in applying convolutional neural networks to the analysis of environmentally and agriculturally relevant images. Specifically, there is a disconnect between the computer scientists and engineers, who build these AI/ML tools, and the potential end users in agricultural research, who may be unsure of how to apply these tools in their work. Additionally, the computing resources required for training and applying deep learning models are unique, more common to computer gaming systems or graphics design work, than to traditional computational systems. To navigate these challenges, we developed a modular workflow for applying convolutional neural networks to X-ray microCT images, using low-cost resources in Googles Colaboratory web application. Here we present the results of the workflow, illustrating how parameters can be optimized to achieve best results using example scans from walnut leaves, almond flower buds, and a soil aggregate. We expect that this framework will accelerate the adoption and use of emerging deep learning techniques within the plant and soil sciences.
Simultaneously Predicting Multiple Plant Traits from Multiple Sensors via Deformable CNN Regression
Dec 06, 2021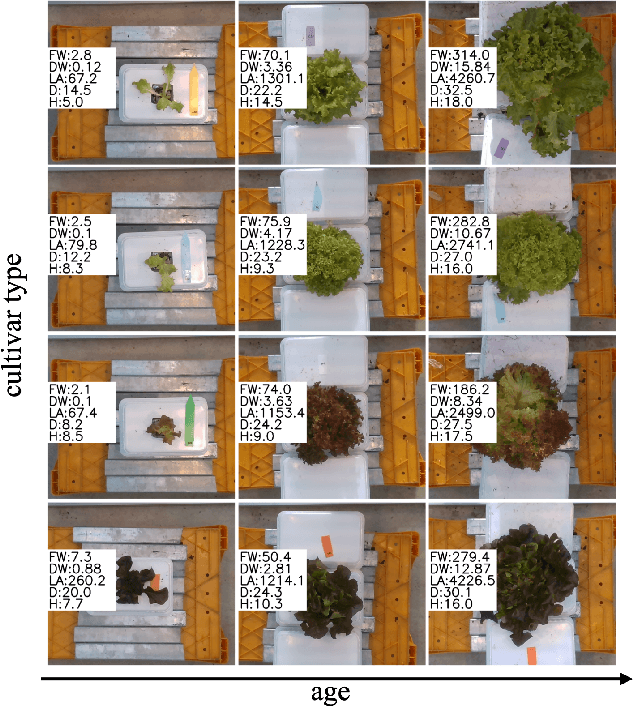
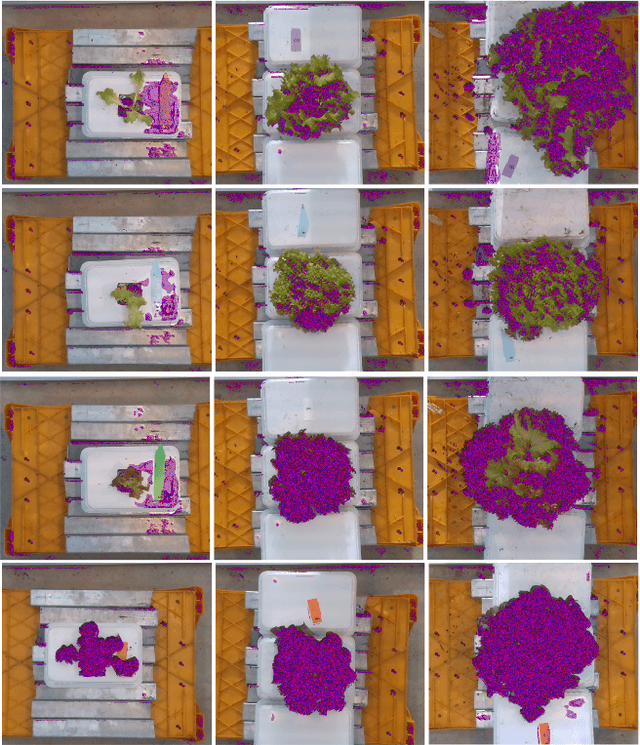
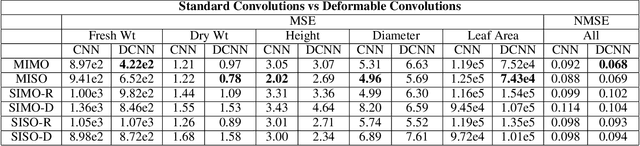
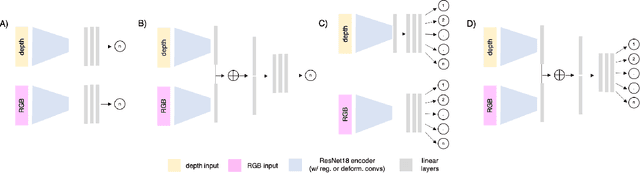

Abstract:Trait measurement is critical for the plant breeding and agricultural production pipeline. Typically, a suite of plant traits is measured using laborious manual measurements and then used to train and/or validate higher throughput trait estimation techniques. Here, we introduce a relatively simple convolutional neural network (CNN) model that accepts multiple sensor inputs and predicts multiple continuous trait outputs - i.e. a multi-input, multi-output CNN (MIMO-CNN). Further, we introduce deformable convolutional layers into this network architecture (MIMO-DCNN) to enable the model to adaptively adjust its receptive field, model complex variable geometric transformations in the data, and fine-tune the continuous trait outputs. We examine how the MIMO-CNN and MIMO-DCNN models perform on a multi-input (i.e. RGB and depth images), multi-trait output lettuce dataset from the 2021 Autonomous Greenhouse Challenge. Ablation studies were conducted to examine the effect of using single versus multiple inputs, and single versus multiple outputs. The MIMO-DCNN model resulted in a normalized mean squared error (NMSE) of 0.068 - a substantial improvement over the top 2021 leaderboard score of 0.081. Open-source code is provided.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge