Hyeong-Geol Shin
Department of Radiology, Johns Hopkins University School of Medicine, Baltimore, MD, USA, F.M. Kirby Research Center for Functional Brain Imaging, Kennedy Krieger Institute, Baltimore, MD, USA
$χ$-sepnet: Deep neural network for magnetic susceptibility source separation
Sep 24, 2024



Abstract:Magnetic susceptibility source separation ($\chi$-separation), an advanced quantitative susceptibility mapping (QSM) method, enables the separate estimation of para- and diamagnetic susceptibility source distributions in the brain. The method utilizes reversible transverse relaxation (R2'=R2*-R2) to complement frequency shift information for estimating susceptibility source concentrations, requiring time-consuming data acquisition for R2 in addition R2*. To address this challenge, we develop a new deep learning network, $\chi$-sepnet, and propose two deep learning-based susceptibility source separation pipelines, $\chi$-sepnet-R2' for inputs with multi-echo GRE and multi-echo spin-echo, and $\chi$-sepnet-R2* for input with multi-echo GRE only. $\chi$-sepnet is trained using multiple head orientation data that provide streaking artifact-free labels, generating high-quality $\chi$-separation maps. The evaluation of the pipelines encompasses both qualitative and quantitative assessments in healthy subjects, and visual inspection of lesion characteristics in multiple sclerosis patients. The susceptibility source-separated maps of the proposed pipelines delineate detailed brain structures with substantially reduced artifacts compared to those from conventional regularization-based reconstruction methods. In quantitative analysis, $\chi$-sepnet-R2' achieves the best outcomes followed by $\chi$-sepnet-R2*, outperforming the conventional methods. When the lesions of multiple sclerosis patients are assessed, both pipelines report identical lesion characteristics in most lesions ($\chi$para: 99.6% and $\chi$dia: 98.4% out of 250 lesions). The $\chi$-sepnet-R2* pipeline, which only requires multi-echo GRE data, has demonstrated its potential to offer broad clinical and scientific applications, although further evaluations for various diseases and pathological conditions are necessary.
A human brain atlas of chi-separation for normative iron and myelin distributions
Nov 08, 2023Abstract:Iron and myelin are primary susceptibility sources in the human brain. These substances are essential for healthy brain, and their abnormalities are often related to various neurological disorders. Recently, an advanced susceptibility mapping technique, which is referred to as chi-separation, has been proposed successfully disentangling paramagnetic iron from diamagnetic myelin, opening a new potential for generating iron map and myelin map in the brain. Utilizing this technique, this study constructs a normative chi-separation atlas from 106 healthy human brains. The resulting atlas provides detailed anatomical structures associated with the distributions of iron and myelin, clearly delineating subcortical nuclei and white matter fiber bundles. Additionally, susceptibility values in a number of regions of interest are reported along with age-dependent changes. This atlas may have direct applications such as localization of subcortical structures for deep brain stimulation or high-intensity focused ultrasound and also serve as a valuable resource for future research.
Coil2Coil: Self-supervised MR image denoising using phased-array coil images
Aug 16, 2022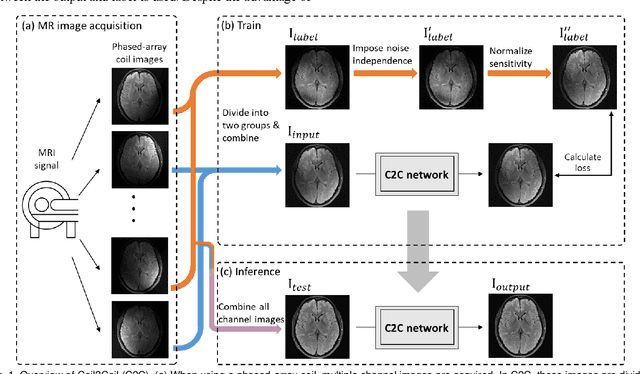
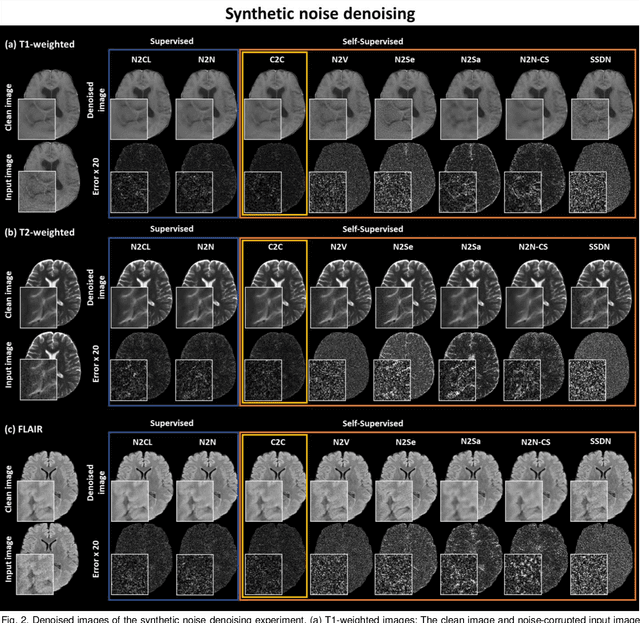
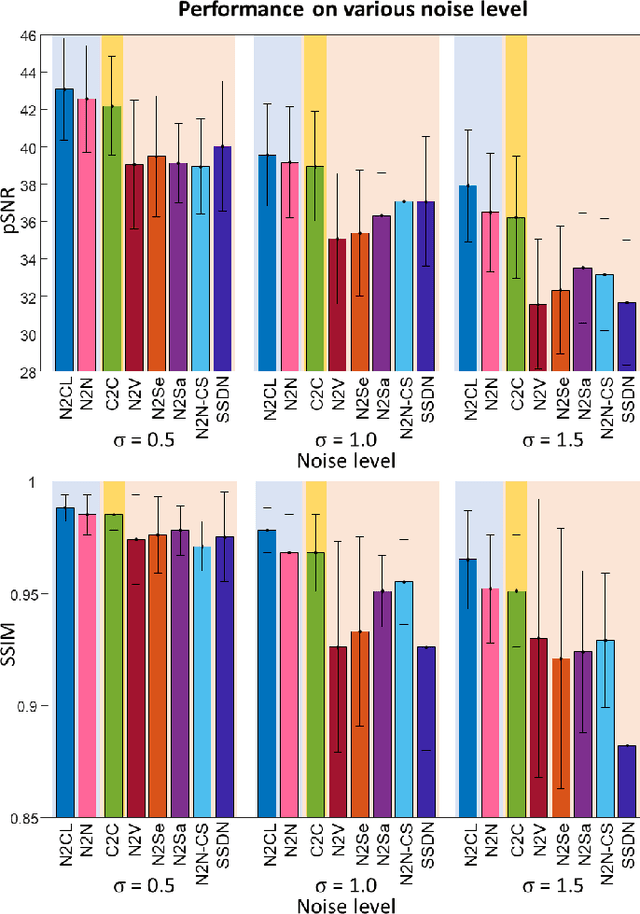
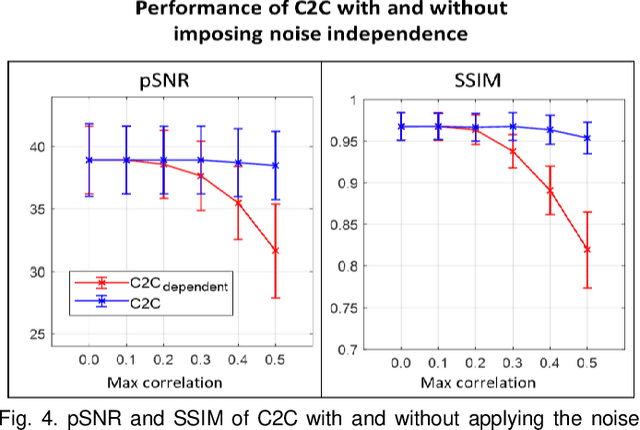
Abstract:Denoising of magnetic resonance images is beneficial in improving the quality of low signal-to-noise ratio images. Recently, denoising using deep neural networks has demonstrated promising results. Most of these networks, however, utilize supervised learning, which requires large training images of noise-corrupted and clean image pairs. Obtaining training images, particularly clean images, is expensive and time-consuming. Hence, methods such as Noise2Noise (N2N) that require only pairs of noise-corrupted images have been developed to reduce the burden of obtaining training datasets. In this study, we propose a new self-supervised denoising method, Coil2Coil (C2C), that does not require the acquisition of clean images or paired noise-corrupted images for training. Instead, the method utilizes multichannel data from phased-array coils to generate training images. First, it divides and combines multichannel coil images into two images, one for input and the other for label. Then, they are processed to impose noise independence and sensitivity normalization such that they can be used for the training images of N2N. For inference, the method inputs a coil-combined image (e.g., DICOM image), enabling a wide application of the method. When evaluated using synthetic noise-added images, C2C shows the best performance against several self-supervised methods, reporting comparable outcomes to supervised methods. When testing the DICOM images, C2C successfully denoised real noise without showing structure-dependent residuals in the error maps. Because of the significant advantage of not requiring additional scans for clean or paired images, the method can be easily utilized for various clinical applications.
BUDA-SAGE with self-supervised denoising enables fast, distortion-free, high-resolution T2, T2*, para- and dia-magnetic susceptibility mapping
Sep 09, 2021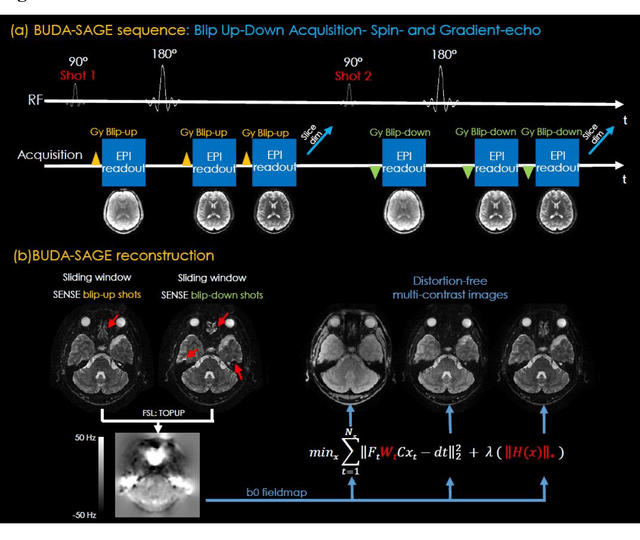
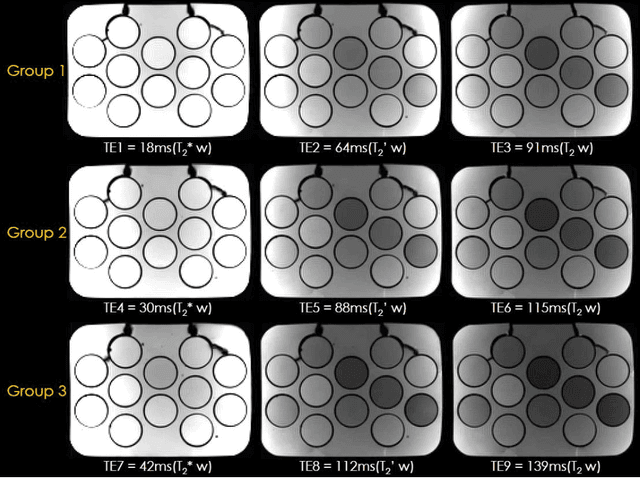
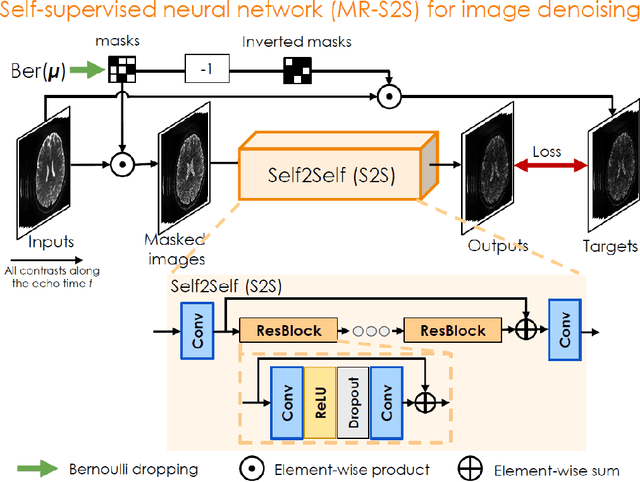

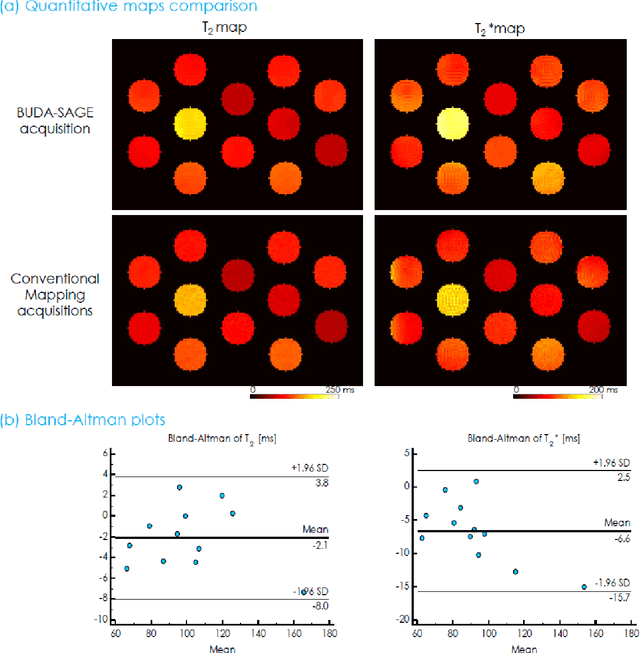
Abstract:To rapidly obtain high resolution T2, T2* and quantitative susceptibility mapping (QSM) source separation maps with whole-brain coverage and high geometric fidelity. We propose Blip Up-Down Acquisition for Spin And Gradient Echo imaging (BUDA-SAGE), an efficient echo-planar imaging (EPI) sequence for quantitative mapping. The acquisition includes multiple T2*-, T2'- and T2-weighted contrasts. We alternate the phase-encoding polarities across the interleaved shots in this multi-shot navigator-free acquisition. A field map estimated from interim reconstructions was incorporated into the joint multi-shot EPI reconstruction with a structured low rank constraint to eliminate geometric distortion. A self-supervised MR-Self2Self (MR-S2S) neural network (NN) was utilized to perform denoising after BUDA reconstruction to boost SNR. Employing Slider encoding allowed us to reach 1 mm isotropic resolution by performing super-resolution reconstruction on BUDA-SAGE volumes acquired with 2 mm slice thickness. Quantitative T2 and T2* maps were obtained using Bloch dictionary matching on the reconstructed echoes. QSM was estimated using nonlinear dipole inversion (NDI) on the gradient echoes. Starting from the estimated R2 and R2* maps, R2' information was derived and used in source separation QSM reconstruction, which provided additional para- and dia-magnetic susceptibility maps. In vivo results demonstrate the ability of BUDA-SAGE to provide whole-brain, distortion-free, high-resolution multi-contrast images and quantitative T2 and T2* maps, as well as yielding para- and dia-magnetic susceptibility maps. Derived quantitative maps showed comparable values to conventional mapping methods in phantom and in vivo measurements. BUDA-SAGE acquisition with self-supervised denoising and Slider encoding enabled rapid, distortion-free, whole-brain T2, T2* mapping at 1 mm3 isotropic resolution in 90 seconds.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge