Huahong Zhang
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Unsupervised Cross-Modality Domain Adaptation for Segmenting Vestibular Schwannoma and Cochlea with Data Augmentation and Model Ensemble
Sep 24, 2021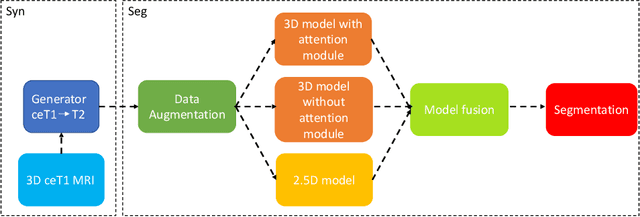
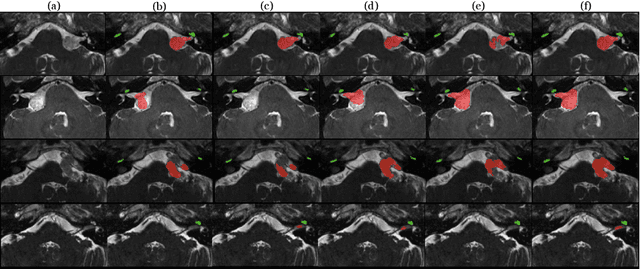
Abstract:Magnetic resonance images (MRIs) are widely used to quantify vestibular schwannoma and the cochlea. Recently, deep learning methods have shown state-of-the-art performance for segmenting these structures. However, training segmentation models may require manual labels in target domain, which is expensive and time-consuming. To overcome this problem, domain adaptation is an effective way to leverage information from source domain to obtain accurate segmentations without requiring manual labels in target domain. In this paper, we propose an unsupervised learning framework to segment the VS and cochlea. Our framework leverages information from contrast-enhanced T1-weighted (ceT1-w) MRIs and its labels, and produces segmentations for T2-weighted MRIs without any labels in the target domain. We first applied a generator to achieve image-to-image translation. Next, we ensemble outputs from an ensemble of different models to obtain final segmentations. To cope with MRIs from different sites/scanners, we applied various 'online' augmentations during training to better capture the geometric variability and the variability in image appearance and quality. Our method is easy to build and produces promising segmentations, with a mean Dice score of 0.7930 and 0.7432 for VS and cochlea respectively in the validation set.
Multiple Sclerosis Lesion Segmentation -- A Survey of Supervised CNN-Based Methods
Dec 26, 2020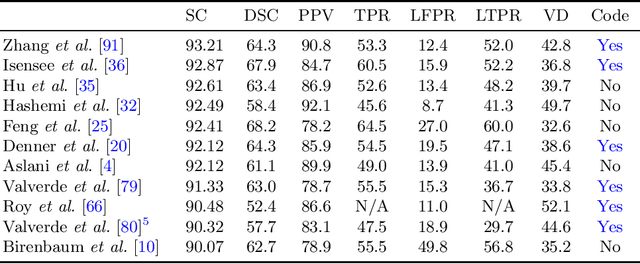
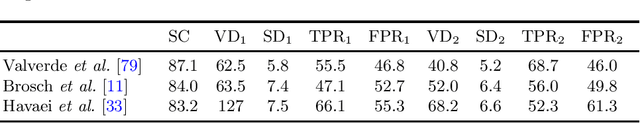
Abstract:Lesion segmentation is a core task for quantitative analysis of MRI scans of Multiple Sclerosis patients. The recent success of deep learning techniques in a variety of medical image analysis applications has renewed community interest in this challenging problem and led to a burst of activity for new algorithm development. In this survey, we investigate the supervised CNN-based methods for MS lesion segmentation. We decouple these reviewed works into their algorithmic components and discuss each separately. For methods that provide evaluations on public benchmark datasets, we report comparisons between their results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge