Houliang Zhou
SDE-Driven Spatio-Temporal Hypergraph Neural Networks for Irregular Longitudinal fMRI Connectome Modeling in Alzheimer's Disease
Mar 20, 2026Abstract:Longitudinal neuroimaging is essential for modeling disease progression in Alzheimer's disease (AD), yet irregular sampling and missing visits pose substantial challenges for learning reliable temporal representations. To address this challenge, we propose SDE-HGNN, a stochastic differential equation (SDE)-driven spatio-temporal hypergraph neural network for irregular longitudinal fMRI connectome modeling. The framework first employs an SDE-based reconstruction module to recover continuous latent trajectories from irregular observations. Based on these reconstructed representations, dynamic hypergraphs are constructed to capture higher-order interactions among brain regions over time. To further model temporal evolution, hypergraph convolution parameters evolve through SDE-controlled recurrent dynamics conditioned on inter-scan intervals, enabling disease-stage-adaptive connectivity modeling. We also incorporate a sparsity-based importance learning mechanism to identify salient brain regions and discriminative connectivity patterns. Extensive experiments on the OASIS-3 and ADNI cohorts demonstrate consistent improvements over state-of-the-art graph and hypergraph baselines in AD progression prediction. The source code is available at https://anonymous.4open.science/r/SDE-HGNN-017F.
Adaptive Clinical-Aware Latent Diffusion for Multimodal Brain Image Generation and Missing Modality Imputation
Mar 10, 2026Abstract:Multimodal neuroimaging provides complementary insights for Alzheimer's disease diagnosis, yet clinical datasets frequently suffer from missing modalities. We propose ACADiff, a framework that synthesizes missing brain imaging modalities through adaptive clinical-aware diffusion. ACADiff learns mappings between incomplete multimodal observations and target modalities by progressively denoising latent representations while attending to available imaging data and clinical metadata. The framework employs adaptive fusion that dynamically reconfigures based on input availability, coupled with semantic clinical guidance via GPT-4o-encoded prompts. Three specialized generators enable bidirectional synthesis among sMRI, FDG-PET, and AV45-PET. Evaluated on ADNI subjects, ACADiff achieves superior generation quality and maintains robust diagnostic performance even under extreme 80\% missing scenarios, outperforming all existing baselines. To promote reproducibility, code is available at https://github.com/rongzhou7/ACADiff
Interpretable Graph Convolutional Network of Multi-Modality Brain Imaging for Alzheimer's Disease Diagnosis
Apr 27, 2022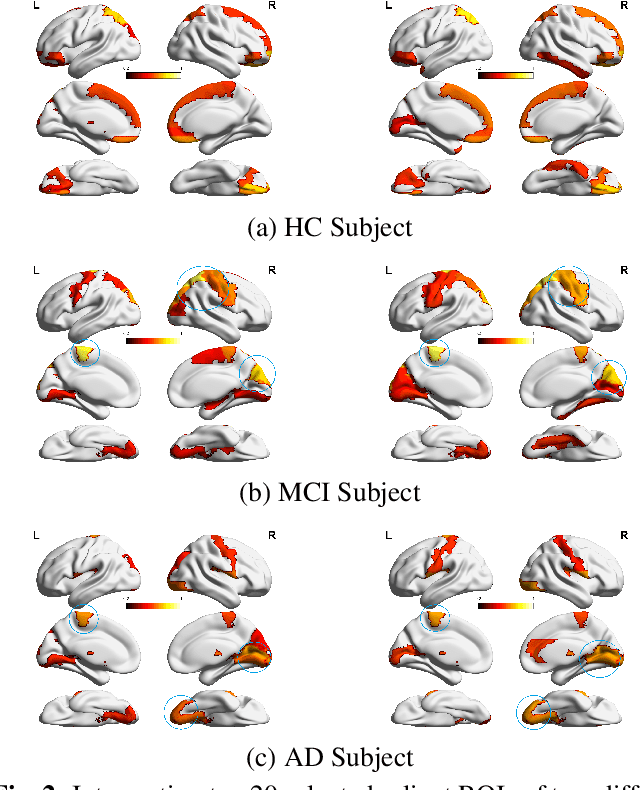
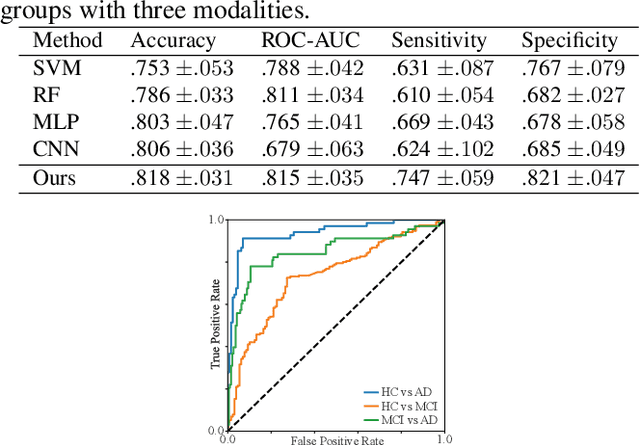
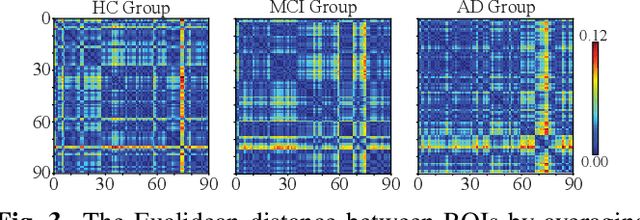
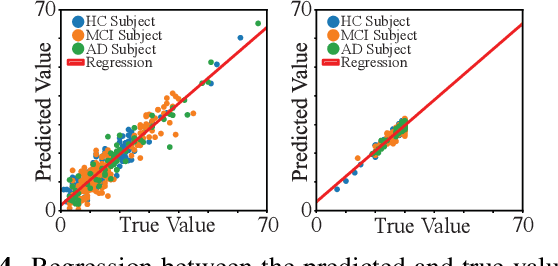
Abstract:Identification of brain regions related to the specific neurological disorders are of great importance for biomarker and diagnostic studies. In this paper, we propose an interpretable Graph Convolutional Network (GCN) framework for the identification and classification of Alzheimer's disease (AD) using multi-modality brain imaging data. Specifically, we extended the Gradient Class Activation Mapping (Grad-CAM) technique to quantify the most discriminative features identified by GCN from brain connectivity patterns. We then utilized them to find signature regions of interest (ROIs) by detecting the difference of features between regions in healthy control (HC), mild cognitive impairment (MCI), and AD groups. We conducted the experiments on the ADNI database with imaging data from three modalities, including VBM-MRI, FDG-PET, and AV45-PET, and showed that the ROI features learned by our method were effective for enhancing the performances of both clinical score prediction and disease status identification. It also successfully identified biomarkers associated with AD and MCI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge