Himashi Peiris
M2Diff: Multi-Modality Multi-Task Enhanced Diffusion Model for MRI-Guided Low-Dose PET Enhancement
Mar 10, 2026Abstract:Positron emission tomography (PET) scans expose patients to radiation, which can be mitigated by reducing the dose, albeit at the cost of diminished quality. This makes low-dose (LD) PET recovery an active research area. Previous studies have focused on standard-dose (SD) PET recovery from LD PET scans and/or multi-modal scans, e.g., PET/CT or PET/MRI, using deep learning. While these studies incorporate multi-modal information through conditioning in a single-task model, such approaches may limit the capacity to extract modality-specific features, potentially leading to early feature dilution. Although recent studies have begun incorporating pathology-rich data, challenges remain in effectively leveraging multi-modality inputs for reconstructing diverse features, particularly in heterogeneous patient populations. To address these limitations, we introduce a multi-modality multi-task diffusion model (M2Diff) that processes MRI and LD PET scans separately to learn modality-specific features and fuse them via hierarchical feature fusion to reconstruct SD PET. This design enables effective integration of complementary structural and functional information, leading to improved reconstruction fidelity. We have validated the effectiveness of our model on both healthy and Alzheimer's disease brain datasets. The M2Diff achieves superior qualitative and quantitative performance on both datasets.
Segment Any Tumour: An Uncertainty-Aware Vision Foundation Model for Whole-Body Analysis
Nov 11, 2025Abstract:Prompt-driven vision foundation models, such as the Segment Anything Model, have recently demonstrated remarkable adaptability in computer vision. However, their direct application to medical imaging remains challenging due to heterogeneous tissue structures, imaging artefacts, and low-contrast boundaries, particularly in tumours and cancer primaries leading to suboptimal segmentation in ambiguous or overlapping lesion regions. Here, we present Segment Any Tumour 3D (SAT3D), a lightweight volumetric foundation model designed to enable robust and generalisable tumour segmentation across diverse medical imaging modalities. SAT3D integrates a shifted-window vision transformer for hierarchical volumetric representation with an uncertainty-aware training pipeline that explicitly incorporates uncertainty estimates as prompts to guide reliable boundary prediction in low-contrast regions. Adversarial learning further enhances model performance for the ambiguous pathological regions. We benchmark SAT3D against three recent vision foundation models and nnUNet across 11 publicly available datasets, encompassing 3,884 tumour and cancer cases for training and 694 cases for in-distribution evaluation. Trained on 17,075 3D volume-mask pairs across multiple modalities and cancer primaries, SAT3D demonstrates strong generalisation and robustness. To facilitate practical use and clinical translation, we developed a 3D Slicer plugin that enables interactive, prompt-driven segmentation and visualisation using the trained SAT3D model. Extensive experiments highlight its effectiveness in improving segmentation accuracy under challenging and out-of-distribution scenarios, underscoring its potential as a scalable foundation model for medical image analysis.
Bilateral Hippocampi Segmentation in Low Field MRIs Using Mutual Feature Learning via Dual-Views
Oct 23, 2024



Abstract:Accurate hippocampus segmentation in brain MRI is critical for studying cognitive and memory functions and diagnosing neurodevelopmental disorders. While high-field MRIs provide detailed imaging, low-field MRIs are more accessible and cost-effective, which eliminates the need for sedation in children, though they often suffer from lower image quality. In this paper, we present a novel deep-learning approach for the automatic segmentation of bilateral hippocampi in low-field MRIs. Extending recent advancements in infant brain segmentation to underserved communities through the use of low-field MRIs ensures broader access to essential diagnostic tools, thereby supporting better healthcare outcomes for all children. Inspired by our previous work, Co-BioNet, the proposed model employs a dual-view structure to enable mutual feature learning via high-frequency masking, enhancing segmentation accuracy by leveraging complementary information from different perspectives. Extensive experiments demonstrate that our method provides reliable segmentation outcomes for hippocampal analysis in low-resource settings. The code is publicly available at: https://github.com/himashi92/LoFiHippSeg.
Motion-Informed Deep Learning for Brain MR Image Reconstruction Framework
May 28, 2024



Abstract:Motion artifacts in Magnetic Resonance Imaging (MRI) are one of the frequently occurring artifacts due to patient movements during scanning. Motion is estimated to be present in approximately 30% of clinical MRI scans; however, motion has not been explicitly modeled within deep learning image reconstruction models. Deep learning (DL) algorithms have been demonstrated to be effective for both the image reconstruction task and the motion correction task, but the two tasks are considered separately. The image reconstruction task involves removing undersampling artifacts such as noise and aliasing artifacts, whereas motion correction involves removing artifacts including blurring, ghosting, and ringing. In this work, we propose a novel method to simultaneously accelerate imaging and correct motion. This is achieved by integrating a motion module into the deep learning-based MRI reconstruction process, enabling real-time detection and correction of motion. We model motion as a tightly integrated auxiliary layer in the deep learning model during training, making the deep learning model 'motion-informed'. During inference, image reconstruction is performed from undersampled raw k-space data using a trained motion-informed DL model. Experimental results demonstrate that the proposed motion-informed deep learning image reconstruction network outperformed the conventional image reconstruction network for motion-degraded MRI datasets.
Perivascular space Identification Nnunet for Generalised Usage (PINGU)
May 17, 2024Abstract:Perivascular spaces(PVSs) form a central component of the brain\'s waste clearance system, the glymphatic system. These structures are visible on MRI images, and their morphology is associated with aging and neurological disease. Manual quantification of PVS is time consuming and subjective. Numerous deep learning methods for PVS segmentation have been developed, however the majority have been developed and evaluated on homogenous datasets and high resolution scans, perhaps limiting their applicability for the wide range of image qualities acquired in clinic and research. In this work we train a nnUNet, a top-performing biomedical image segmentation algorithm, on a heterogenous training sample of manually segmented MRI images of a range of different qualities and resolutions from 6 different datasets. These are compared to publicly available deep learning methods for 3D segmentation of PVS. The resulting model, PINGU (Perivascular space Identification Nnunet for Generalised Usage), achieved voxel and cluster level dice scores of 0.50(SD=0.15), 0.63(0.17) in the white matter(WM), and 0.54(0.11), 0.66(0.17) in the basal ganglia(BG). Performance on data from unseen sites was substantially lower for both PINGU(0.20-0.38(WM, voxel), 0.29-0.58(WM, cluster), 0.22-0.36(BG, voxel), 0.46-0.60(BG, cluster)) and the publicly available algorithms(0.18-0.30(WM, voxel), 0.29-0.38(WM cluster), 0.10-0.20(BG, voxel), 0.15-0.37(BG, cluster)), but PINGU strongly outperformed the publicly available algorithms, particularly in the BG. Finally, training PINGU on manual segmentations from a single site with homogenous scan properties gave marginally lower performances on internal cross-validation, but in some cases gave higher performance on external validation. PINGU stands out as broad-use PVS segmentation tool, with particular strength in the BG, an area of PVS related to vascular disease and pathology.
Hybrid Window Attention Based Transformer Architecture for Brain Tumor Segmentation
Sep 16, 2022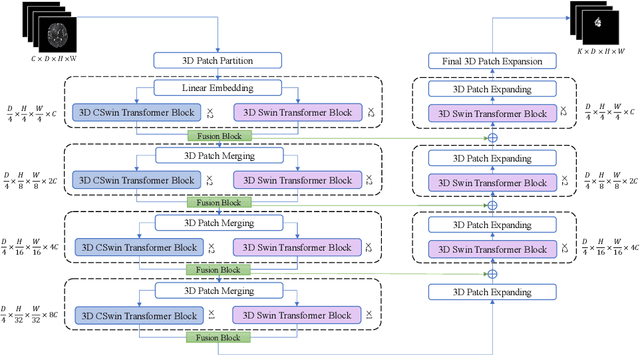
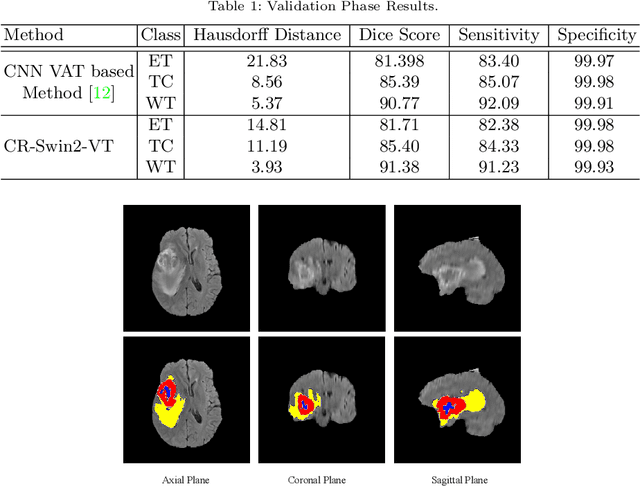
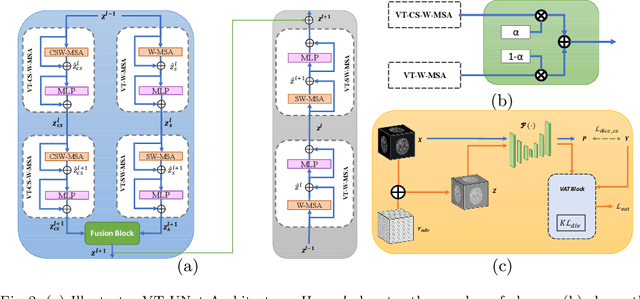
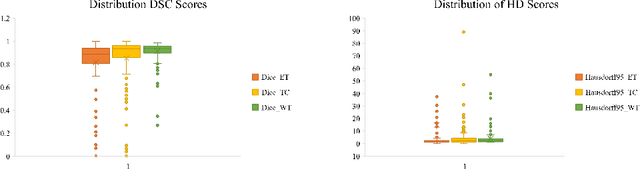
Abstract:As intensities of MRI volumes are inconsistent across institutes, it is essential to extract universal features of multi-modal MRIs to precisely segment brain tumors. In this concept, we propose a volumetric vision transformer that follows two windowing strategies in attention for extracting fine features and local distributional smoothness (LDS) during model training inspired by virtual adversarial training (VAT) to make the model robust. We trained and evaluated network architecture on the FeTS Challenge 2022 dataset. Our performance on the online validation dataset is as follows: Dice Similarity Score of 81.71%, 91.38% and 85.40%; Hausdorff Distance (95%) of 14.81 mm, 3.93 mm, 11.18 mm for the enhancing tumor, whole tumor, and tumor core, respectively. Overall, the experimental results verify our method's effectiveness by yielding better performance in segmentation accuracy for each tumor sub-region. Our code implementation is publicly available : https://github.com/himashi92/vizviva_fets_2022
Reciprocal Adversarial Learning for Brain Tumor Segmentation: A Solution to BraTS Challenge 2021 Segmentation Task
Jan 11, 2022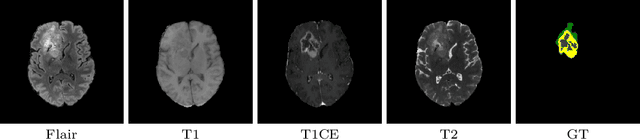
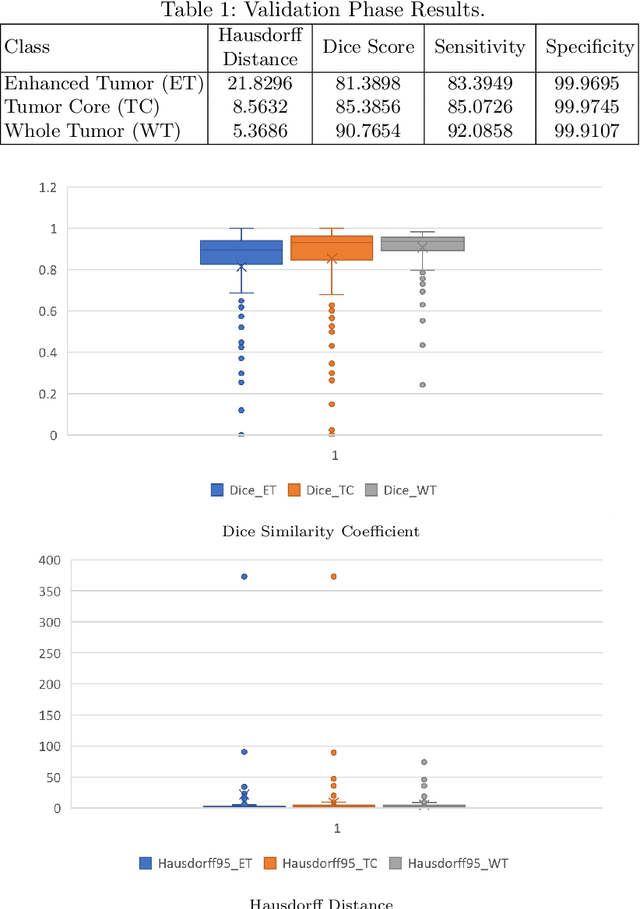
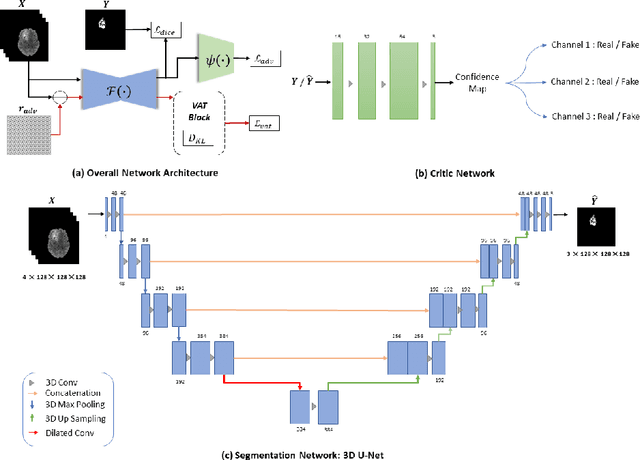
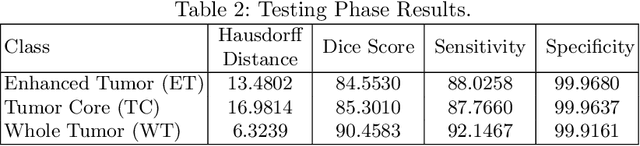
Abstract:This paper proposes an adversarial learning based training approach for brain tumor segmentation task. In this concept, the 3D segmentation network learns from dual reciprocal adversarial learning approaches. To enhance the generalization across the segmentation predictions and to make the segmentation network robust, we adhere to the Virtual Adversarial Training approach by generating more adversarial examples via adding some noise on original patient data. By incorporating a critic that acts as a quantitative subjective referee, the segmentation network learns from the uncertainty information associated with segmentation results. We trained and evaluated network architecture on the RSNA-ASNR-MICCAI BraTS 2021 dataset. Our performance on the online validation dataset is as follows: Dice Similarity Score of 81.38%, 90.77% and 85.39%; Hausdorff Distance (95\%) of 21.83 mm, 5.37 mm, 8.56 mm for the enhancing tumor, whole tumor and tumor core, respectively. Similarly, our approach achieved a Dice Similarity Score of 84.55%, 90.46% and 85.30%, as well as Hausdorff Distance (95\%) of 13.48 mm, 6.32 mm and 16.98 mm on the final test dataset. Overall, our proposed approach yielded better performance in segmentation accuracy for each tumor sub-region. Our code implementation is publicly available at https://github.com/himashi92/vizviva_brats_2021
A Volumetric Transformer for Accurate 3D Tumor Segmentation
Nov 26, 2021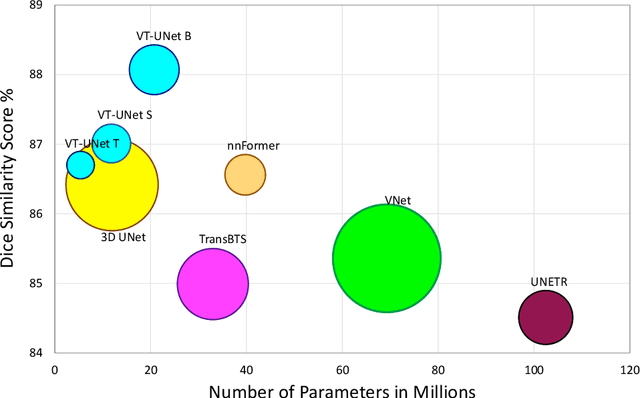
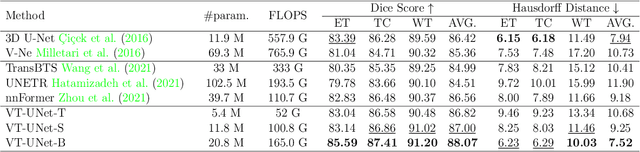
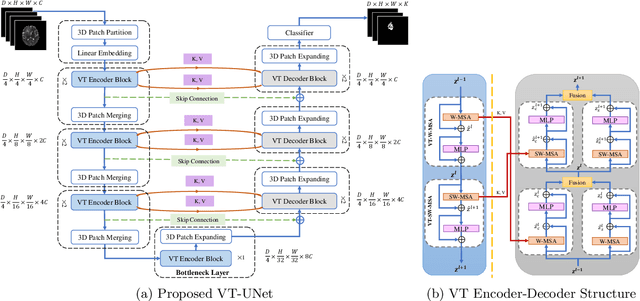
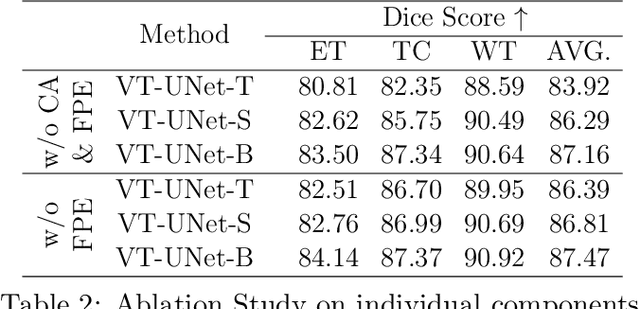
Abstract:This paper presents a Transformer architecture for volumetric medical image segmentation. Designing a computationally efficient Transformer architecture for volumetric segmentation is a challenging task. It requires keeping a complex balance in encoding local and global spatial cues, and preserving information along all axes of the volumetric data. The proposed volumetric Transformer has a U-shaped encoder-decoder design that processes the input voxels in their entirety. Our encoder has two consecutive self-attention layers to simultaneously encode local and global cues, and our decoder has novel parallel shifted window based self and cross attention blocks to capture fine details for boundary refinement by subsuming Fourier position encoding. Our proposed design choices result in a computationally efficient architecture, which demonstrates promising results on Brain Tumor Segmentation (BraTS) 2021, and Medical Segmentation Decathlon (Pancreas and Liver) datasets for tumor segmentation. We further show that the representations learned by our model transfer better across-datasets and are robust against data corruptions. \href{https://github.com/himashi92/VT-UNet}{Our code implementation is publicly available}.
Duo-SegNet: Adversarial Dual-Views for Semi-Supervised Medical Image Segmentation
Aug 25, 2021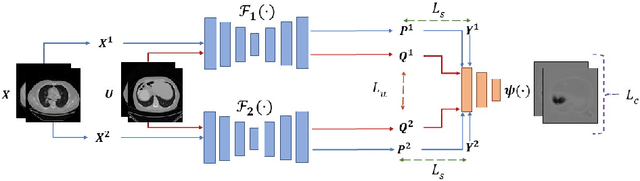
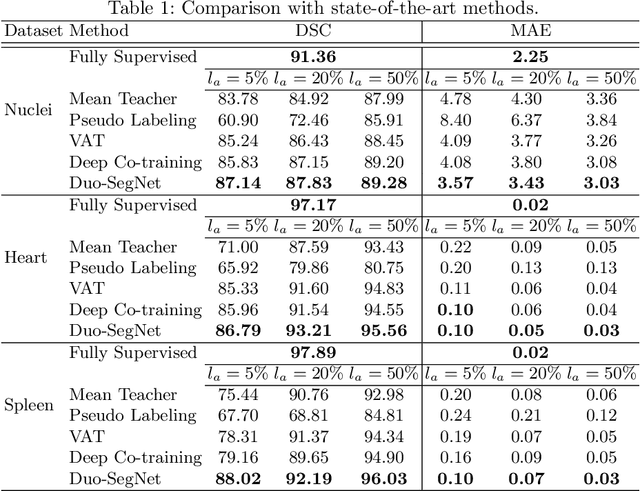
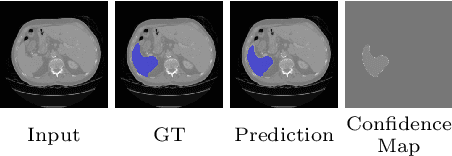
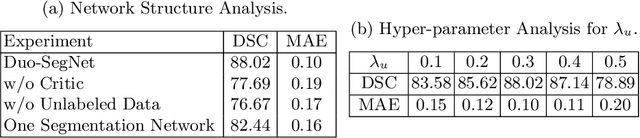
Abstract:Segmentation of images is a long-standing challenge in medical AI. This is mainly due to the fact that training a neural network to perform image segmentation requires a significant number of pixel-level annotated data, which is often unavailable. To address this issue, we propose a semi-supervised image segmentation technique based on the concept of multi-view learning. In contrast to the previous art, we introduce an adversarial form of dual-view training and employ a critic to formulate the learning problem in multi-view training as a min-max problem. Thorough quantitative and qualitative evaluations on several datasets indicate that our proposed method outperforms state-of-the-art medical image segmentation algorithms consistently and comfortably. The code is publicly available at https://github.com/himashi92/Duo-SegNet
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge