Hideki Nakayama
Graph-based Heuristic Search for Module Selection Procedure in Neural Module Network
Sep 30, 2020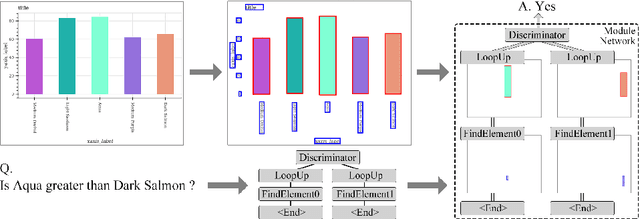

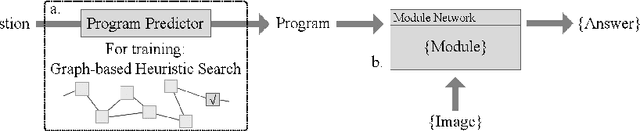
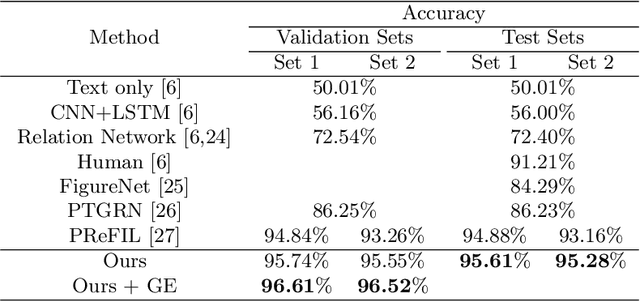
Abstract:Neural Module Network (NMN) is a machine learning model for solving the visual question answering tasks. NMN uses programs to encode modules' structures, and its modularized architecture enables it to solve logical problems more reasonably. However, because of the non-differentiable procedure of module selection, NMN is hard to be trained end-to-end. To overcome this problem, existing work either included ground-truth program into training data or applied reinforcement learning to explore the program. However, both of these methods still have weaknesses. In consideration of this, we proposed a new learning framework for NMN. Graph-based Heuristic Search is the algorithm we proposed to discover the optimal program through a heuristic search on the data structure named Program Graph. Our experiments on FigureQA and CLEVR dataset show that our methods can realize the training of NMN without ground-truth programs and achieve superior efficiency over existing reinforcement learning methods in program exploration.
MADGAN: unsupervised Medical Anomaly Detection GAN using multiple adjacent brain MRI slice reconstruction
Jul 24, 2020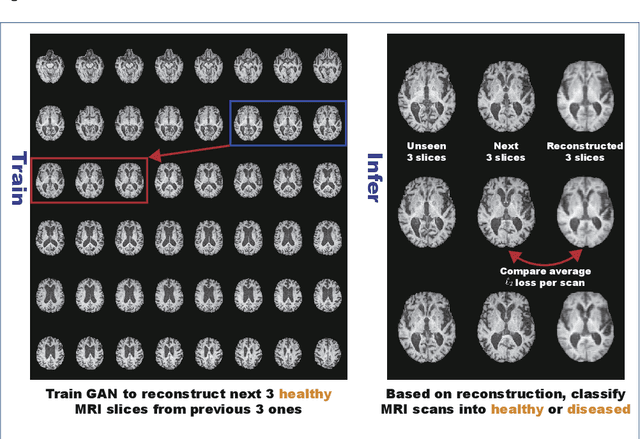
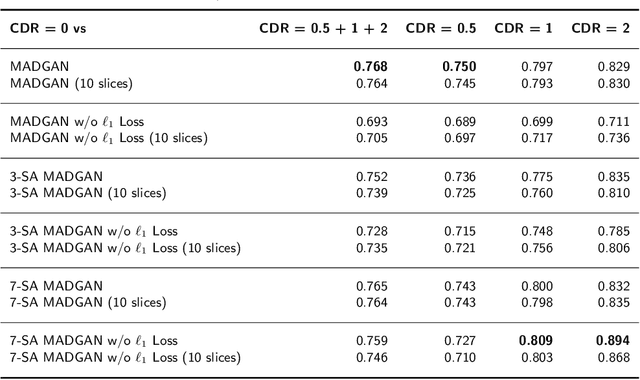
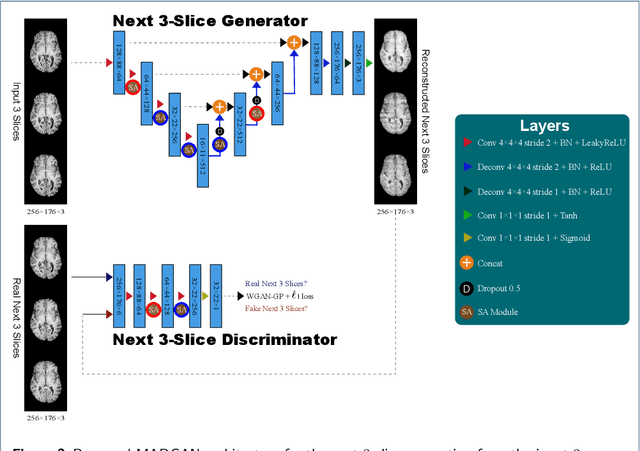
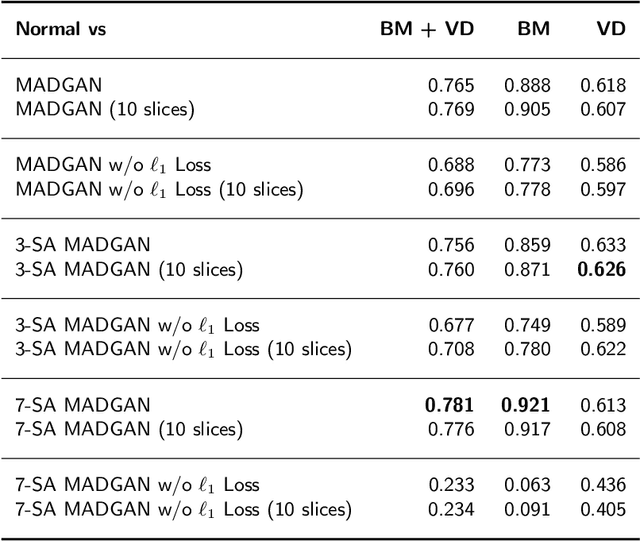
Abstract:Unsupervised learning can discover various unseen diseases, relying on large-scale unannotated medical images of healthy subjects. Towards this, unsupervised methods reconstruct a 2D/3D single medical image to detect outliers either in the learned feature space or from high reconstruction loss. However, without considering continuity between multiple adjacent slices, they cannot directly discriminate diseases composed of the accumulation of subtle anatomical anomalies, such as Alzheimer's Disease (AD). Moreover, no study has shown how unsupervised anomaly detection is associated with either disease stages, various (i.e., more than two types of) diseases, or multi-sequence Magnetic Resonance Imaging (MRI) scans. Therefore, we propose unsupervised Medical Anomaly Detection Generative Adversarial Network (MADGAN), a novel two-step method using GAN-based multiple adjacent brain MRI slice reconstruction to detect various diseases at different stages on multi-sequence structural MRI: (Reconstruction) Wasserstein loss with Gradient Penalty + 100 L1 loss-trained on 3 healthy brain axial MRI slices to reconstruct the next 3 ones-reconstructs unseen healthy/abnormal scans; (Diagnosis) Average L2 loss per scan discriminates them, comparing the ground truth/reconstructed slices. For training, we use 1,133 healthy T1-weighted (T1) and 135 healthy contrast-enhanced T1 (T1c) brain MRI scans. Our Self-Attention MADGAN can detect AD on T1 scans at a very early stage, Mild Cognitive Impairment (MCI), with Area Under the Curve (AUC) 0.727, and AD at a late stage with AUC 0.894, while detecting brain metastases on T1c scans with AUC 0.921.
Meta Approach to Data Augmentation Optimization
Jun 14, 2020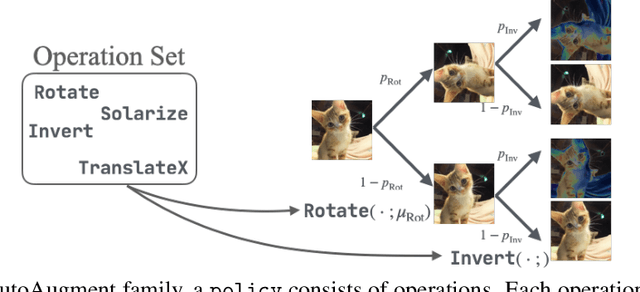
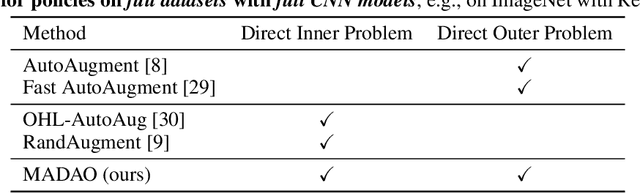

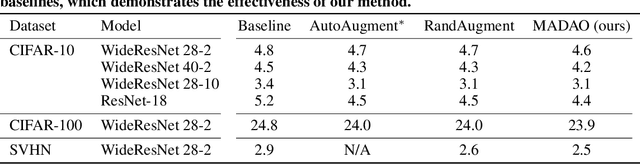
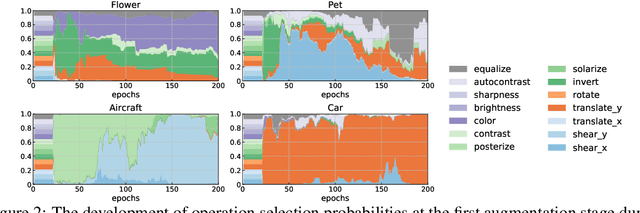
Abstract:Data augmentation policies drastically improve the performance of image recognition tasks, especially when the policies are optimized for the target data and tasks. In this paper, we propose to optimize image recognition models and data augmentation policies simultaneously to improve the performance using gradient descent. Unlike prior methods, our approach avoids using proxy tasks or reducing search space, and can directly improve the validation performance. Our method achieves efficient and scalable training by approximating the gradient of policies by implicit gradient with Neumann series approximation. We demonstrate that our approach can improve the performance of various image classification tasks, including ImageNet classification and fine-grained recognition, without using dataset-specific hyperparameter tuning.
Single Model Ensemble using Pseudo-Tags and Distinct Vectors
May 02, 2020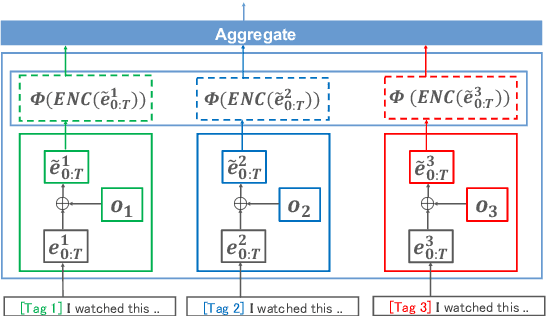
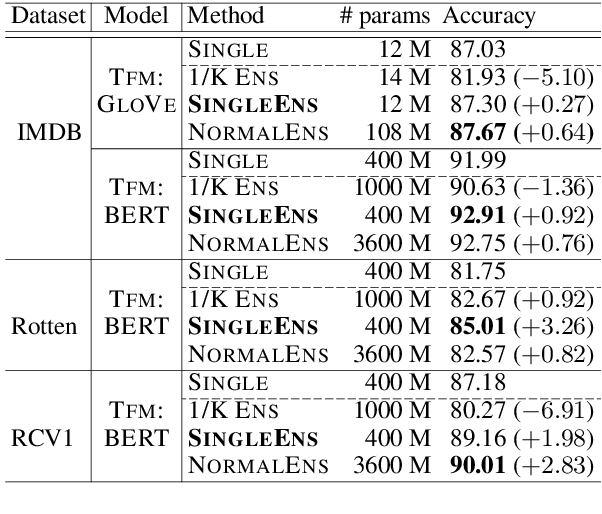
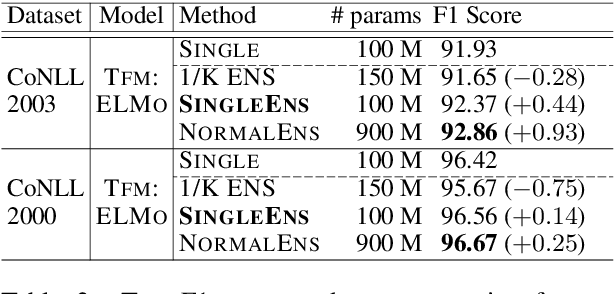
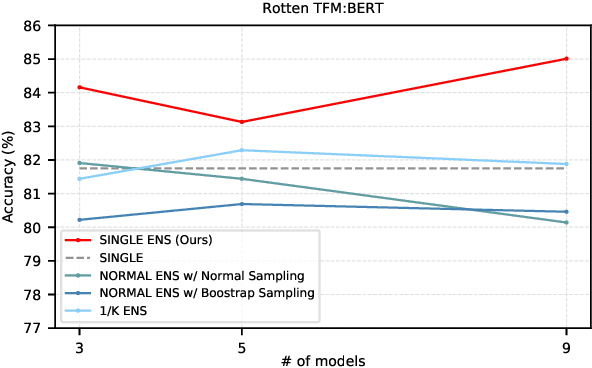
Abstract:Model ensemble techniques often increase task performance in neural networks; however, they require increased time, memory, and management effort. In this study, we propose a novel method that replicates the effects of a model ensemble with a single model. Our approach creates K-virtual models within a single parameter space using K-distinct pseudo-tags and K-distinct vectors. Experiments on text classification and sequence labeling tasks on several datasets demonstrate that our method emulates or outperforms a traditional model ensemble with 1/K-times fewer parameters.
Bridging the gap between AI and Healthcare sides: towards developing clinically relevant AI-powered diagnosis systems
Jan 12, 2020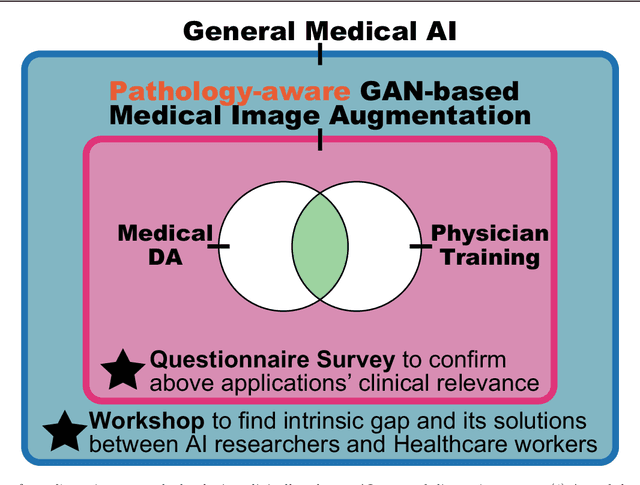

Abstract:This work aims to identify/bridge the gap between Artificial Intelligence (AI) and Healthcare sides in Japan towards developing medical AI fitting into a clinical environment in five years. Moreover, we attempt to confirm the clinical relevance for diagnosis of our research-proven pathology-aware Generative Adversarial Network (GAN)-based medical image augmentation: a data wrangling and information conversion technique to address data paucity. We hold a clinically valuable AI-envisioning workshop among 2 Medical Imaging experts, 2 physicians, and 3 Healthcare/Informatics generalists. A qualitative/quantitative questionnaire survey for 3 project-related physicians and 6 project non-related radiologists evaluates the GAN projects in terms of Data Augmentation (DA) and physician training. The workshop reveals the intrinsic gap between AI/Healthcare sides and its preliminary solutions on Why (i.e., clinical significance/interpretation) and How (i.e., data acquisition, commercial deployment, and safety/feeling safe). The survey confirms our pathology-aware GANs' clinical relevance as a clinical decision support system and non-expert physician training tool. Radiologists generally have high expectations for AI-based diagnosis as a reliable second opinion and abnormal candidate detection, instead of replacing them. Our findings would play a key role in connecting inter-disciplinary research and clinical applications, not limited to the Japanese medical context and pathology-aware GANs. We find that better DA and expert physician training would require atypical image generation via further GAN-based extrapolation.
Empirical Study of Easy and Hard Examples in CNN Training
Nov 25, 2019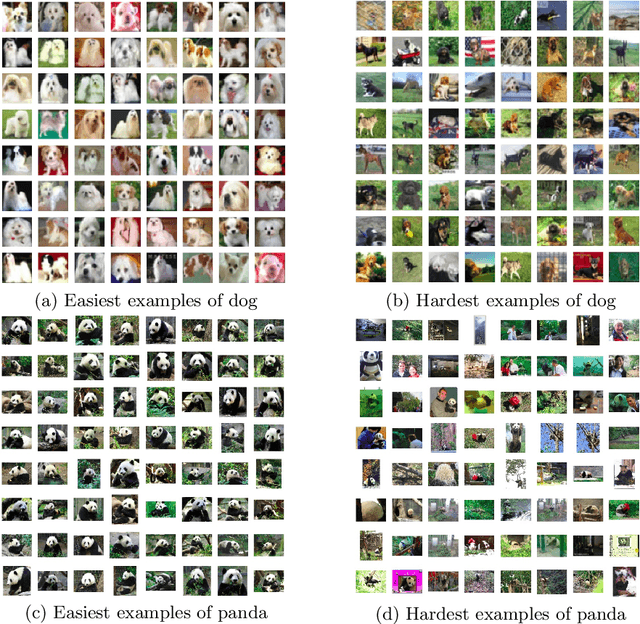
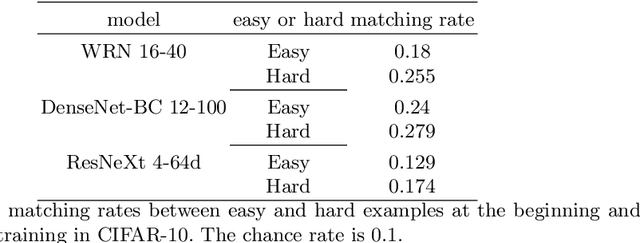
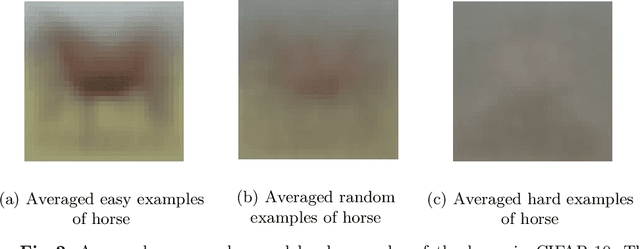


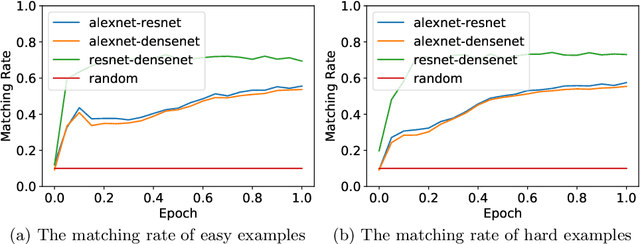
Abstract:Deep Neural Networks (DNNs) generalize well despite their massive size and capability of memorizing all examples. There is a hypothesis that DNNs start learning from simple patterns and the hypothesis is based on the existence of examples that are consistently well-classified at the early training stage (i.e., easy examples) and examples misclassified (i.e., hard examples). Easy examples are the evidence that DNNs start learning from specific patterns and there is a consistent learning process. It is important to know how DNNs learn patterns and obtain generalization ability, however, properties of easy and hard examples are not thoroughly investigated (e.g., contributions to generalization and visual appearances). In this work, we study the similarities of easy and hard examples respectively for different Convolutional Neural Network (CNN) architectures, assessing how those examples contribute to generalization. Our results show that easy examples are visually similar to each other and hard examples are visually diverse, and both examples are largely shared across different CNN architectures. Moreover, while hard examples tend to contribute more to generalization than easy examples, removing a large number of easy examples leads to poor generalization. By analyzing those results, we hypothesize that biases in a dataset and Stochastic Gradient Descent (SGD) are the reasons why CNNs have consistent easy and hard examples. Furthermore, we show that large scale classification datasets can be efficiently compressed by using easiness proposed in this work.
* Accepted to ICONIP 2019
Faster AutoAugment: Learning Augmentation Strategies using Backpropagation
Nov 16, 2019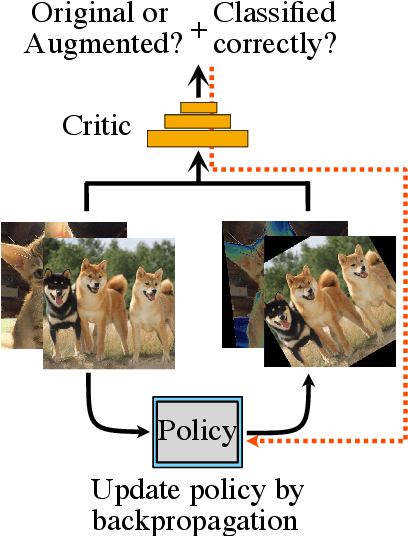
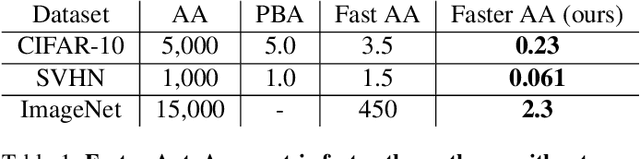
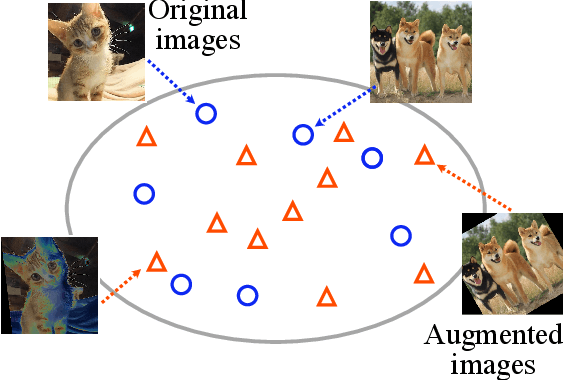
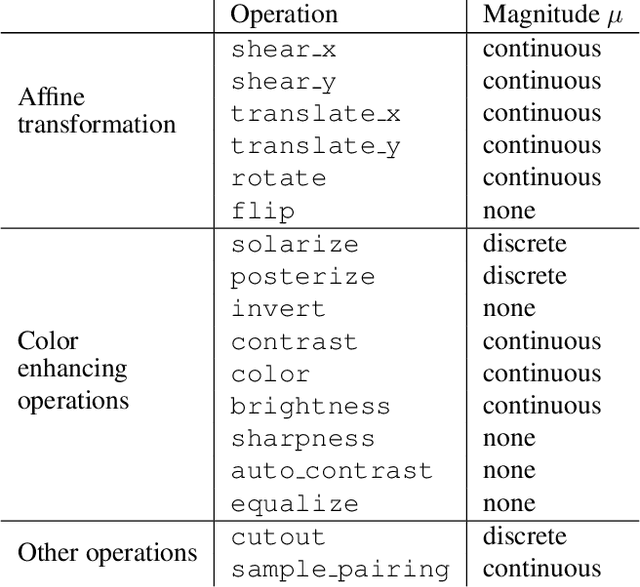
Abstract:Data augmentation methods are indispensable heuristics to boost the performance of deep neural networks, especially in image recognition tasks. Recently, several studies have shown that augmentation strategies found by search algorithms outperform hand-made strategies. Such methods employ black-box search algorithms over image transformations with continuous or discrete parameters and require a long time to obtain better strategies. In this paper, we propose a differentiable policy search pipeline for data augmentation, which is much faster than previous methods. We introduce approximate gradients for several transformation operations with discrete parameters as well as the differentiable mechanism for selecting operations. As the objective of training, we minimize the distance between the distributions of augmented data and the original data, which can be differentiated. We show that our method, Faster AutoAugment, achieves significantly faster searching than prior work without a performance drop.
Latent-Variable Non-Autoregressive Neural Machine Translation with Deterministic Inference using a Delta Posterior
Sep 10, 2019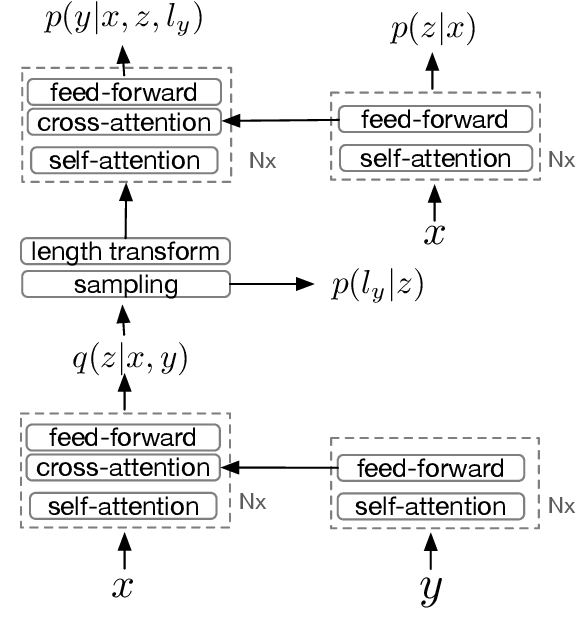
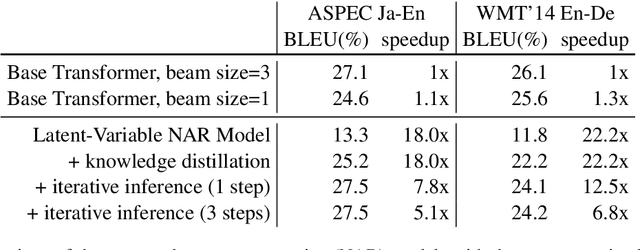
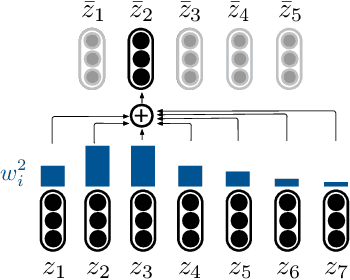
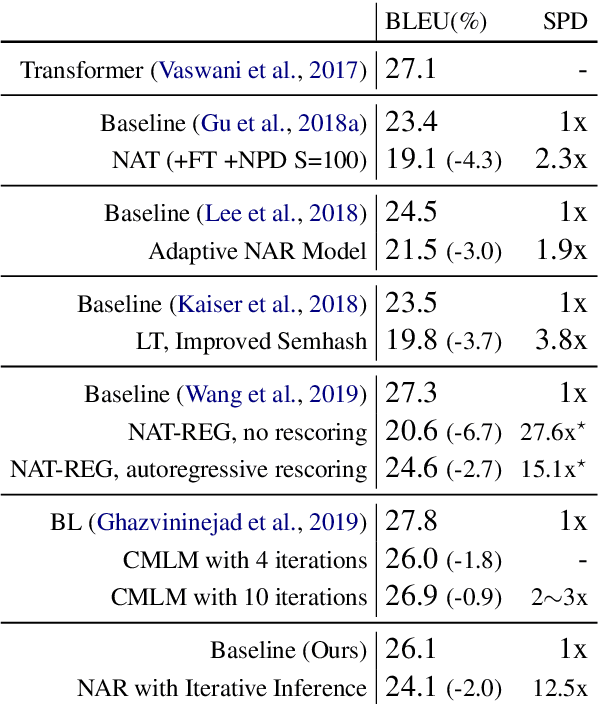
Abstract:Although neural machine translation models reached high translation quality, the autoregressive nature makes inference difficult to parallelize and leads to high translation latency. Inspired by recent refinement-based approaches, we propose a latent-variable non-autoregressive model with continuous latent variables and deterministic inference procedure. In contrast to existing approaches, we use a deterministic iterative inference algorithm to find a target sequence that maximizes the lowerbound to the log-probability. During inference, the length of translation automatically adapts itself. Our experiments show that the lowerbound can be greatly increased by running the inference algorithm for only one step, resulting in significantly improved translation quality. Our proposed model closes the gap between non-autoregressive and autoregressive approaches on ASPEC Ja-En dataset with 7.8x faster decoding. On WMT'14 En-De dataset, our model narrows the performance gap with autoregressive baseline down to 2.0 BLEU points with 12.5x speedup.
GAN-based Multiple Adjacent Brain MRI Slice Reconstruction for Unsupervised Alzheimer's Disease Diagnosis
Jul 08, 2019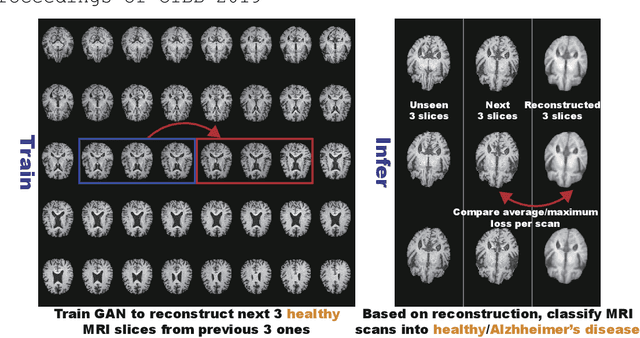
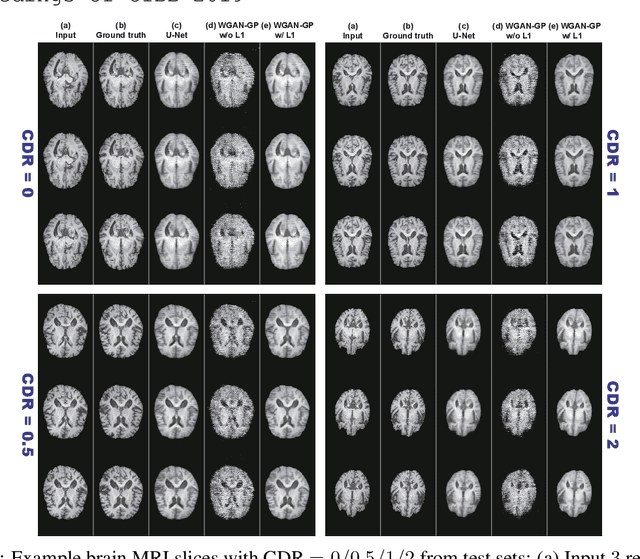
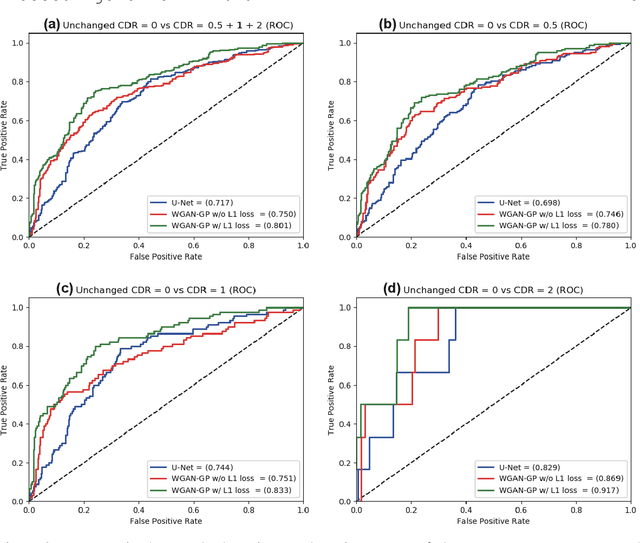
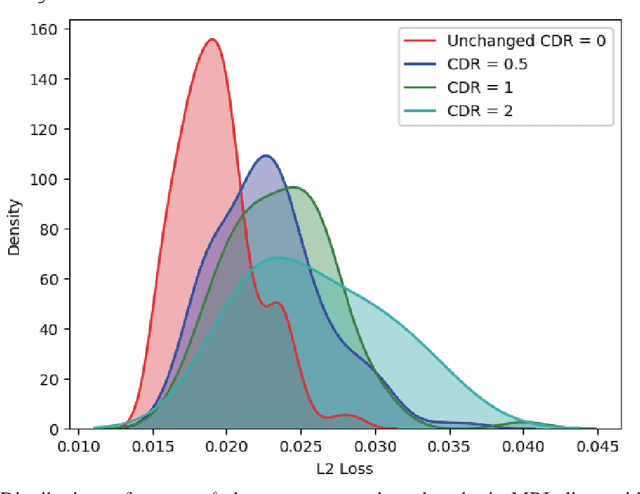
Abstract:Unsupervised learning can discover various unseen diseases, relying on large-scale unannotated medical images of healthy subjects. Towards this, unsupervised methods reconstruct a single medical image to detect outliers either in the learned feature space or from high reconstruction loss. However, without considering continuity between multiple adjacent images, they cannot directly discriminate diseases composed of the accumulation of subtle anatomical anomalies, such as Alzheimer's Disease (AD). Moreover, no study shows how unsupervised anomaly detection is associated with disease stages. Therefore, we propose a two-step method using Generative Adversarial Network-based multiple adjacent brain MRI slice reconstruction to detect AD at various stages: (Reconstruction) Wasserstein loss with Gradient Penalty + L1 loss---trained on 3 healthy slices to reconstruct the next 3 ones---reconstructs unseen healthy/AD cases; (Diagnosis) Average/Maximum loss (e.g., L2 loss) per scan discriminates them, comparing the reconstructed/ground truth images. The results show that we can reliably detect AD at a very early stage with Area Under the Curve (AUC) 0.780 while also detecting AD at a late stage much more accurately with AUC 0.917; since our method is unsupervised, it should also discover and alert any anomalies including rare disease.
Synthesizing Diverse Lung Nodules Wherever Massively: 3D Multi-Conditional GAN-based CT Image Augmentation for Object Detection
Jun 12, 2019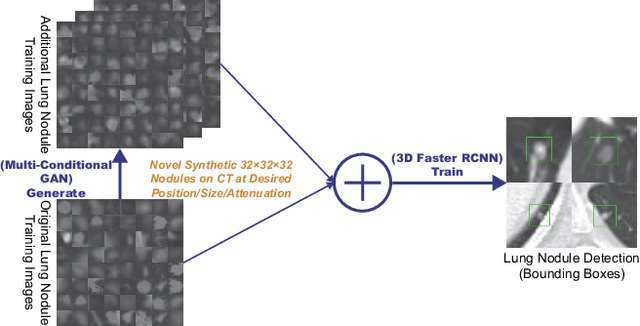
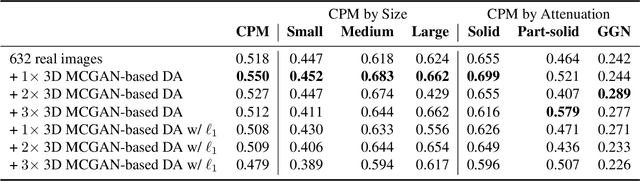
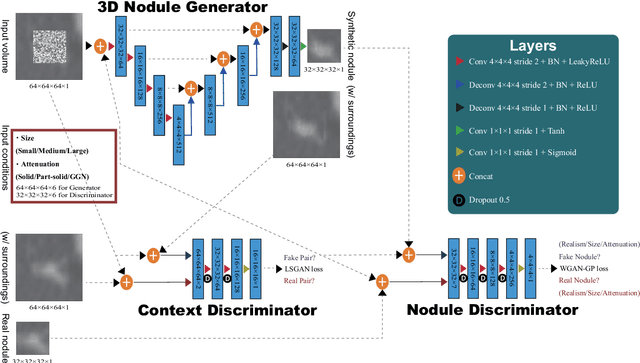
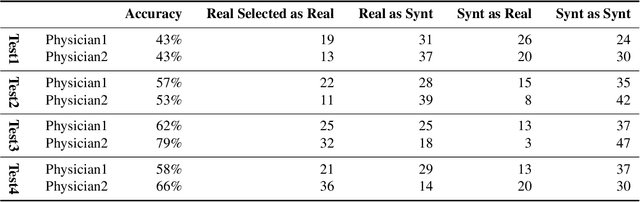
Abstract:Accurate computer-assisted diagnosis, relying on large-scale annotated pathological images, can alleviate the risk of overlooking the diagnosis. Unfortunately, in medical imaging, most available datasets are small/fragmented. To tackle this, as a Data Augmentation (DA) method, 3D conditional Generative Adversarial Networks (GANs) can synthesize desired realistic/diverse 3D images as additional training data. However, no 3D conditional GAN-based DA approach exists for general bounding box-based 3D object detection, while it can locate disease areas with physicians' minimum annotation cost, unlike rigorous 3D segmentation. Moreover, since lesions vary in position/size/attenuation, further GAN-based DA performance requires multiple conditions. Therefore, we propose 3D Multi-Conditional GAN (MCGAN) to generate realistic/diverse 32 x 32 x 32 nodules placed naturally on lung Computed Tomography images to boost sensitivity in 3D object detection. Our MCGAN adopts two discriminators for conditioning: the context discriminator learns to classify real vs synthetic nodule/surrounding pairs with noise box-centered surroundings; the nodule discriminator attempts to classify real vs synthetic nodules with size/attenuation conditions. The results show that 3D Convolutional Neural Network-based detection can achieve higher sensitivity under any nodule size/attenuation at fixed False Positive rates and overcome the medical data paucity with the MCGAN-generated realistic nodules---even expert physicians fail to distinguish them from the real ones in Visual Turing Test.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge