Hideki Nakayama
StoryER: Automatic Story Evaluation via Ranking, Rating and Reasoning
Oct 16, 2022
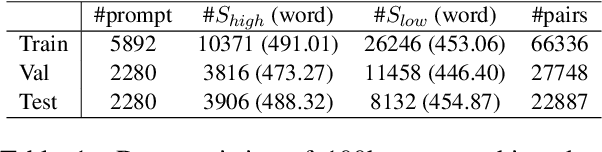
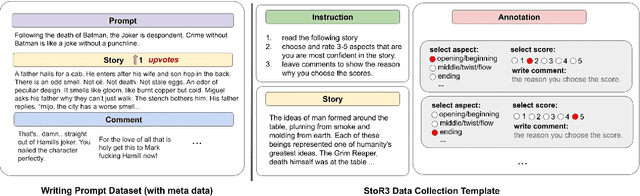
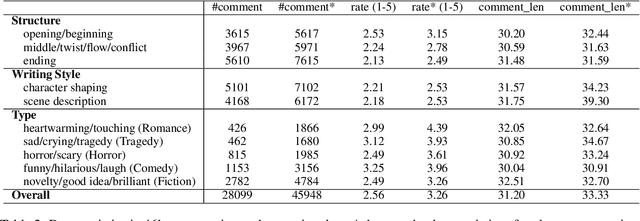
Abstract:Existing automatic story evaluation methods place a premium on story lexical level coherence, deviating from human preference. We go beyond this limitation by considering a novel \textbf{Story} \textbf{E}valuation method that mimics human preference when judging a story, namely \textbf{StoryER}, which consists of three sub-tasks: \textbf{R}anking, \textbf{R}ating and \textbf{R}easoning. Given either a machine-generated or a human-written story, StoryER requires the machine to output 1) a preference score that corresponds to human preference, 2) specific ratings and their corresponding confidences and 3) comments for various aspects (e.g., opening, character-shaping). To support these tasks, we introduce a well-annotated dataset comprising (i) 100k ranked story pairs; and (ii) a set of 46k ratings and comments on various aspects of the story. We finetune Longformer-Encoder-Decoder (LED) on the collected dataset, with the encoder responsible for preference score and aspect prediction and the decoder for comment generation. Our comprehensive experiments result in a competitive benchmark for each task, showing the high correlation to human preference. In addition, we have witnessed the joint learning of the preference scores, the aspect ratings, and the comments brings gain in each single task. Our dataset and benchmarks are publicly available to advance the research of story evaluation tasks.\footnote{Dataset and pre-trained model demo are available at anonymous website \url{http://storytelling-lab.com/eval} and \url{https://github.com/sairin1202/StoryER}}
Towards Parameter-Efficient Integration of Pre-Trained Language Models In Temporal Video Grounding
Sep 26, 2022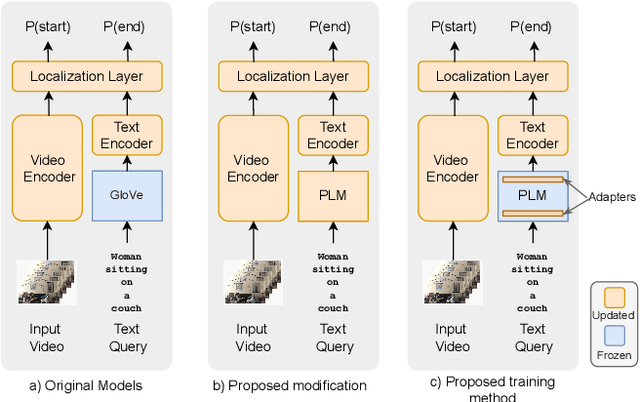
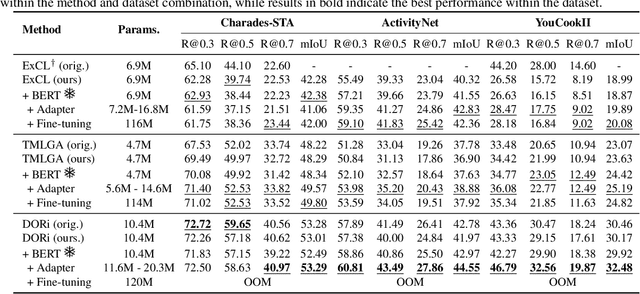
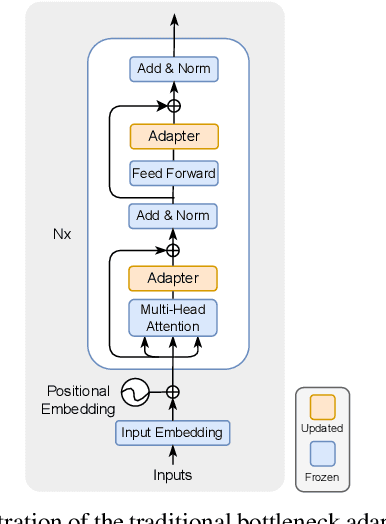
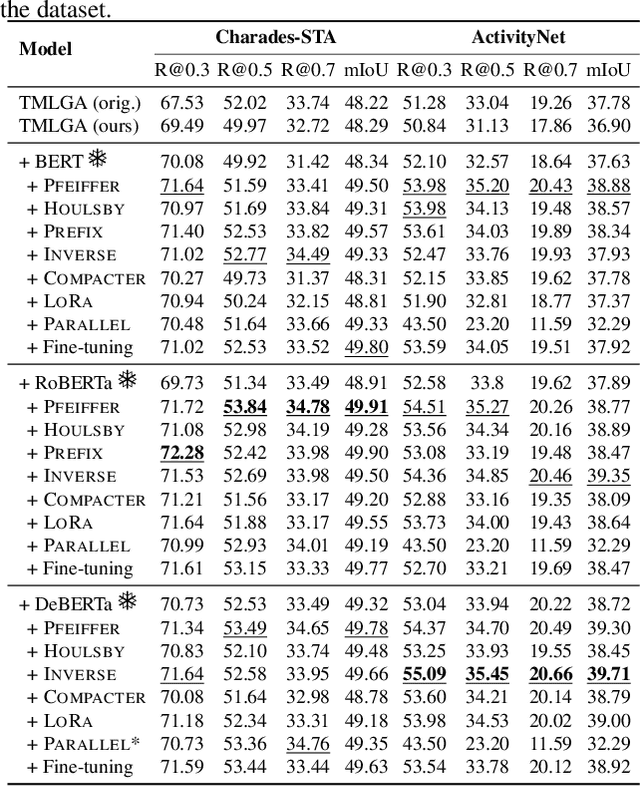
Abstract:This paper explores the task of Temporal Video Grounding (TVG) where, given an untrimmed video and a query sentence, the goal is to recognize and determine temporal boundaries of action instances in the video described by the provided natural language queries. Recent works solve this task by directly encoding the query using large pre-trained language models (PLM). However, isolating the effects of the improved language representations is difficult, as these works also propose improvements in the visual inputs. Furthermore, these PLMs significantly increase the computational cost of training TVG models. Therefore, this paper studies the effects of PLMs in the TVG task and assesses the applicability of NLP parameter-efficient training alternatives based on adapters. We couple popular PLMs with a selection of existing approaches and test different adapters to reduce the impact of the additional parameters. Our results on three challenging datasets show that TVG models could greatly benefit from PLMs when these are fine-tuned for the task and that adapters are an effective alternative to full fine-tuning, even though they are not tailored for our task. Concretely, adapters helped save on computational cost, allowing PLM integration in larger TVG models and delivering results comparable to the state-of-the-art models. Finally, through benchmarking different types of adapters in TVG, our results shed light on what kind of adapters work best for each studied case.
Pixel to Binary Embedding Towards Robustness for CNNs
Jun 13, 2022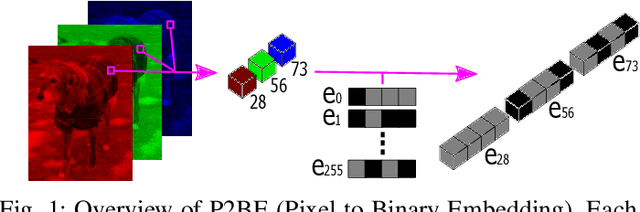
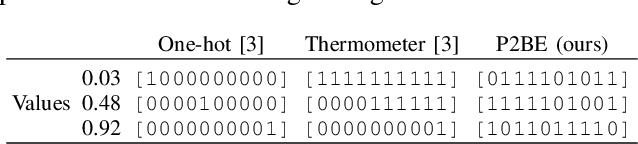
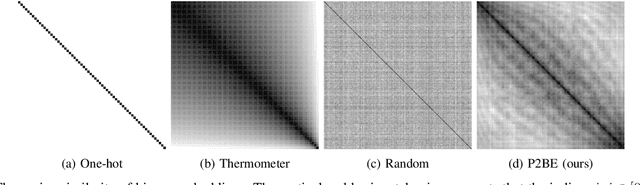
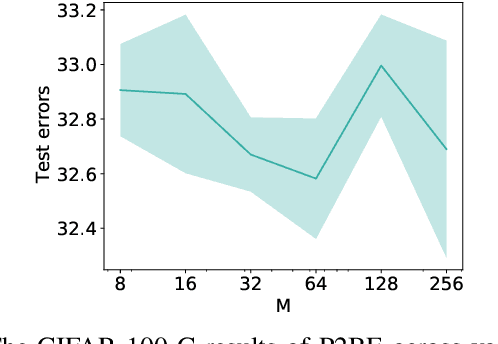

Abstract:There are several problems with the robustness of Convolutional Neural Networks (CNNs). For example, the prediction of CNNs can be changed by adding a small magnitude of noise to an input, and the performances of CNNs are degraded when the distribution of input is shifted by a transformation never seen during training (e.g., the blur effect). There are approaches to replace pixel values with binary embeddings to tackle the problem of adversarial perturbations, which successfully improve robustness. In this work, we propose Pixel to Binary Embedding (P2BE) to improve the robustness of CNNs. P2BE is a learnable binary embedding method as opposed to previous hand-coded binary embedding methods. P2BE outperforms other binary embedding methods in robustness against adversarial perturbations and visual corruptions that are not shown during training.
OSSGAN: Open-Set Semi-Supervised Image Generation
Apr 29, 2022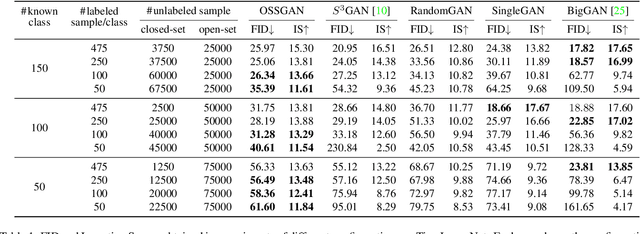
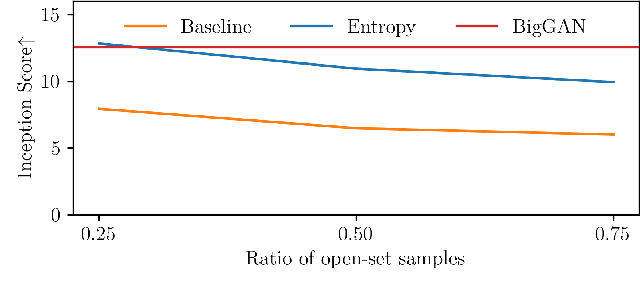
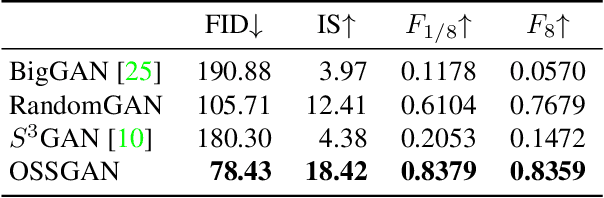
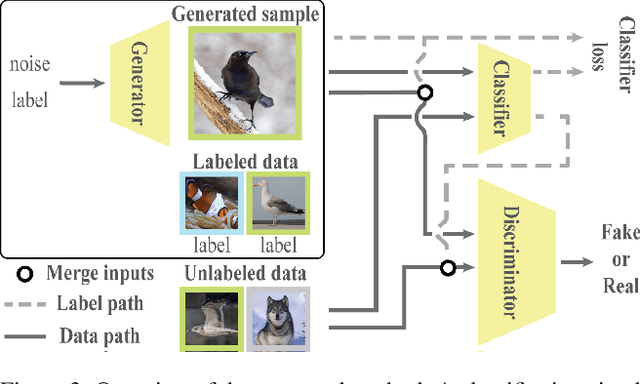
Abstract:We introduce a challenging training scheme of conditional GANs, called open-set semi-supervised image generation, where the training dataset consists of two parts: (i) labeled data and (ii) unlabeled data with samples belonging to one of the labeled data classes, namely, a closed-set, and samples not belonging to any of the labeled data classes, namely, an open-set. Unlike the existing semi-supervised image generation task, where unlabeled data only contain closed-set samples, our task is more general and lowers the data collection cost in practice by allowing open-set samples to appear. Thanks to entropy regularization, the classifier that is trained on labeled data is able to quantify sample-wise importance to the training of cGAN as confidence, allowing us to use all samples in unlabeled data. We design OSSGAN, which provides decision clues to the discriminator on the basis of whether an unlabeled image belongs to one or none of the classes of interest, smoothly integrating labeled and unlabeled data during training. The results of experiments on Tiny ImageNet and ImageNet show notable improvements over supervised BigGAN and semi-supervised methods. Our code is available at https://github.com/raven38/OSSGAN.
NOC-REK: Novel Object Captioning with Retrieved Vocabulary from External Knowledge
Mar 28, 2022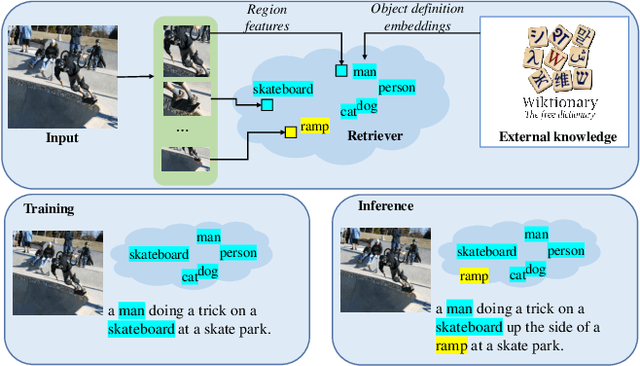
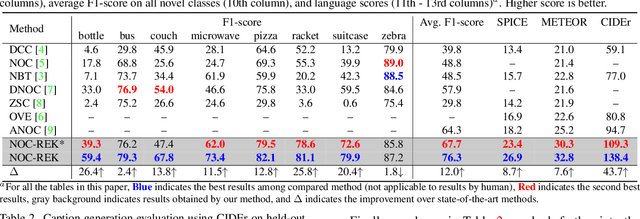
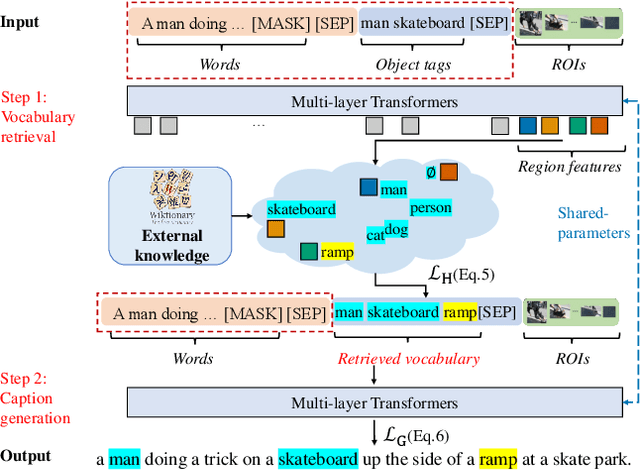

Abstract:Novel object captioning aims at describing objects absent from training data, with the key ingredient being the provision of object vocabulary to the model. Although existing methods heavily rely on an object detection model, we view the detection step as vocabulary retrieval from an external knowledge in the form of embeddings for any object's definition from Wiktionary, where we use in the retrieval image region features learned from a transformers model. We propose an end-to-end Novel Object Captioning with Retrieved vocabulary from External Knowledge method (NOC-REK), which simultaneously learns vocabulary retrieval and caption generation, successfully describing novel objects outside of the training dataset. Furthermore, our model eliminates the requirement for model retraining by simply updating the external knowledge whenever a novel object appears. Our comprehensive experiments on held-out COCO and Nocaps datasets show that our NOC-REK is considerably effective against SOTAs.
PPCD-GAN: Progressive Pruning and Class-Aware Distillation for Large-Scale Conditional GANs Compression
Mar 16, 2022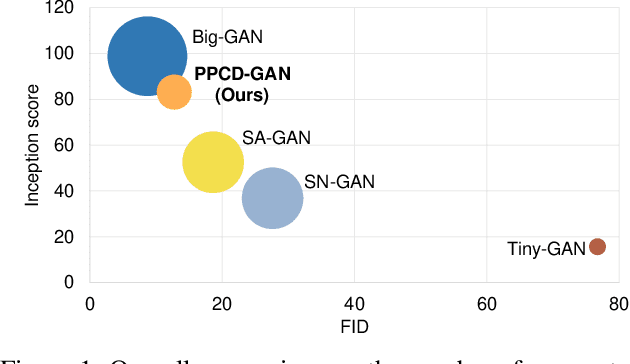
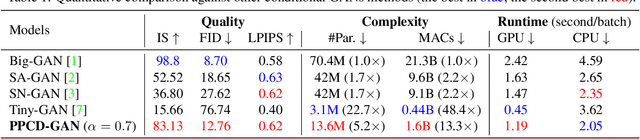
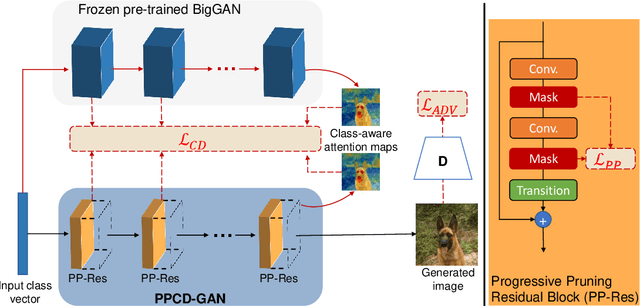
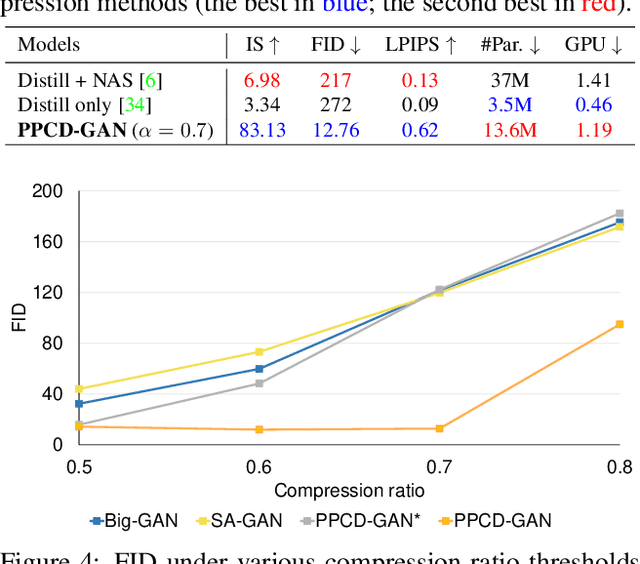
Abstract:We push forward neural network compression research by exploiting a novel challenging task of large-scale conditional generative adversarial networks (GANs) compression. To this end, we propose a gradually shrinking GAN (PPCD-GAN) by introducing progressive pruning residual block (PP-Res) and class-aware distillation. The PP-Res is an extension of the conventional residual block where each convolutional layer is followed by a learnable mask layer to progressively prune network parameters as training proceeds. The class-aware distillation, on the other hand, enhances the stability of training by transferring immense knowledge from a well-trained teacher model through instructive attention maps. We train the pruning and distillation processes simultaneously on a well-known GAN architecture in an end-to-end manner. After training, all redundant parameters as well as the mask layers are discarded, yielding a lighter network while retaining the performance. We comprehensively illustrate, on ImageNet 128x128 dataset, PPCD-GAN reduces up to 5.2x (81%) parameters against state-of-the-arts while keeping better performance.
SciXGen: A Scientific Paper Dataset for Context-Aware Text Generation
Oct 20, 2021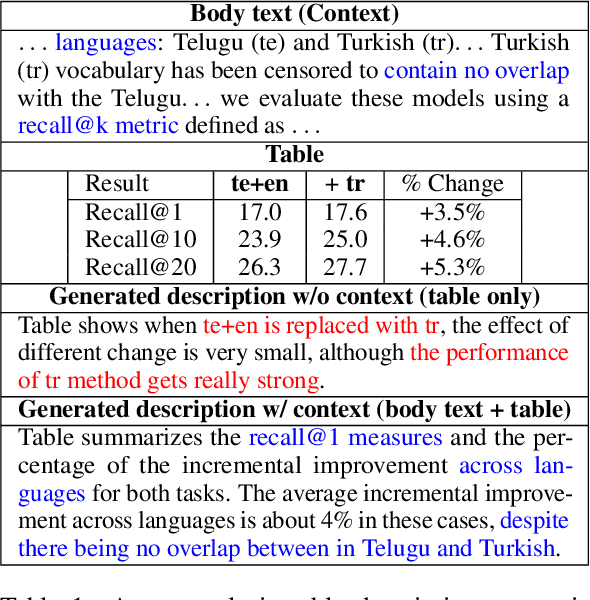
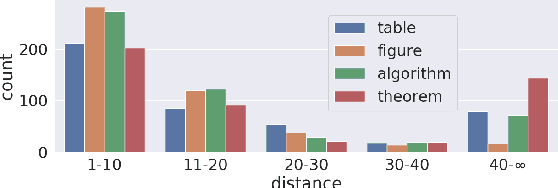
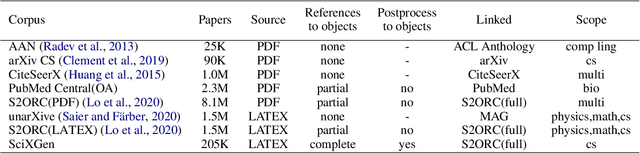
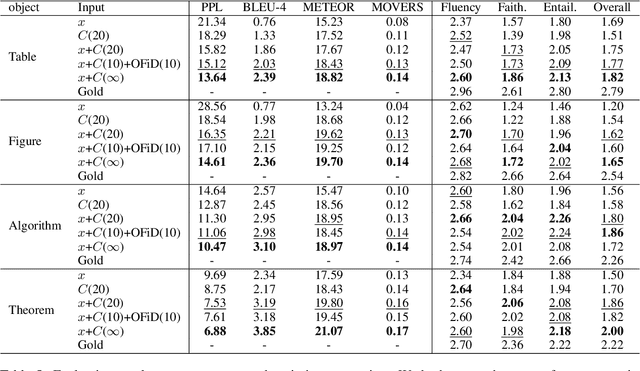
Abstract:Generating texts in scientific papers requires not only capturing the content contained within the given input but also frequently acquiring the external information called \textit{context}. We push forward the scientific text generation by proposing a new task, namely \textbf{context-aware text generation} in the scientific domain, aiming at exploiting the contributions of context in generated texts. To this end, we present a novel challenging large-scale \textbf{Sci}entific Paper Dataset for Conte\textbf{X}t-Aware Text \textbf{Gen}eration (SciXGen), consisting of well-annotated 205,304 papers with full references to widely-used objects (e.g., tables, figures, algorithms) in a paper. We comprehensively benchmark, using state-of-the-arts, the efficacy of our newly constructed SciXGen dataset in generating description and paragraph. Our dataset and benchmarks will be made publicly available to hopefully facilitate the scientific text generation research.
Graph Energy-based Model for Substructure Preserving Molecular Design
Feb 09, 2021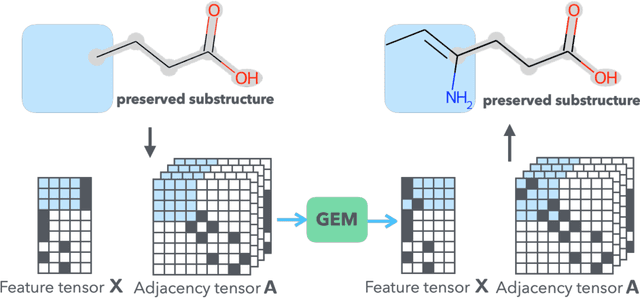
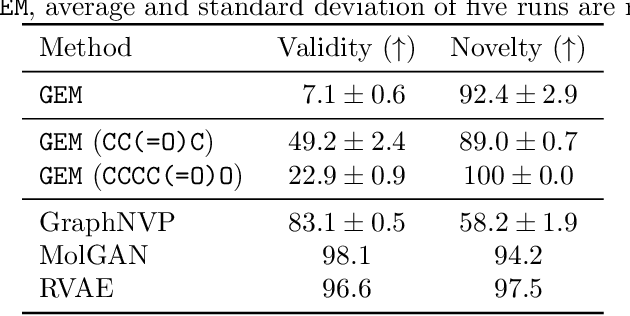
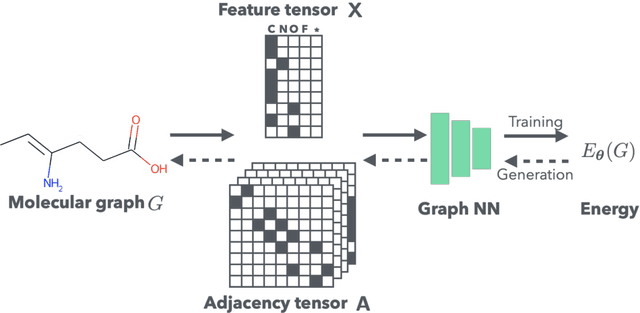
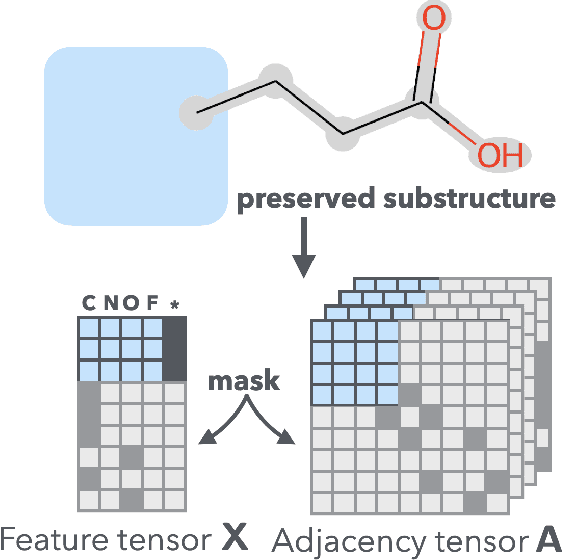
Abstract:It is common practice for chemists to search chemical databases based on substructures of compounds for finding molecules with desired properties. The purpose of de novo molecular generation is to generate instead of search. Existing machine learning based molecular design methods have no or limited ability in generating novel molecules that preserves a target substructure. Our Graph Energy-based Model, or GEM, can fix substructures and generate the rest. The experimental results show that the GEMs trained from chemistry datasets successfully generate novel molecules while preserving the target substructures. This method would provide a new way of incorporating the domain knowledge of chemists in molecular design.
GraphPlan: Story Generation by Planning with Event Graph
Feb 05, 2021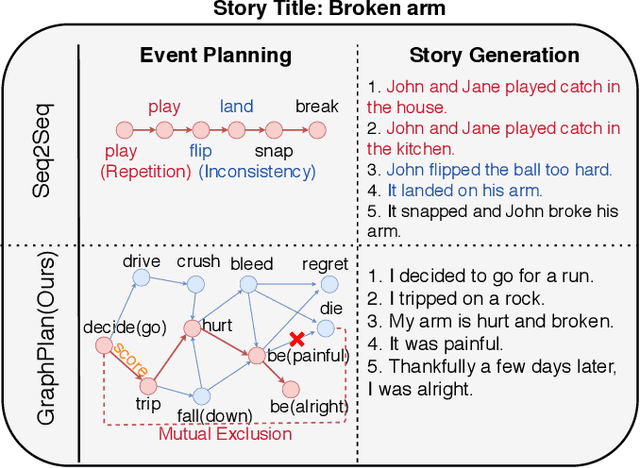
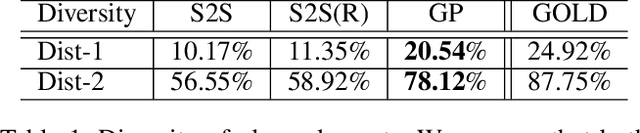
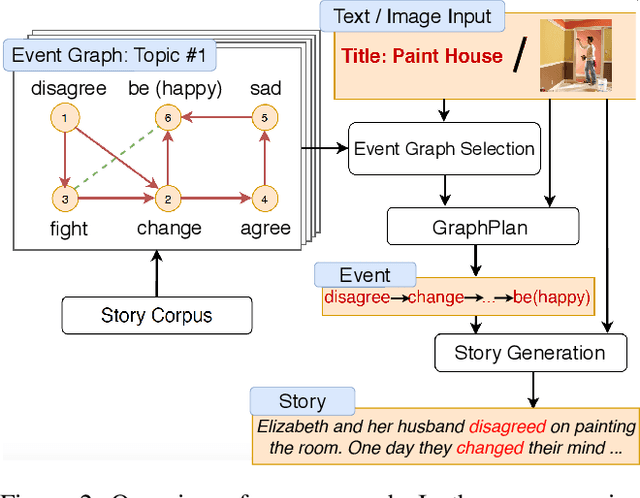
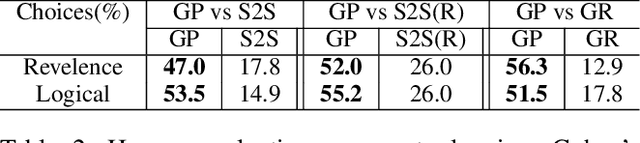
Abstract:Story generation is a task that aims to automatically produce multiple sentences to make up a meaningful story. This task is challenging because it requires high-level understanding of semantic meaning of sentences and causality of story events. Naive sequence-to-sequence models generally fail to acquire such knowledge, as the logical correctness can hardly be guaranteed in a text generation model without the strategic planning. In this paper, we focus on planning a sequence of events assisted by event graphs, and use the events to guide the generator. Instead of using a sequence-to-sequence model to output a storyline as in some existing works, we propose to generate an event sequence by walking on an event graph. The event graphs are built automatically based on the corpus. To evaluate the proposed approach, we conduct human evaluation both on event planning and story generation. Based on large-scale human annotation results, our proposed approach is shown to produce more logically correct event sequences and stories.
Commonsense Knowledge Aware Concept Selection For Diverse and Informative Visual Storytelling
Feb 05, 2021
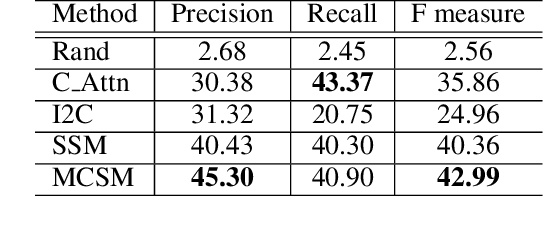
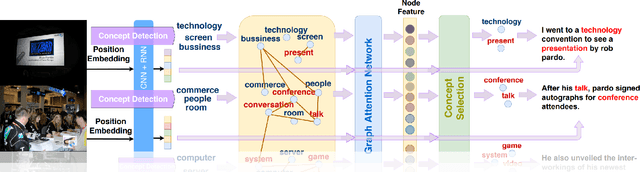
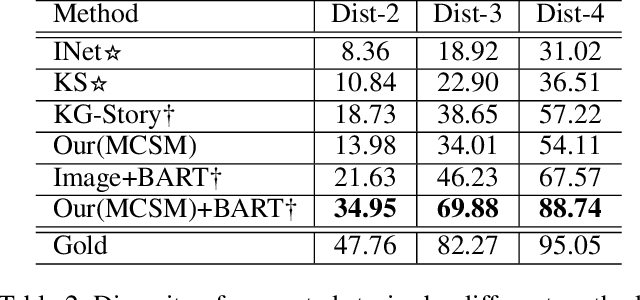
Abstract:Visual storytelling is a task of generating relevant and interesting stories for given image sequences. In this work we aim at increasing the diversity of the generated stories while preserving the informative content from the images. We propose to foster the diversity and informativeness of a generated story by using a concept selection module that suggests a set of concept candidates. Then, we utilize a large scale pre-trained model to convert concepts and images into full stories. To enrich the candidate concepts, a commonsense knowledge graph is created for each image sequence from which the concept candidates are proposed. To obtain appropriate concepts from the graph, we propose two novel modules that consider the correlation among candidate concepts and the image-concept correlation. Extensive automatic and human evaluation results demonstrate that our model can produce reasonable concepts. This enables our model to outperform the previous models by a large margin on the diversity and informativeness of the story, while retaining the relevance of the story to the image sequence.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge