Haslina Makmur
HistDiT: A Structure-Aware Latent Conditional Diffusion Model for High-Fidelity Virtual Staining in Histopathology
Apr 09, 2026Abstract:Immunohistochemistry (IHC) is essential for assessing specific immune biomarkers like Human Epidermal growth-factor Receptor 2 (HER2) in breast cancer. However, the traditional protocols of obtaining IHC stains are resource-intensive, time-consuming, and prone to structural damages. Virtual staining has emerged as a scalable alternative, but it faces significant challenges in preserving fine-grained cellular structures while accurately translating biochemical expressions. Current state-of-the-art methods still rely on Generative Adversarial Networks (GANs) or standard convolutional U-Net diffusion models that often struggle with "structure and staining trade-offs". The generated samples are either structurally relevant but blurry, or texturally realistic but have artifacts that compromise their diagnostic use. In this paper, we introduce HistDiT, a novel latent conditional Diffusion Transformer (DiT) architecture that establishes a new benchmark for visual fidelity in virtual histological staining. The novelty introduced in this work is, a) the Dual-Stream Conditioning strategy that explicitly maintains a balance between spatial constraints via VAE-encoded latents and semantic phenotype guidance via UNI embeddings; b) the multi-objective loss function that contributes to sharper images with clear morphological structure; and c) the use of the Structural Correlation Metric (SCM) to focus on the core morphological structure for precise assessment of sample quality. Consequently, our model outperforms existing baselines, as demonstrated through rigorous quantitative and qualitative evaluations.
Multi-modal AI for comprehensive breast cancer prognostication
Oct 28, 2024


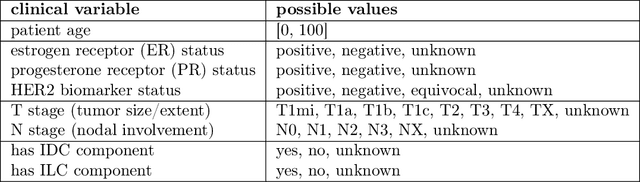
Abstract:Treatment selection in breast cancer is guided by molecular subtypes and clinical characteristics. Recurrence risk assessment plays a crucial role in personalizing treatment. Current methods, including genomic assays, have limited accuracy and clinical utility, leading to suboptimal decisions for many patients. We developed a test for breast cancer patient stratification based on digital pathology and clinical characteristics using novel AI methods. Specifically, we utilized a vision transformer-based pan-cancer foundation model trained with self-supervised learning to extract features from digitized H&E-stained slides. These features were integrated with clinical data to form a multi-modal AI test predicting cancer recurrence and death. The test was developed and evaluated using data from a total of 8,161 breast cancer patients across 15 cohorts originating from seven countries. Of these, 3,502 patients from five cohorts were used exclusively for evaluation, while the remaining patients were used for training. Our test accurately predicted our primary endpoint, disease-free interval, in the five external cohorts (C-index: 0.71 [0.68-0.75], HR: 3.63 [3.02-4.37, p<0.01]). In a direct comparison (N=858), the AI test was more accurate than Oncotype DX, the standard-of-care 21-gene assay, with a C-index of 0.67 [0.61-0.74] versus 0.61 [0.49-0.73], respectively. Additionally, the AI test added independent information to Oncotype DX in a multivariate analysis (HR: 3.11 [1.91-5.09, p<0.01)]). The test demonstrated robust accuracy across all major breast cancer subtypes, including TNBC (C-index: 0.71 [0.62-0.81], HR: 3.81 [2.35-6.17, p=0.02]), where no diagnostic tools are currently recommended by clinical guidelines. These results suggest that our AI test can improve accuracy, extend applicability to a wider range of patients, and enhance access to treatment selection tools.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge