Gene Myers
Towards Interpretable Semantic Segmentation via Gradient-weighted Class Activation Mapping
Feb 26, 2020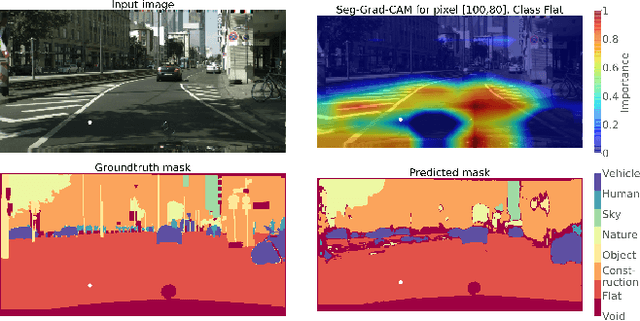
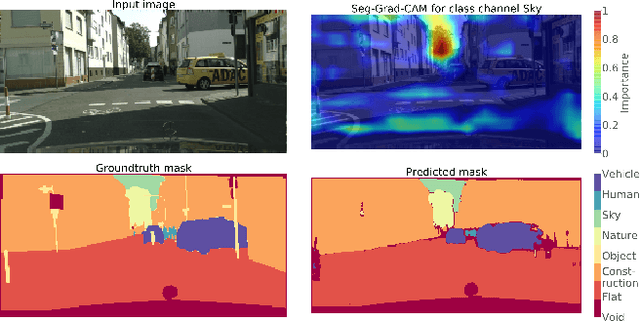
Abstract:Convolutional neural networks have become state-of-the-art in a wide range of image recognition tasks. The interpretation of their predictions, however, is an active area of research. Whereas various interpretation methods have been suggested for image classification, the interpretation of image segmentation still remains largely unexplored. To that end, we propose SEG-GRAD-CAM, a gradient-based method for interpreting semantic segmentation. Our method is an extension of the widely-used Grad-CAM method, applied locally to produce heatmaps showing the relevance of individual pixels for semantic segmentation.
* 2 pages, 2 figures. AAAI 2020 camera-ready
Star-convex Polyhedra for 3D Object Detection and Segmentation in Microscopy
Aug 09, 2019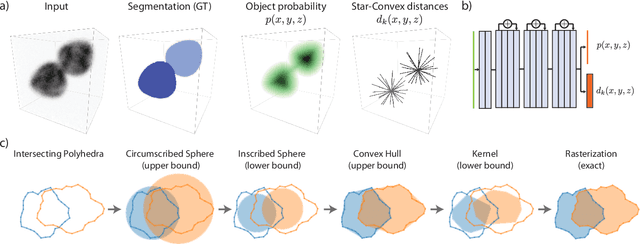
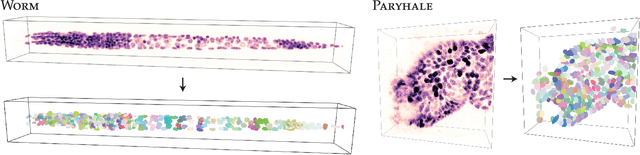
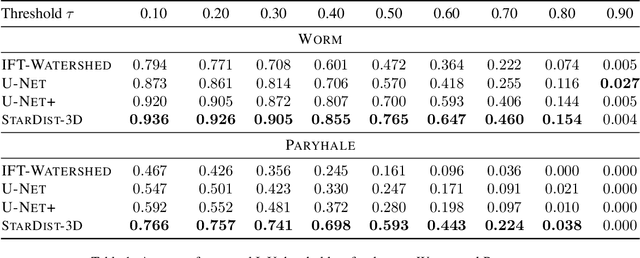
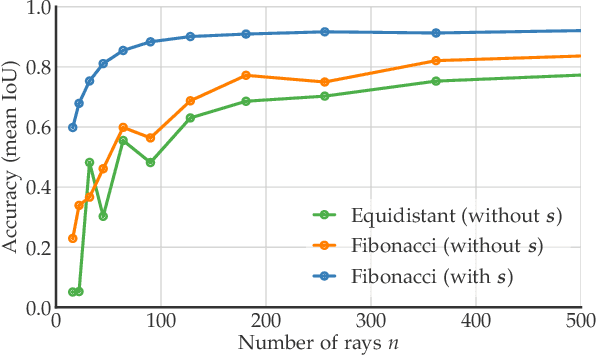

Abstract:Accurate detection and segmentation of cell nuclei in volumetric (3D) fluorescence microscopy datasets is an important step in many biomedical research projects. Although many automated methods for these tasks exist, they often struggle for images with low signal-to-noise ratios and/or dense packing of nuclei. It was recently shown for 2D microscopy images that these issues can be alleviated by training a neural network to directly predict a suitable shape representation (star-convex polygon) for cell nuclei. In this paper, we adopt and extend this approach to 3D volumes by using star-convex polyhedra to represent cell nuclei and similar shapes. To that end, we overcome the challenges of 1) finding parameter-efficient star-convex polyhedra representations that can faithfully describe cell nuclei shapes, 2) adapting to anisotropic voxel sizes often found in fluorescence microscopy datasets, and 3) efficiently computing intersections between pairs of star-convex polyhedra (required for non-maximum suppression). Although our approach is quite general, since star-convex polyhedra subsume common shapes like bounding boxes and spheres as special cases, our focus is on accurate detection and segmentation of cell nuclei. That that end, we demonstrate on two challenging datasets that our approach (StarDist-3D) leads to superior results when compared to classical and deep-learning based methods.
Cell Detection with Star-convex Polygons
Jun 09, 2018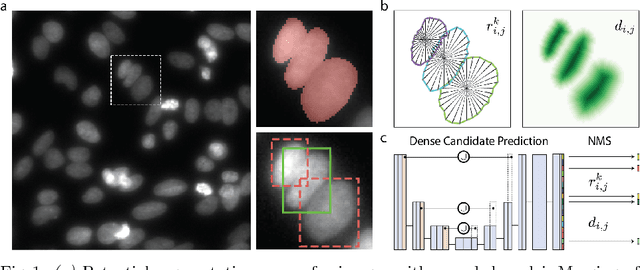
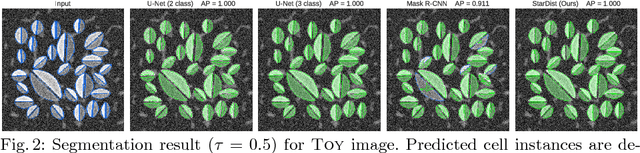
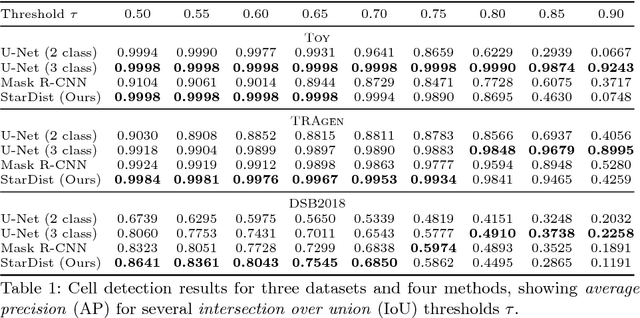


Abstract:Automatic detection and segmentation of cells and nuclei in microscopy images is important for many biological applications. Recent successful learning-based approaches include per-pixel cell segmentation with subsequent pixel grouping, or localization of bounding boxes with subsequent shape refinement. In situations of crowded cells, these can be prone to segmentation errors, such as falsely merging bordering cells or suppressing valid cell instances due to the poor approximation with bounding boxes. To overcome these issues, we propose to localize cell nuclei via star-convex polygons, which are a much better shape representation as compared to bounding boxes and thus do not need shape refinement. To that end, we train a convolutional neural network that predicts for every pixel a polygon for the cell instance at that position. We demonstrate the merits of our approach on two synthetic datasets and one challenging dataset of diverse fluorescence microscopy images.
Isotropic reconstruction of 3D fluorescence microscopy images using convolutional neural networks
Apr 05, 2017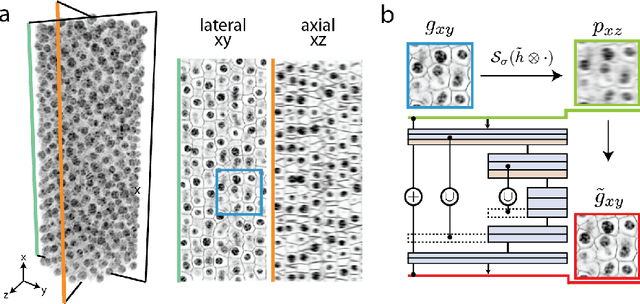
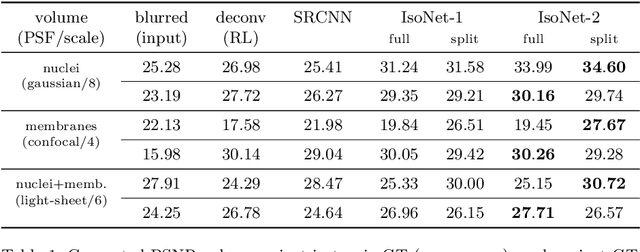

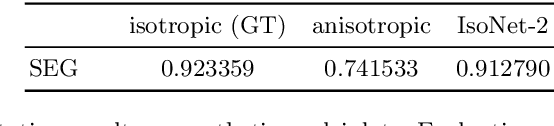
Abstract:Fluorescence microscopy images usually show severe anisotropy in axial versus lateral resolution. This hampers downstream processing, i.e. the automatic extraction of quantitative biological data. While deconvolution methods and other techniques to address this problem exist, they are either time consuming to apply or limited in their ability to remove anisotropy. We propose a method to recover isotropic resolution from readily acquired anisotropic data. We achieve this using a convolutional neural network that is trained end-to-end from the same anisotropic body of data we later apply the network to. The network effectively learns to restore the full isotropic resolution by restoring the image under a trained, sample specific image prior. We apply our method to $3$ synthetic and $3$ real datasets and show that our results improve on results from deconvolution and state-of-the-art super-resolution techniques. Finally, we demonstrate that a standard 3D segmentation pipeline performs on the output of our network with comparable accuracy as on the full isotropic data.
Crowd Sourcing Image Segmentation with iaSTAPLE
Feb 21, 2017
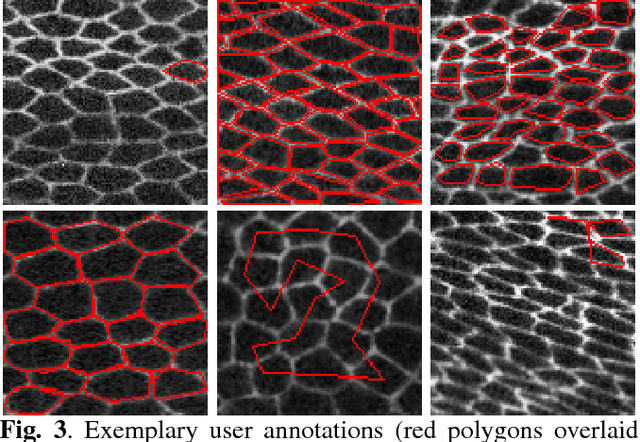
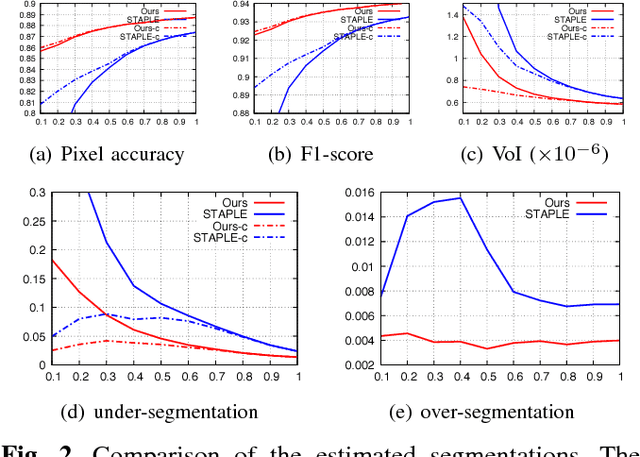

Abstract:We propose a novel label fusion technique as well as a crowdsourcing protocol to efficiently obtain accurate epithelial cell segmentations from non-expert crowd workers. Our label fusion technique simultaneously estimates the true segmentation, the performance levels of individual crowd workers, and an image segmentation model in the form of a pairwise Markov random field. We term our approach image-aware STAPLE (iaSTAPLE) since our image segmentation model seamlessly integrates into the well-known and widely used STAPLE approach. In an evaluation on a light microscopy dataset containing more than 5000 membrane labeled epithelial cells of a fly wing, we show that iaSTAPLE outperforms STAPLE in terms of segmentation accuracy as well as in terms of the accuracy of estimated crowd worker performance levels, and is able to correctly segment 99% of all cells when compared to expert segmentations. These results show that iaSTAPLE is a highly useful tool for crowd sourcing image segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge