Kira Vinogradova
Estimation of Optical Aberrations in 3D Microscopic Bioimages
Sep 16, 2022
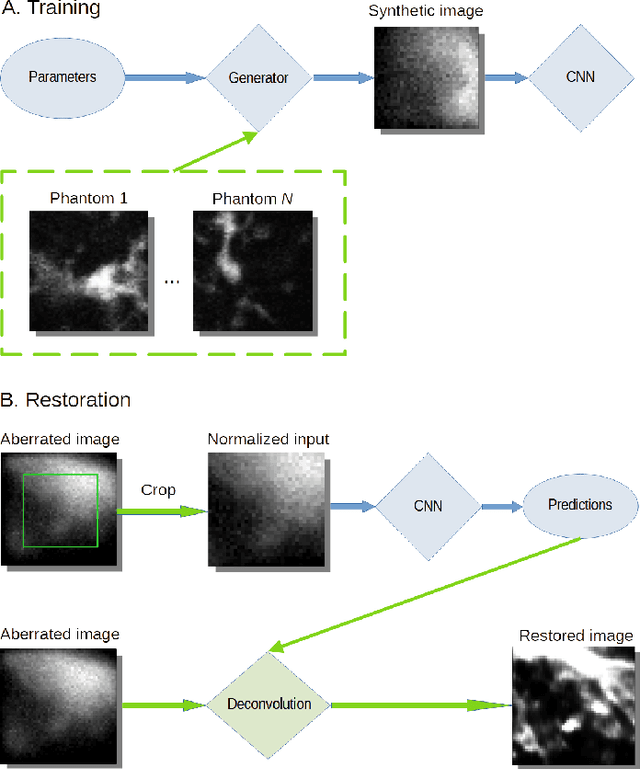
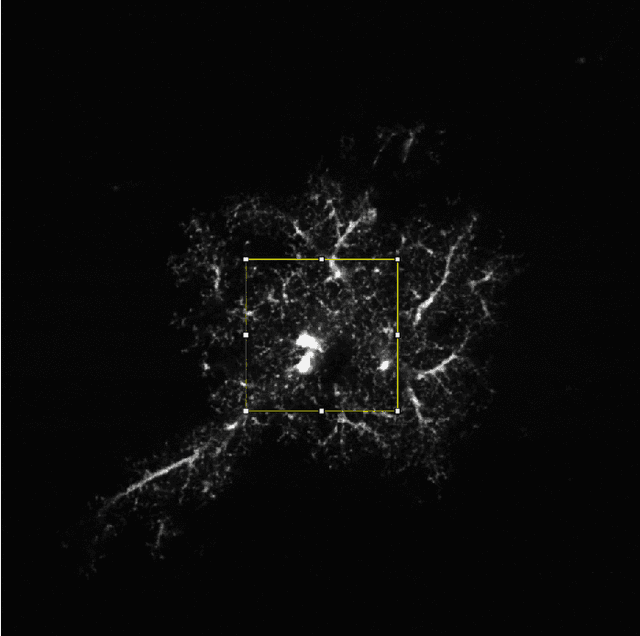
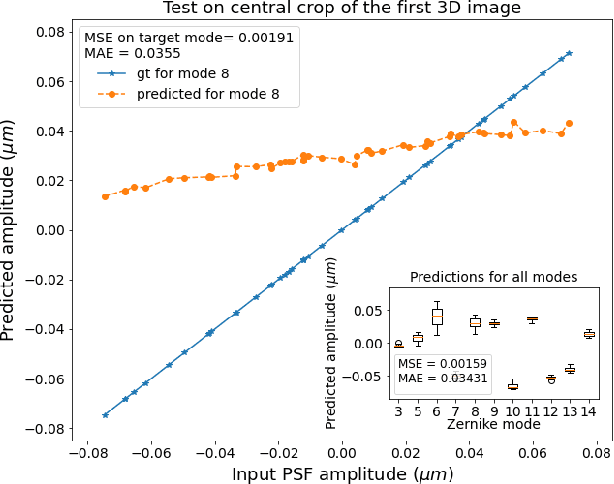
Abstract:The quality of microscopy images often suffers from optical aberrations. These aberrations and their associated point spread functions have to be quantitatively estimated to restore aberrated images. The recent state-of-the-art method PhaseNet, based on a convolutional neural network, can quantify aberrations accurately but is limited to images of point light sources, e.g. fluorescent beads. In this research, we describe an extension of PhaseNet enabling its use on 3D images of biological samples. To this end, our method incorporates object-specific information into the simulated images used for training the network. Further, we add a Python-based restoration of images via Richardson-Lucy deconvolution. We demonstrate that the deconvolution with the predicted PSF can not only remove the simulated aberrations but also improve the quality of the real raw microscopic images with unknown residual PSF. We provide code for fast and convenient prediction and correction of aberrations.
Towards Interpretable Semantic Segmentation via Gradient-weighted Class Activation Mapping
Feb 26, 2020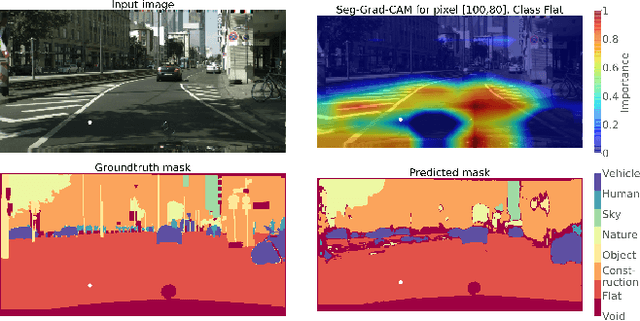
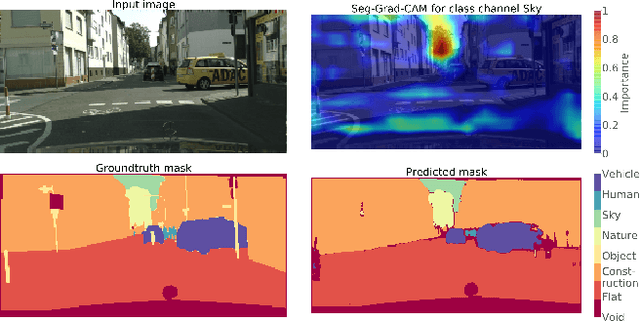
Abstract:Convolutional neural networks have become state-of-the-art in a wide range of image recognition tasks. The interpretation of their predictions, however, is an active area of research. Whereas various interpretation methods have been suggested for image classification, the interpretation of image segmentation still remains largely unexplored. To that end, we propose SEG-GRAD-CAM, a gradient-based method for interpreting semantic segmentation. Our method is an extension of the widely-used Grad-CAM method, applied locally to produce heatmaps showing the relevance of individual pixels for semantic segmentation.
* 2 pages, 2 figures. AAAI 2020 camera-ready
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge