Gari D. Clifford
Department of Biomedical Informatics, Emory University, Atlanta, USA, Department of Biomedical Engineering, Georgia Institute of Technology and Emory University, Atlanta, USA
Beyond Heart Murmur Detection: Automatic Murmur Grading from Phonocardiogram
Sep 27, 2022
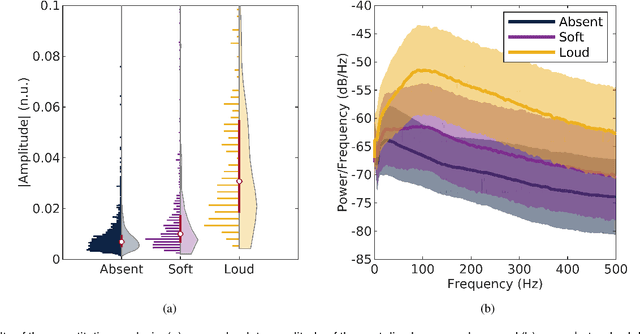
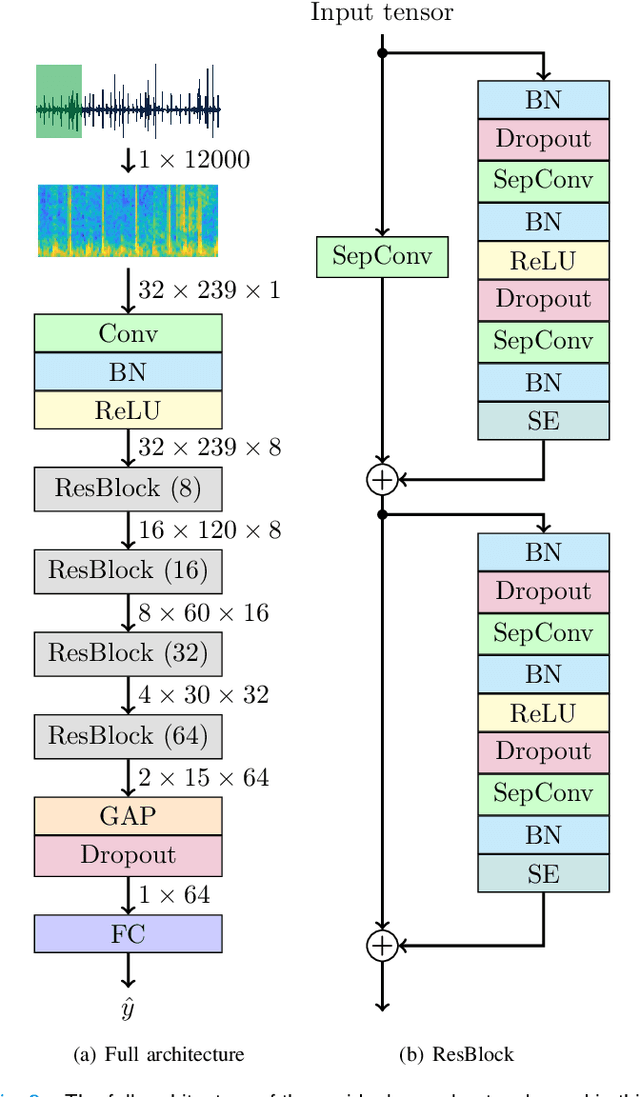
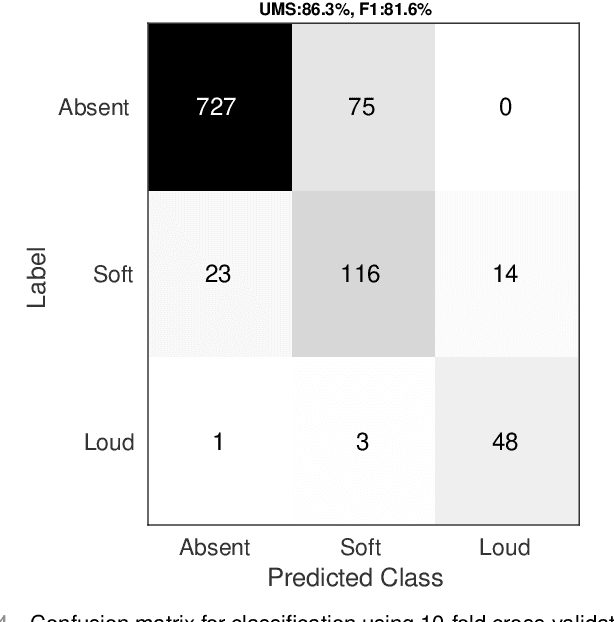
Abstract:Objective: Murmurs are abnormal heart sounds, identified by experts through cardiac auscultation. The murmur grade, a quantitative measure of the murmur intensity, is strongly correlated with the patient's clinical condition. This work aims to estimate each patient's murmur grade (i.e., absent, soft, loud) from multiple auscultation location phonocardiograms (PCGs) of a large population of pediatric patients from a low-resource rural area. Methods: The Mel spectrogram representation of each PCG recording is given to an ensemble of 15 convolutional residual neural networks with channel-wise attention mechanisms to classify each PCG recording. The final murmur grade for each patient is derived based on the proposed decision rule and considering all estimated labels for available recordings. The proposed method is cross-validated on a dataset consisting of 3456 PCG recordings from 1007 patients using a stratified ten-fold cross-validation. Additionally, the method was tested on a hidden test set comprised of 1538 PCG recordings from 442 patients. Results: The overall cross-validation performances for patient-level murmur gradings are 86.3% and 81.6% in terms of the unweighted average of sensitivities and F1-scores, respectively. The sensitivities (and F1-scores) for absent, soft, and loud murmurs are 90.7% (93.6%), 75.8% (66.8%), and 92.3% (84.2%), respectively. On the test set, the algorithm achieves an unweighted average of sensitivities of 80.4% and an F1-score of 75.8%. Conclusions: This study provides a potential approach for algorithmic pre-screening in low-resource settings with relatively high expert screening costs. Significance: The proposed method represents a significant step beyond detection of murmurs, providing characterization of intensity which may provide a enhanced classification of clinical outcomes.
Mythological Medical Machine Learning: Boosting the Performance of a Deep Learning Medical Data Classifier Using Realistic Physiological Models
Dec 28, 2021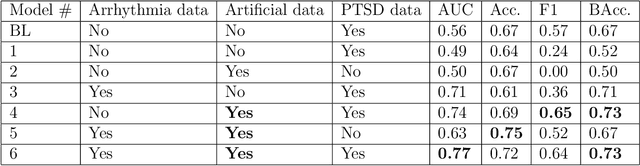
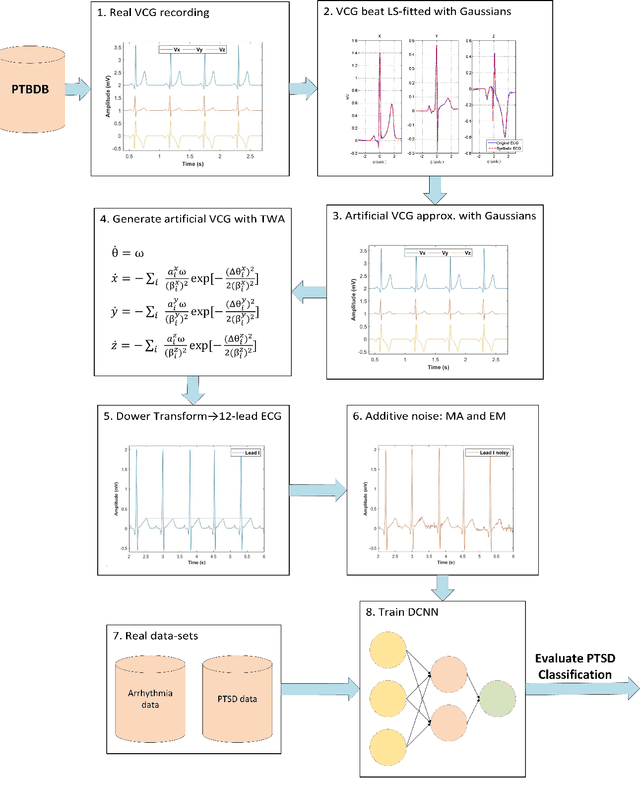
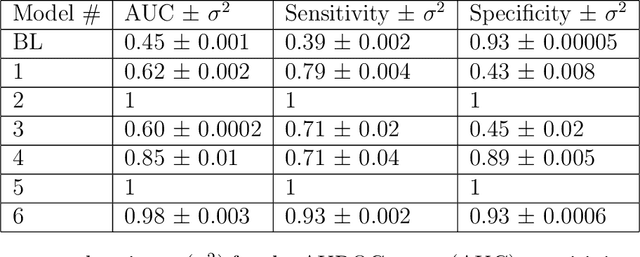
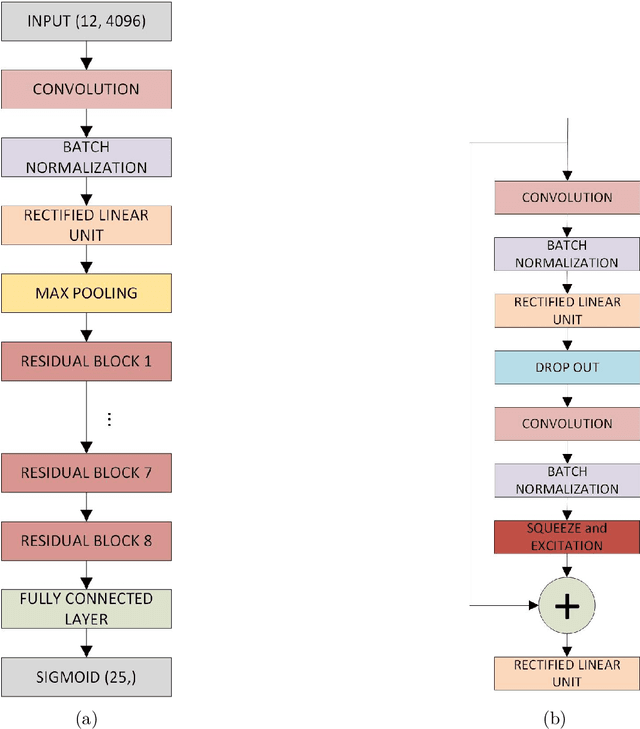
Abstract:Objective: To determine if a realistic, but computationally efficient model of the electrocardiogram can be used to pre-train a deep neural network (DNN) with a wide range of morphologies and abnormalities specific to a given condition - T-wave Alternans (TWA) as a result of Post-Traumatic Stress Disorder, or PTSD - and significantly boost performance on a small database of rare individuals. Approach: Using a previously validated artificial ECG model, we generated 180,000 artificial ECGs with or without significant TWA, with varying heart rate, breathing rate, TWA amplitude, and ECG morphology. A DNN, trained on over 70,000 patients to classify 25 different rhythms, was modified the output layer to a binary class (TWA or no-TWA, or equivalently, PTSD or no-PTSD), and transfer learning was performed on the artificial ECG. In a final transfer learning step, the DNN was trained and cross-validated on ECG from 12 PTSD and 24 controls for all combinations of using the three databases. Main results: The best performing approach (AUROC = 0.77, Accuracy = 0.72, F1-score = 0.64) was found by performing both transfer learning steps, using the pre-trained arrhythmia DNN, the artificial data and the real PTSD-related ECG data. Removing the artificial data from training led to the largest drop in performance. Removing the arrhythmia data from training provided a modest, but significant, drop in performance. The final model showed no significant drop in performance on the artificial data, indicating no overfitting. Significance: In healthcare, it is common to only have a small collection of high-quality data and labels, or a larger database with much lower quality (and less relevant) labels. The paradigm presented here, involving model-based performance boosting, provides a solution through transfer learning on a large realistic artificial database, and a partially relevant real database.
Privacy-Preserving Eye-tracking Using Deep Learning
Jun 22, 2021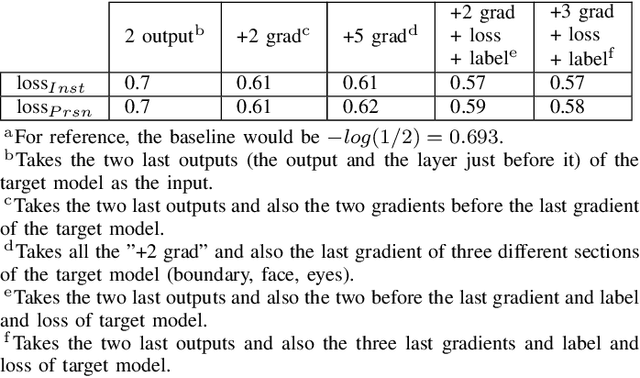
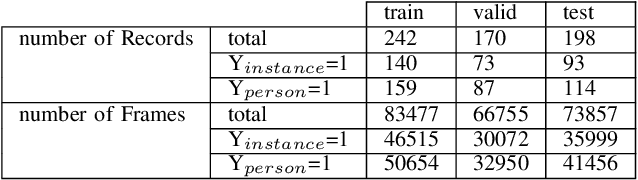

Abstract:The expanding usage of complex machine learning methods like deep learning has led to an explosion in human activity recognition, particularly applied to health. In particular, as part of a larger body sensor network system, face and full-body analysis is becoming increasingly common for evaluating health status. However, complex models which handle private and sometimes protected data, raise concerns about the potential leak of identifiable data. In this work, we focus on the case of a deep network model trained on images of individual faces. Full-face video recordings taken from 493 individuals undergoing an eye-tracking based evaluation of neurological function were used. Outputs, gradients, intermediate layer outputs, loss, and labels were used as inputs for a deep network with an added support vector machine emission layer to recognize membership in the training data. The inference attack method and associated mathematical analysis indicate that there is a low likelihood of unintended memorization of facial features in the deep learning model. In this study, it is showed that the named model preserves the integrity of training data with reasonable confidence. The same process can be implemented in similar conditions for different models.
Late fusion of machine learning models using passively captured interpersonal social interactions and motion from smartphones predicts decompensation in heart failure
Apr 04, 2021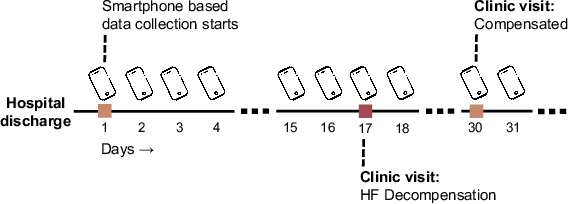
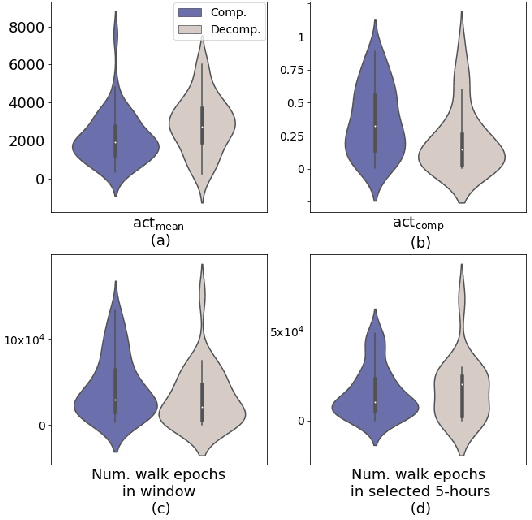
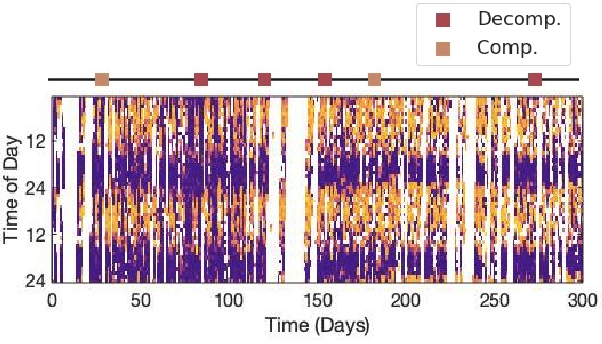
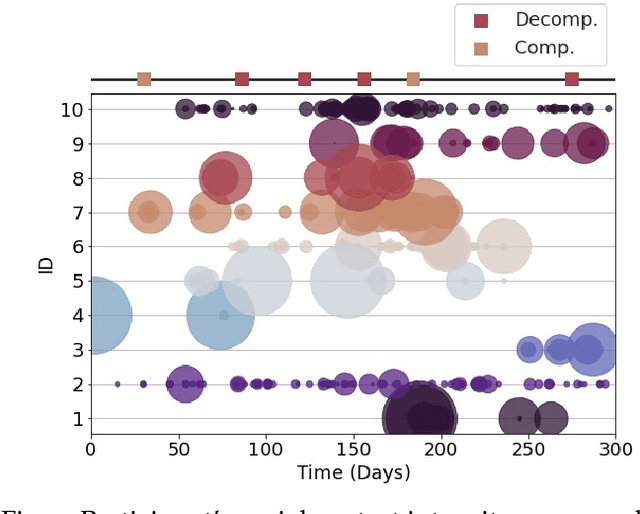
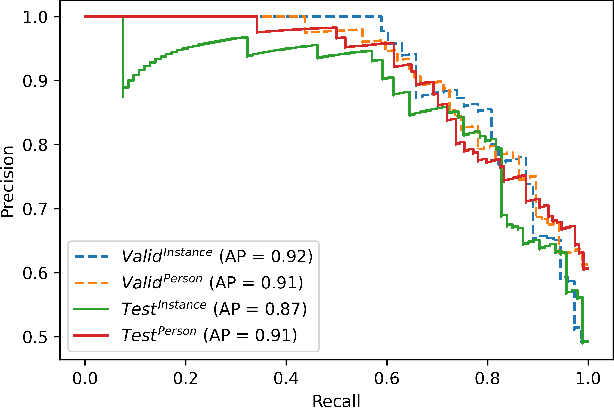
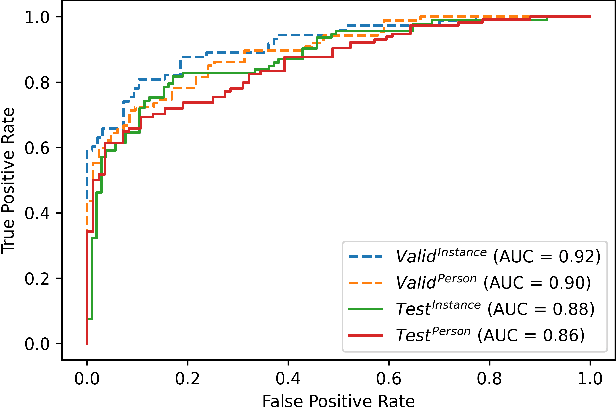
Abstract:Objective: Worldwide, heart failure (HF) is a major cause of morbidity and mortality and one of the leading causes of hospitalization. Early detection of HF symptoms and pro-active management may reduce adverse events. Approach: Twenty-eight participants were monitored using a smartphone app after discharge from hospitals, and each clinical event during the enrollment (N=110 clinical events) was recorded. Motion, social, location, and clinical survey data collected via the smartphone-based monitoring system were used to develop and validate an algorithm for predicting or classifying HF decompensation events (hospitalizations or clinic visit) versus clinic monitoring visits in which they were determined to be compensated or stable. Models based on single modality as well as early and late fusion approaches combining patient-reported outcomes and passive smartphone data were evaluated. Results: The highest AUCPr for classifying decompensation with a late fusion approach was 0.80 using leave one subject out cross-validation. Significance: Passively collected data from smartphones, especially when combined with weekly patient-reported outcomes, may reflect behavioral and physiological changes due to HF and thus could enable prediction of HF decompensation.
An Analysis Of Protected Health Information Leakage In Deep-Learning Based De-Identification Algorithms
Jan 28, 2021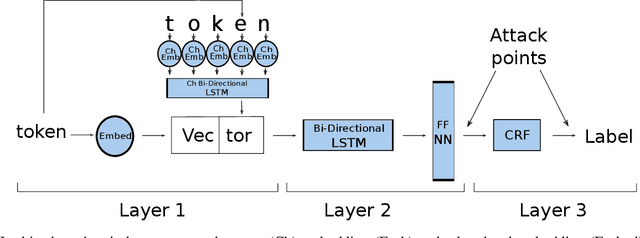
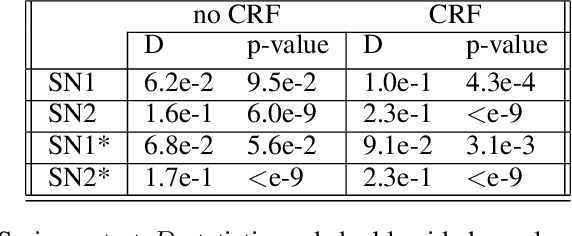
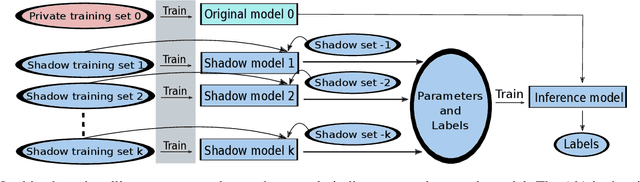
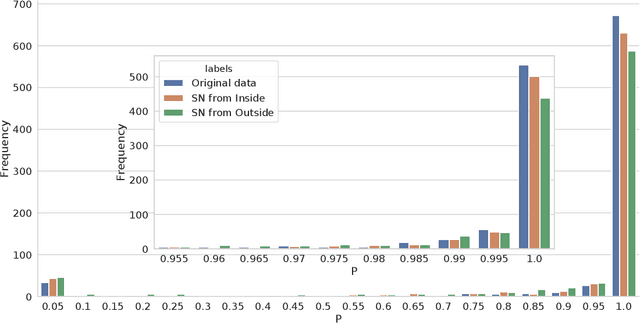
Abstract:The increasing complexity of algorithms for analyzing medical data, including de-identification tasks, raises the possibility that complex algorithms are learning not just the general representation of the problem, but specifics of given individuals within the data. Modern legal frameworks specifically prohibit the intentional or accidental distribution of patient data, but have not addressed this potential avenue for leakage of such protected health information. Modern deep learning algorithms have the highest potential of such leakage due to complexity of the models. Recent research in the field has highlighted such issues in non-medical data, but all analysis is likely to be data and algorithm specific. We, therefore, chose to analyze a state-of-the-art free-text de-identification algorithm based on LSTM (Long Short-Term Memory) and its potential in encoding any individual in the training set. Using the i2b2 Challenge Data, we trained, then analyzed the model to assess whether the output of the LSTM, before the compression layer of the classifier, could be used to estimate the membership of the training data. Furthermore, we used different attacks including membership inference attack method to attack the model. Results indicate that the attacks could not identify whether members of the training data were distinguishable from non-members based on the model output. This indicates that the model does not provide any strong evidence into the identification of the individuals in the training data set and there is not yet empirical evidence it is unsafe to distribute the model for general use.
Voting of predictive models for clinical outcomes: consensus of algorithms for the early prediction of sepsis from clinical data and an analysis of the PhysioNet/Computing in Cardiology Challenge 2019
Dec 20, 2020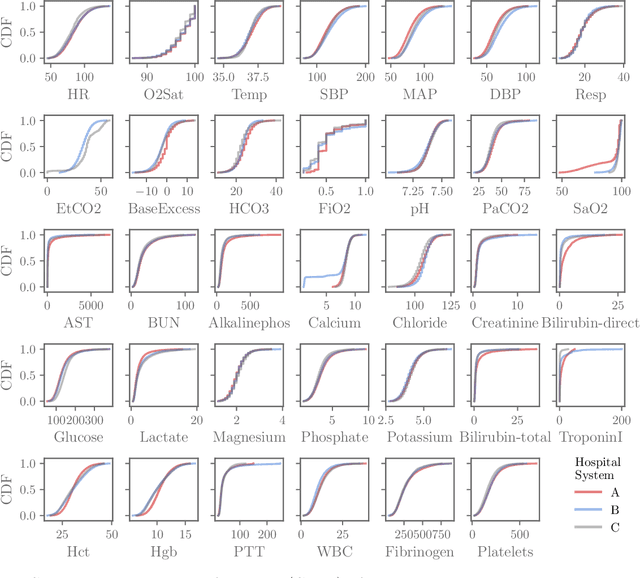
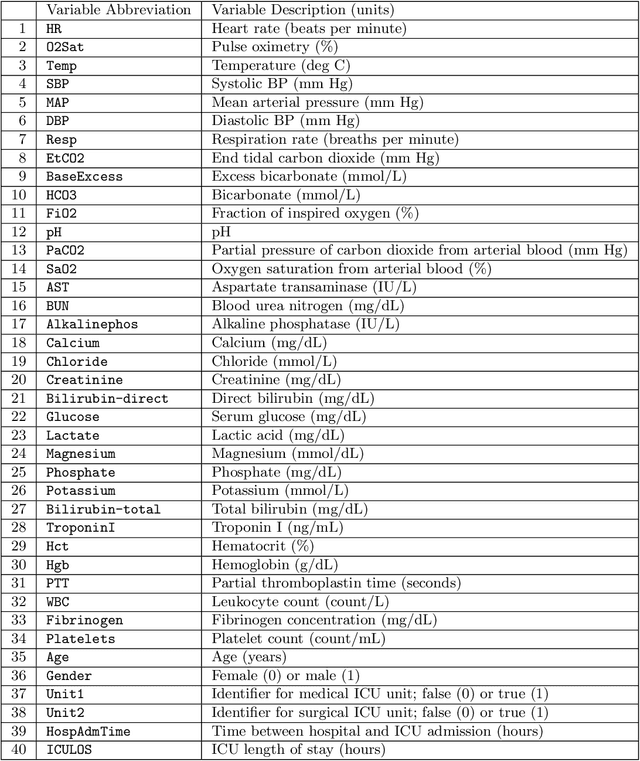
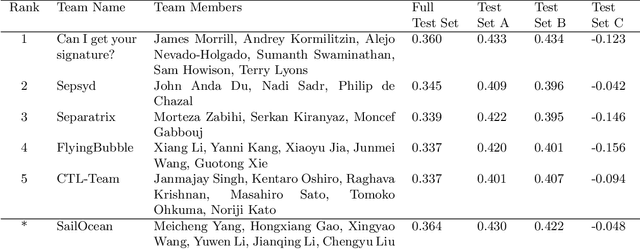
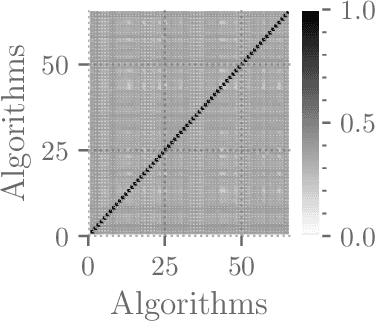
Abstract:Although there has been significant research in boosting of weak learners, there has been little work in the field of boosting from strong learners. This latter paradigm is a form of weighted voting with learned weights. In this work, we consider the problem of constructing an ensemble algorithm from 70 individual algorithms for the early prediction of sepsis from clinical data. We find that this ensemble algorithm outperforms separate algorithms, especially on a hidden test set on which most algorithms failed to generalize.
Using Convolutional Variational Autoencoders to Predict Post-Trauma Health Outcomes from Actigraphy Data
Nov 20, 2020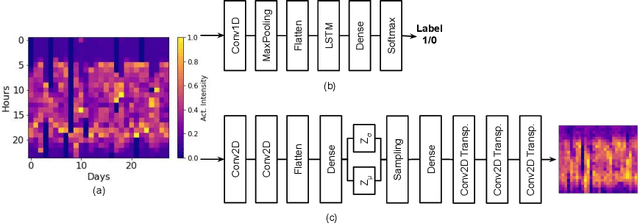
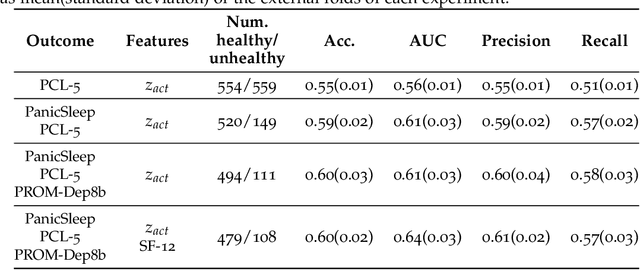
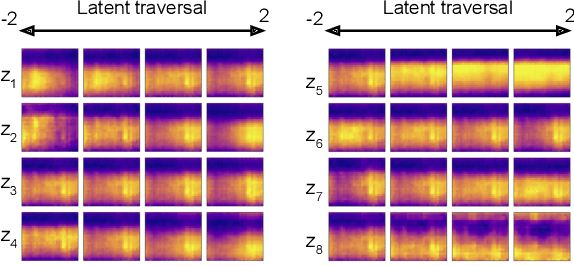
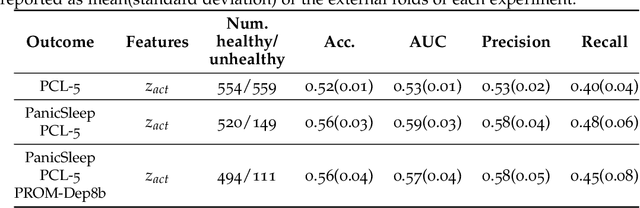
Abstract:Depression and post-traumatic stress disorder (PTSD) are psychiatric conditions commonly associated with experiencing a traumatic event. Estimating mental health status through non-invasive techniques such as activity-based algorithms can help to identify successful early interventions. In this work, we used locomotor activity captured from 1113 individuals who wore a research grade smartwatch post-trauma. A convolutional variational autoencoder (VAE) architecture was used for unsupervised feature extraction from four weeks of actigraphy data. By using VAE latent variables and the participant's pre-trauma physical health status as features, a logistic regression classifier achieved an area under the receiver operating characteristic curve (AUC) of 0.64 to estimate mental health outcomes. The results indicate that the VAE model is a promising approach for actigraphy data analysis for mental health outcomes in long-term studies.
Addressing Class Imbalance in Classification Problems of Noisy Signals by using Fourier Transform Surrogates
Jun 20, 2018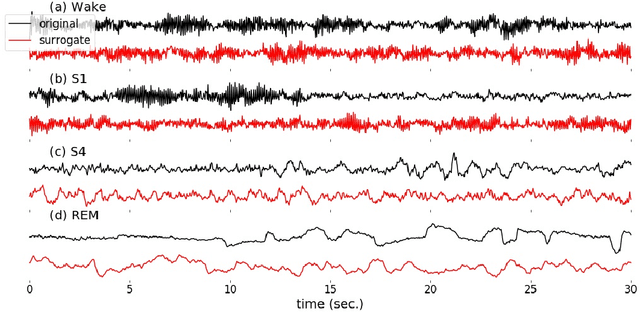
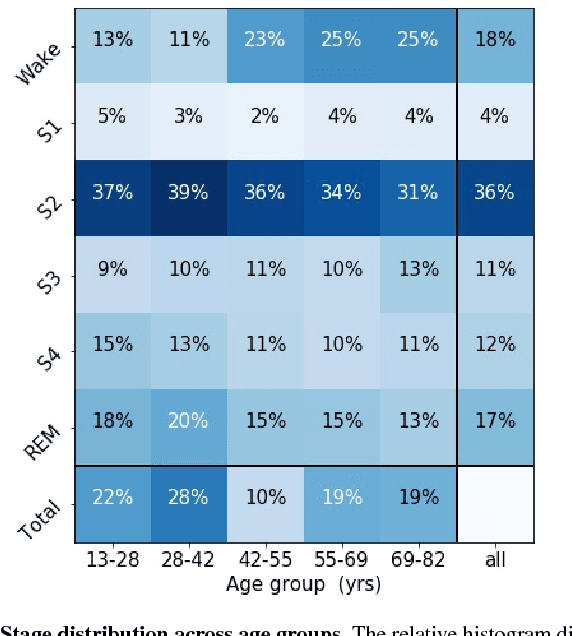
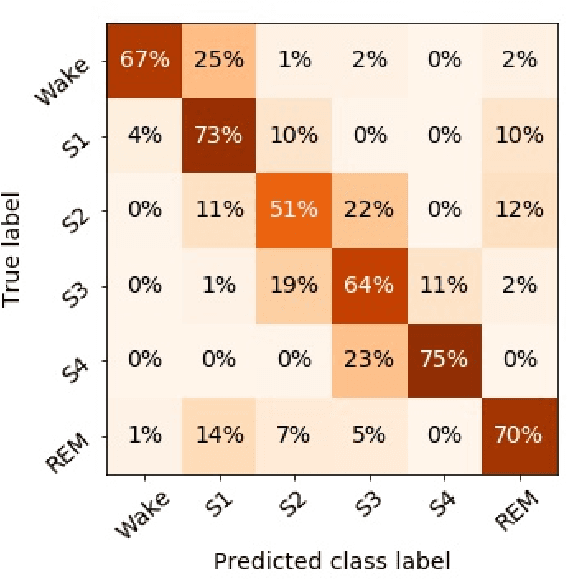
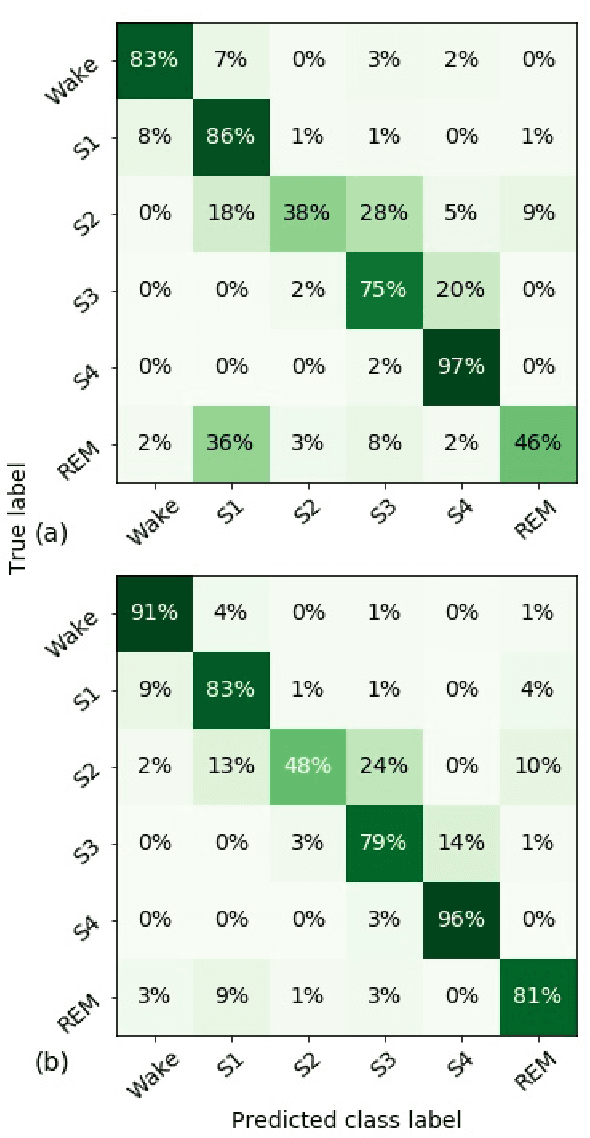
Abstract:Randomizing the Fourier-transform (FT) phases of temporal-spatial data generates surrogates that approximate examples from the data-generating distribution. We propose such FT surrogates as a novel tool to augment and analyze training of neural networks and explore the approach in the example of sleep-stage classification. By computing FT surrogates of raw EEG, EOG, and EMG signals of under-represented sleep stages, we balanced the CAPSLPDB sleep database. We then trained and tested a convolutional neural network for sleep stage classification, and found that our surrogate-based augmentation improved the mean F1-score by 7%. As another application of FT surrogates, we formulated an approach to compute saliency maps for individual sleep epochs. The visualization is based on the response of inferred class probabilities under replacement of short data segments by partial surrogates. To quantify how well the distributions of the surrogates and the original data match, we evaluated a trained classifier on surrogates of correctly classified examples, and summarized these conditional predictions in a confusion matrix. We show how such conditional confusion matrices can qualitatively explain the performance of surrogates in class balancing. The FT-surrogate augmentation approach may improve classification on noisy signals if carefully adapted to the data distribution under analysis.
Detection of Paroxysmal Atrial Fibrillation using Attention-based Bidirectional Recurrent Neural Networks
May 07, 2018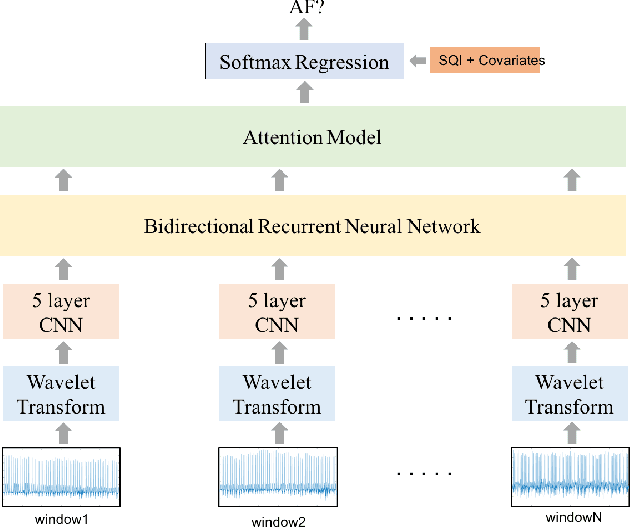
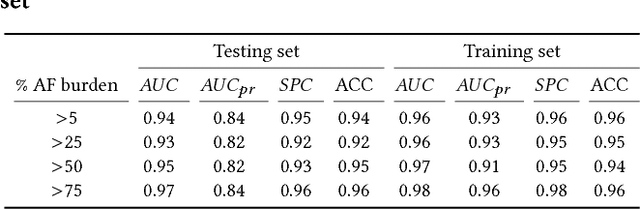
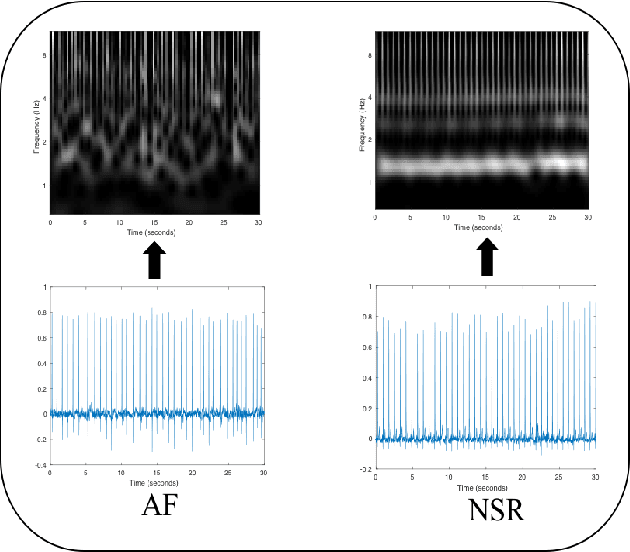
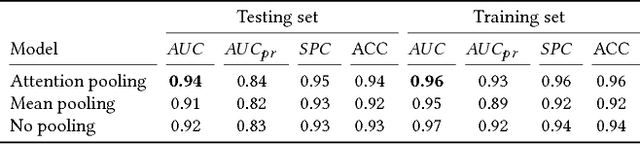
Abstract:Detection of atrial fibrillation (AF), a type of cardiac arrhythmia, is difficult since many cases of AF are usually clinically silent and undiagnosed. In particular paroxysmal AF is a form of AF that occurs occasionally, and has a higher probability of being undetected. In this work, we present an attention based deep learning framework for detection of paroxysmal AF episodes from a sequence of windows. Time-frequency representation of 30 seconds recording windows, over a 10 minute data segment, are fed sequentially into a deep convolutional neural network for image-based feature extraction, which are then presented to a bidirectional recurrent neural network with an attention layer for AF detection. To demonstrate the effectiveness of the proposed framework for transient AF detection, we use a database of 24 hour Holter Electrocardiogram (ECG) recordings acquired from 2850 patients at the University of Virginia heart station. The algorithm achieves an AUC of 0.94 on the testing set, which exceeds the performance of baseline models. We also demonstrate the cross-domain generalizablity of the approach by adapting the learned model parameters from one recording modality (ECG) to another (photoplethysmogram) with improved AF detection performance. The proposed high accuracy, low false alarm algorithm for detecting paroxysmal AF has potential applications in long-term monitoring using wearable sensors.
Fusing Continuous-valued Medical Labels using a Bayesian Model
Jun 13, 2015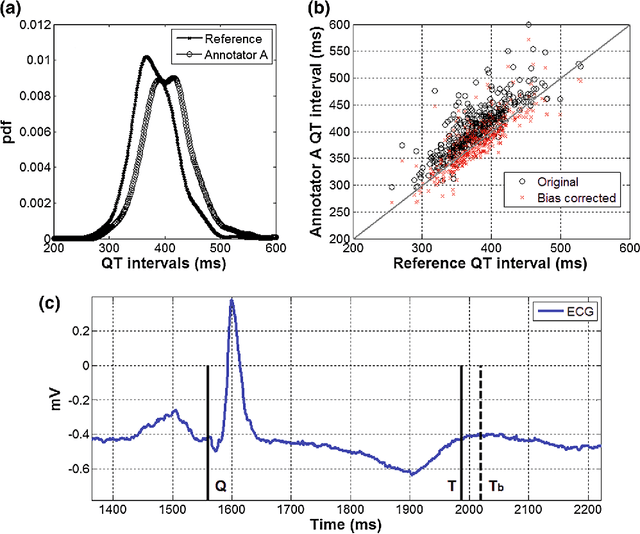
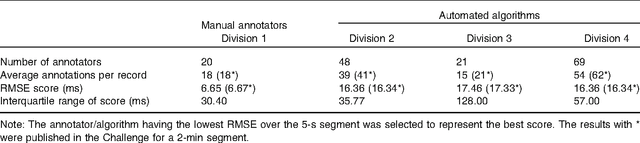
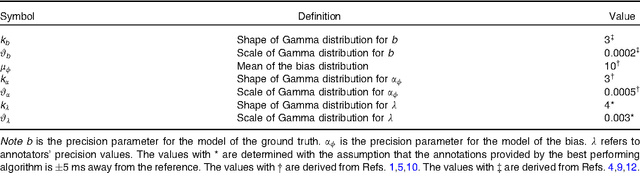

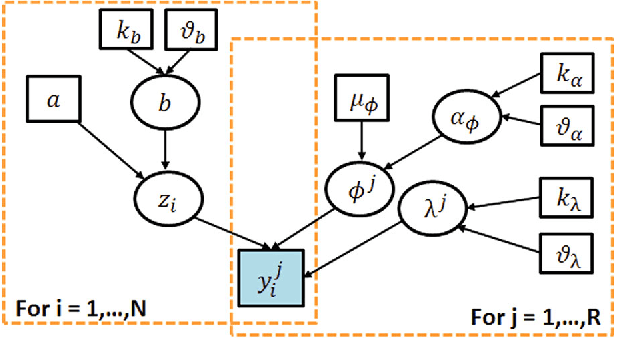

Abstract:With the rapid increase in volume of time series medical data available through wearable devices, there is a need to employ automated algorithms to label data. Examples of labels include interventions, changes in activity (e.g. sleep) and changes in physiology (e.g. arrhythmias). However, automated algorithms tend to be unreliable resulting in lower quality care. Expert annotations are scarce, expensive, and prone to significant inter- and intra-observer variance. To address these problems, a Bayesian Continuous-valued Label Aggregator(BCLA) is proposed to provide a reliable estimation of label aggregation while accurately infer the precision and bias of each algorithm. The BCLA was applied to QT interval (pro-arrhythmic indicator) estimation from the electrocardiogram using labels from the 2006 PhysioNet/Computing in Cardiology Challenge database. It was compared to the mean, median, and a previously proposed Expectation Maximization (EM) label aggregation approaches. While accurately predicting each labelling algorithm's bias and precision, the root-mean-square error of the BCLA was 11.78$\pm$0.63ms, significantly outperforming the best Challenge entry (15.37$\pm$2.13ms) as well as the EM, mean, and median voting strategies (14.76$\pm$0.52ms, 17.61$\pm$0.55ms, and 14.43$\pm$0.57ms respectively with $p<0.0001$).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge