Fangxu Xing
MomentaMorph: Unsupervised Spatial-Temporal Registration with Momenta, Shooting, and Correction
Aug 05, 2023



Abstract:Tagged magnetic resonance imaging (tMRI) has been employed for decades to measure the motion of tissue undergoing deformation. However, registration-based motion estimation from tMRI is difficult due to the periodic patterns in these images, particularly when the motion is large. With a larger motion the registration approach gets trapped in a local optima, leading to motion estimation errors. We introduce a novel "momenta, shooting, and correction" framework for Lagrangian motion estimation in the presence of repetitive patterns and large motion. This framework, grounded in Lie algebra and Lie group principles, accumulates momenta in the tangent vector space and employs exponential mapping in the diffeomorphic space for rapid approximation towards true optima, circumventing local optima. A subsequent correction step ensures convergence to true optima. The results on a 2D synthetic dataset and a real 3D tMRI dataset demonstrate our method's efficiency in estimating accurate, dense, and diffeomorphic 2D/3D motion fields amidst large motion and repetitive patterns.
Incremental Learning for Heterogeneous Structure Segmentation in Brain Tumor MRI
May 30, 2023



Abstract:Deep learning (DL) models for segmenting various anatomical structures have achieved great success via a static DL model that is trained in a single source domain. Yet, the static DL model is likely to perform poorly in a continually evolving environment, requiring appropriate model updates. In an incremental learning setting, we would expect that well-trained static models are updated, following continually evolving target domain data -- e.g., additional lesions or structures of interest -- collected from different sites, without catastrophic forgetting. This, however, poses challenges, due to distribution shifts, additional structures not seen during the initial model training, and the absence of training data in a source domain. To address these challenges, in this work, we seek to progressively evolve an ``off-the-shelf" trained segmentation model to diverse datasets with additional anatomical categories in a unified manner. Specifically, we first propose a divergence-aware dual-flow module with balanced rigidity and plasticity branches to decouple old and new tasks, which is guided by continuous batch renormalization. Then, a complementary pseudo-label training scheme with self-entropy regularized momentum MixUp decay is developed for adaptive network optimization. We evaluated our framework on a brain tumor segmentation task with continually changing target domains -- i.e., new MRI scanners/modalities with incremental structures. Our framework was able to well retain the discriminability of previously learned structures, hence enabling the realistic life-long segmentation model extension along with the widespread accumulation of big medical data.
Attentive Continuous Generative Self-training for Unsupervised Domain Adaptive Medical Image Translation
May 23, 2023



Abstract:Self-training is an important class of unsupervised domain adaptation (UDA) approaches that are used to mitigate the problem of domain shift, when applying knowledge learned from a labeled source domain to unlabeled and heterogeneous target domains. While self-training-based UDA has shown considerable promise on discriminative tasks, including classification and segmentation, through reliable pseudo-label filtering based on the maximum softmax probability, there is a paucity of prior work on self-training-based UDA for generative tasks, including image modality translation. To fill this gap, in this work, we seek to develop a generative self-training (GST) framework for domain adaptive image translation with continuous value prediction and regression objectives. Specifically, we quantify both aleatoric and epistemic uncertainties within our GST using variational Bayes learning to measure the reliability of synthesized data. We also introduce a self-attention scheme that de-emphasizes the background region to prevent it from dominating the training process. The adaptation is then carried out by an alternating optimization scheme with target domain supervision that focuses attention on the regions with reliable pseudo-labels. We evaluated our framework on two cross-scanner/center, inter-subject translation tasks, including tagged-to-cine magnetic resonance (MR) image translation and T1-weighted MR-to-fractional anisotropy translation. Extensive validations with unpaired target domain data showed that our GST yielded superior synthesis performance in comparison to adversarial training UDA methods.
Synthesizing audio from tongue motion during speech using tagged MRI via transformer
Feb 14, 2023Abstract:Investigating the relationship between internal tissue point motion of the tongue and oropharyngeal muscle deformation measured from tagged MRI and intelligible speech can aid in advancing speech motor control theories and developing novel treatment methods for speech related-disorders. However, elucidating the relationship between these two sources of information is challenging, due in part to the disparity in data structure between spatiotemporal motion fields (i.e., 4D motion fields) and one-dimensional audio waveforms. In this work, we present an efficient encoder-decoder translation network for exploring the predictive information inherent in 4D motion fields via 2D spectrograms as a surrogate of the audio data. Specifically, our encoder is based on 3D convolutional spatial modeling and transformer-based temporal modeling. The extracted features are processed by an asymmetric 2D convolution decoder to generate spectrograms that correspond to 4D motion fields. Furthermore, we incorporate a generative adversarial training approach into our framework to further improve synthesis quality on our generated spectrograms. We experiment on 63 paired motion field sequences and speech waveforms, demonstrating that our framework enables the generation of clear audio waveforms from a sequence of motion fields. Thus, our framework has the potential to improve our understanding of the relationship between these two modalities and inform the development of treatments for speech disorders.
Successive Subspace Learning for Cardiac Disease Classification with Two-phase Deformation Fields from Cine MRI
Jan 21, 2023



Abstract:Cardiac cine magnetic resonance imaging (MRI) has been used to characterize cardiovascular diseases (CVD), often providing a noninvasive phenotyping tool.~While recently flourished deep learning based approaches using cine MRI yield accurate characterization results, the performance is often degraded by small training samples. In addition, many deep learning models are deemed a ``black box," for which models remain largely elusive in how models yield a prediction and how reliable they are. To alleviate this, this work proposes a lightweight successive subspace learning (SSL) framework for CVD classification, based on an interpretable feedforward design, in conjunction with a cardiac atlas. Specifically, our hierarchical SSL model is based on (i) neighborhood voxel expansion, (ii) unsupervised subspace approximation, (iii) supervised regression, and (iv) multi-level feature integration. In addition, using two-phase 3D deformation fields, including end-diastolic and end-systolic phases, derived between the atlas and individual subjects as input offers objective means of assessing CVD, even with small training samples. We evaluate our framework on the ACDC2017 database, comprising one healthy group and four disease groups. Compared with 3D CNN-based approaches, our framework achieves superior classification performance with 140$\times$ fewer parameters, which supports its potential value in clinical use.
Deep Unsupervised Phase-based 3D Incompressible Motion Estimation in Tagged-MRI
Jan 18, 2023



Abstract:Tagged magnetic resonance imaging (MRI) has been used for decades to observe and quantify the detailed motion of deforming tissue. However, this technique faces several challenges such as tag fading, large motion, long computation times, and difficulties in obtaining diffeomorphic incompressible flow fields. To address these issues, this paper presents a novel unsupervised phase-based 3D motion estimation technique for tagged MRI. We introduce two key innovations. First, we apply a sinusoidal transformation to the harmonic phase input, which enables end-to-end training and avoids the need for phase interpolation. Second, we propose a Jacobian determinant-based learning objective to encourage incompressible flow fields for deforming biological tissues. Our method efficiently estimates 3D motion fields that are accurate, dense, and approximately diffeomorphic and incompressible. The efficacy of the method is assessed using human tongue motion during speech, and includes both healthy controls and patients that have undergone glossectomy. We show that the method outperforms existing approaches, and also exhibits improvements in speed, robustness to tag fading, and large tongue motion.
Memory Consistent Unsupervised Off-the-Shelf Model Adaptation for Source-Relaxed Medical Image Segmentation
Sep 16, 2022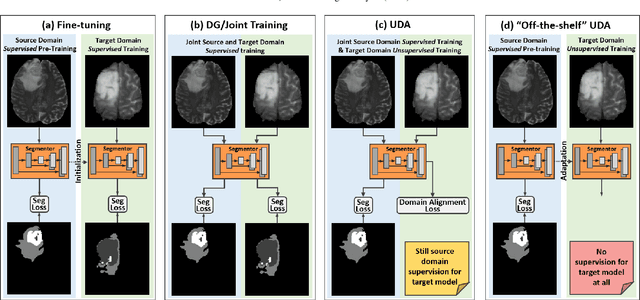
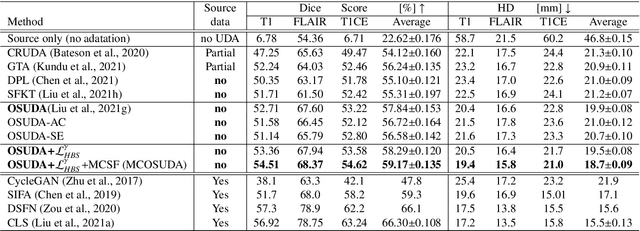
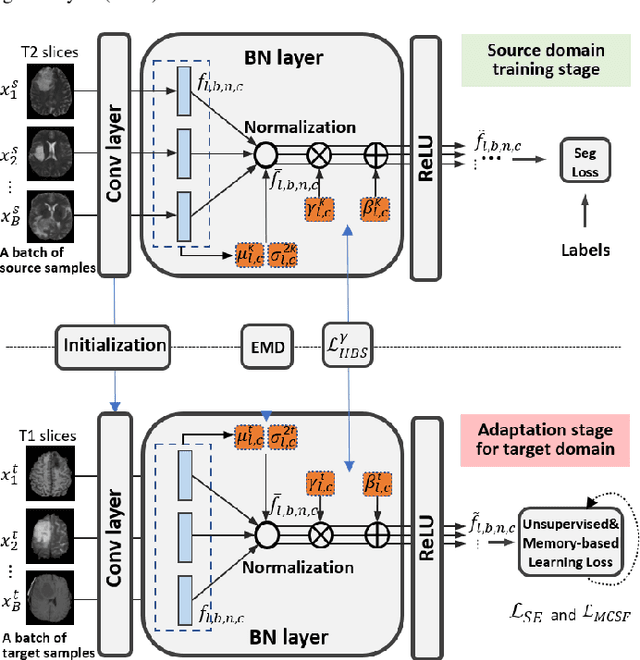
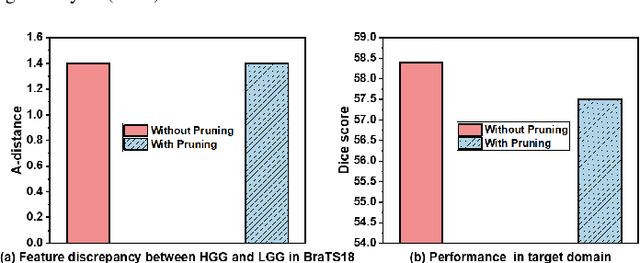
Abstract:Unsupervised domain adaptation (UDA) has been a vital protocol for migrating information learned from a labeled source domain to facilitate the implementation in an unlabeled heterogeneous target domain. Although UDA is typically jointly trained on data from both domains, accessing the labeled source domain data is often restricted, due to concerns over patient data privacy or intellectual property. To sidestep this, we propose "off-the-shelf (OS)" UDA (OSUDA), aimed at image segmentation, by adapting an OS segmentor trained in a source domain to a target domain, in the absence of source domain data in adaptation. Toward this goal, we aim to develop a novel batch-wise normalization (BN) statistics adaptation framework. In particular, we gradually adapt the domain-specific low-order BN statistics, e.g., mean and variance, through an exponential momentum decay strategy, while explicitly enforcing the consistency of the domain shareable high-order BN statistics, e.g., scaling and shifting factors, via our optimization objective. We also adaptively quantify the channel-wise transferability to gauge the importance of each channel, via both low-order statistics divergence and a scaling factor.~Furthermore, we incorporate unsupervised self-entropy minimization into our framework to boost performance alongside a novel queued, memory-consistent self-training strategy to utilize the reliable pseudo label for stable and efficient unsupervised adaptation. We evaluated our OSUDA-based framework on both cross-modality and cross-subtype brain tumor segmentation and cardiac MR to CT segmentation tasks. Our experimental results showed that our memory consistent OSUDA performs better than existing source-relaxed UDA methods and yields similar performance to UDA methods with source data.
Unsupervised Domain Adaptation for Segmentation with Black-box Source Model
Aug 16, 2022Abstract:Unsupervised domain adaptation (UDA) has been widely used to transfer knowledge from a labeled source domain to an unlabeled target domain to counter the difficulty of labeling in a new domain. The training of conventional solutions usually relies on the existence of both source and target domain data. However, privacy of the large-scale and well-labeled data in the source domain and trained model parameters can become the major concern of cross center/domain collaborations. In this work, to address this, we propose a practical solution to UDA for segmentation with a black-box segmentation model trained in the source domain only, rather than original source data or a white-box source model. Specifically, we resort to a knowledge distillation scheme with exponential mixup decay (EMD) to gradually learn target-specific representations. In addition, unsupervised entropy minimization is further applied to regularization of the target domain confidence. We evaluated our framework on the BraTS 2018 database, achieving performance on par with white-box source model adaptation approaches.
Subtype-Aware Dynamic Unsupervised Domain Adaptation
Aug 16, 2022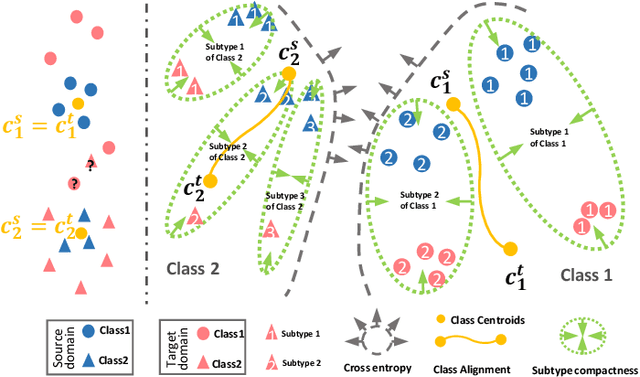
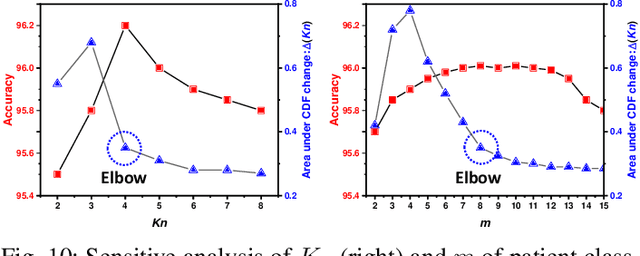
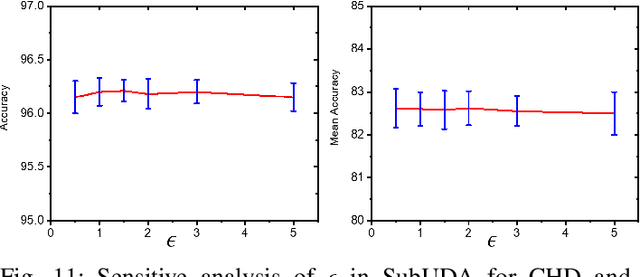
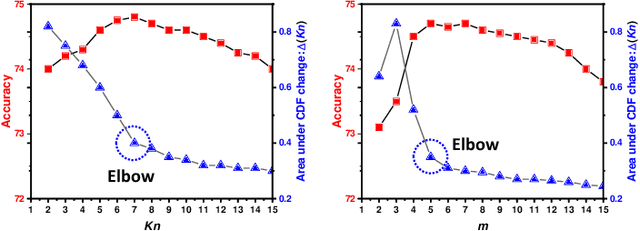
Abstract:Unsupervised domain adaptation (UDA) has been successfully applied to transfer knowledge from a labeled source domain to target domains without their labels. Recently introduced transferable prototypical networks (TPN) further addresses class-wise conditional alignment. In TPN, while the closeness of class centers between source and target domains is explicitly enforced in a latent space, the underlying fine-grained subtype structure and the cross-domain within-class compactness have not been fully investigated. To counter this, we propose a new approach to adaptively perform a fine-grained subtype-aware alignment to improve performance in the target domain without the subtype label in both domains. The insight of our approach is that the unlabeled subtypes in a class have the local proximity within a subtype, while exhibiting disparate characteristics, because of different conditional and label shifts. Specifically, we propose to simultaneously enforce subtype-wise compactness and class-wise separation, by utilizing intermediate pseudo-labels. In addition, we systematically investigate various scenarios with and without prior knowledge of subtype numbers, and propose to exploit the underlying subtype structure. Furthermore, a dynamic queue framework is developed to evolve the subtype cluster centroids steadily using an alternative processing scheme. Experimental results, carried out with multi-view congenital heart disease data and VisDA and DomainNet, show the effectiveness and validity of our subtype-aware UDA, compared with state-of-the-art UDA methods.
Deep Unsupervised Domain Adaptation: A Review of Recent Advances and Perspectives
Aug 15, 2022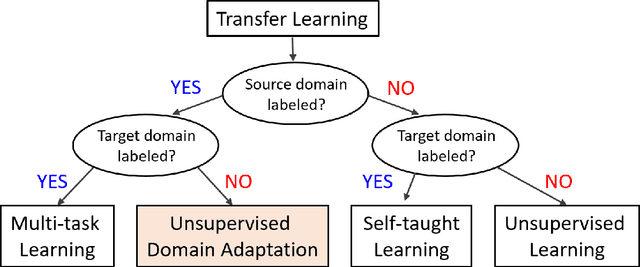

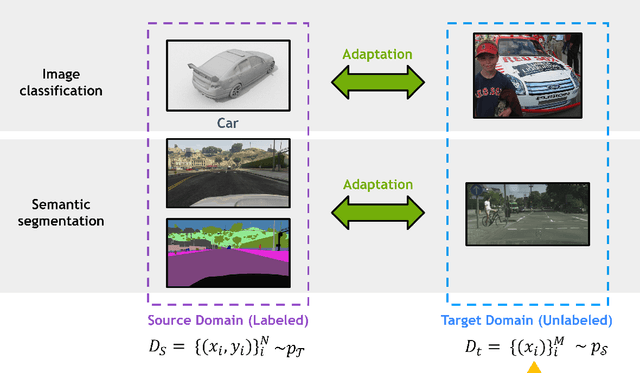
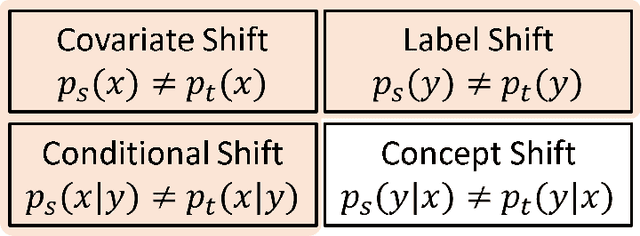
Abstract:Deep learning has become the method of choice to tackle real-world problems in different domains, partly because of its ability to learn from data and achieve impressive performance on a wide range of applications. However, its success usually relies on two assumptions: (i) vast troves of labeled datasets are required for accurate model fitting, and (ii) training and testing data are independent and identically distributed. Its performance on unseen target domains, thus, is not guaranteed, especially when encountering out-of-distribution data at the adaptation stage. The performance drop on data in a target domain is a critical problem in deploying deep neural networks that are successfully trained on data in a source domain. Unsupervised domain adaptation (UDA) is proposed to counter this, by leveraging both labeled source domain data and unlabeled target domain data to carry out various tasks in the target domain. UDA has yielded promising results on natural image processing, video analysis, natural language processing, time-series data analysis, medical image analysis, etc. In this review, as a rapidly evolving topic, we provide a systematic comparison of its methods and applications. In addition, the connection of UDA with its closely related tasks, e.g., domain generalization and out-of-distribution detection, has also been discussed. Furthermore, deficiencies in current methods and possible promising directions are highlighted.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge