Eva L. Dyer
The good, the bad and the ugly sides of data augmentation: An implicit spectral regularization perspective
Oct 10, 2022
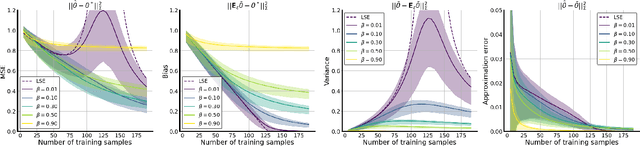
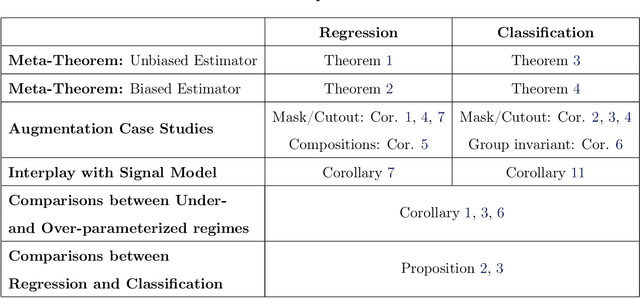
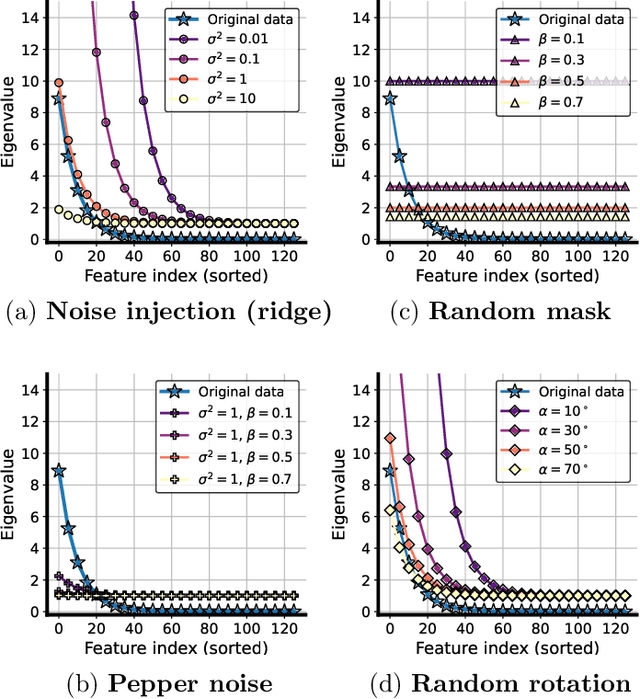
Abstract:Data augmentation (DA) is a powerful workhorse for bolstering performance in modern machine learning. Specific augmentations like translations and scaling in computer vision are traditionally believed to improve generalization by generating new (artificial) data from the same distribution. However, this traditional viewpoint does not explain the success of prevalent augmentations in modern machine learning (e.g. randomized masking, cutout, mixup), that greatly alter the training data distribution. In this work, we develop a new theoretical framework to characterize the impact of a general class of DA on underparameterized and overparameterized linear model generalization. Our framework reveals that DA induces implicit spectral regularization through a combination of two distinct effects: a) manipulating the relative proportion of eigenvalues of the data covariance matrix in a training-data-dependent manner, and b) uniformly boosting the entire spectrum of the data covariance matrix through ridge regression. These effects, when applied to popular augmentations, give rise to a wide variety of phenomena, including discrepancies in generalization between over-parameterized and under-parameterized regimes and differences between regression and classification tasks. Our framework highlights the nuanced and sometimes surprising impacts of DA on generalization, and serves as a testbed for novel augmentation design.
Learning Behavior Representations Through Multi-Timescale Bootstrapping
Jun 14, 2022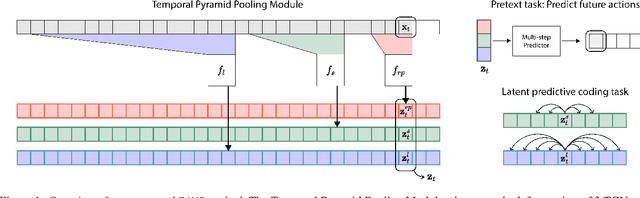
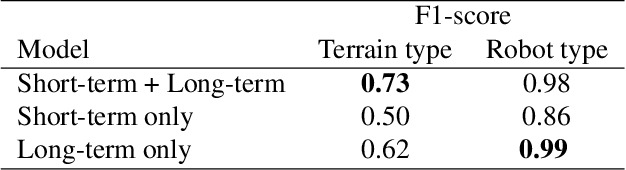
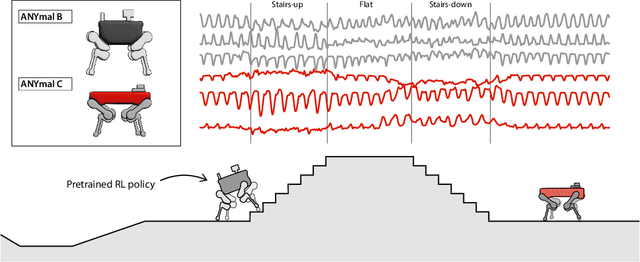
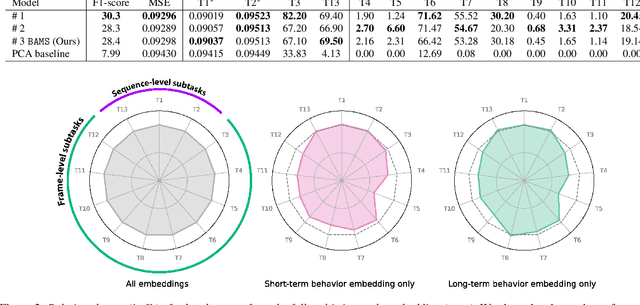
Abstract:Natural behavior consists of dynamics that are both unpredictable, can switch suddenly, and unfold over many different timescales. While some success has been found in building representations of behavior under constrained or simplified task-based conditions, many of these models cannot be applied to free and naturalistic settings due to the fact that they assume a single scale of temporal dynamics. In this work, we introduce Bootstrap Across Multiple Scales (BAMS), a multi-scale representation learning model for behavior: we combine a pooling module that aggregates features extracted over encoders with different temporal receptive fields, and design a set of latent objectives to bootstrap the representations in each respective space to encourage disentanglement across different timescales. We first apply our method on a dataset of quadrupeds navigating in different terrain types, and show that our model captures the temporal complexity of behavior. We then apply our method to the MABe 2022 Multi-agent behavior challenge, where our model ranks 3rd overall and 1st on two subtasks, and show the importance of incorporating multi-timescales when analyzing behavior.
Seeing the forest and the tree: Building representations of both individual and collective dynamics with transformers
Jun 10, 2022Abstract:Complex time-varying systems are often studied by abstracting away from the dynamics of individual components to build a model of the population-level dynamics from the start. However, when building a population-level description, it can be easy to lose sight of each individual and how each contributes to the larger picture. In this paper, we present a novel transformer architecture for learning from time-varying data that builds descriptions of both the individual as well as the collective population dynamics. Rather than combining all of our data into our model at the onset, we develop a separable architecture that operates on individual time-series first before passing them forward; this induces a permutation-invariance property and can be used to transfer across systems of different size and order. After demonstrating that our model can be applied to successfully recover complex interactions and dynamics in many-body systems, we apply our approach to populations of neurons in the nervous system. On neural activity datasets, we show that our multi-scale transformer not only yields robust decoding performance, but also provides impressive performance in transfer. Our results show that it is possible to learn from neurons in one animal's brain and transfer the model on neurons in a different animal's brain, with interpretable neuron correspondence across sets and animals. This finding opens up a new path to decode from and represent large collections of neurons.
Learning Sinkhorn divergences for supervised change point detection
Feb 10, 2022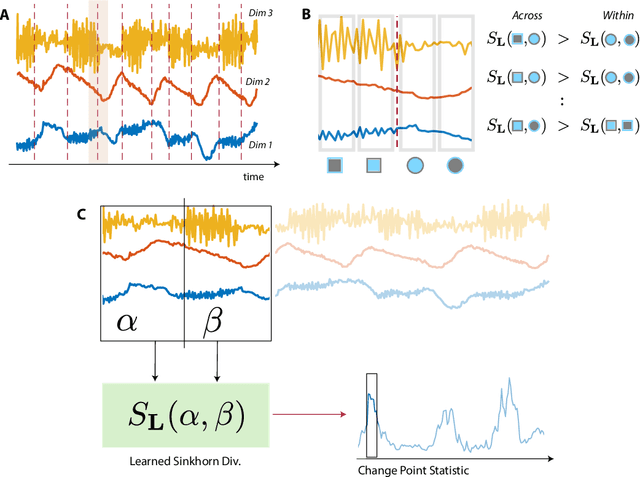
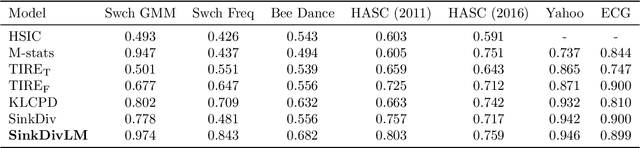
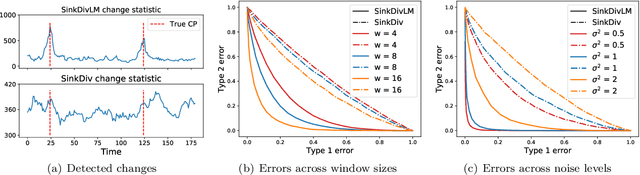
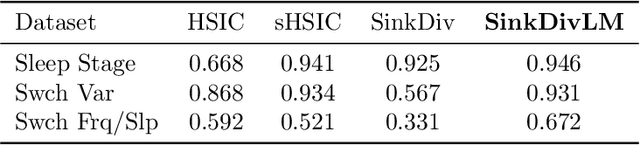
Abstract:Many modern applications require detecting change points in complex sequential data. Most existing methods for change point detection are unsupervised and, as a consequence, lack any information regarding what kind of changes we want to detect or if some kinds of changes are safe to ignore. This often results in poor change detection performance. We present a novel change point detection framework that uses true change point instances as supervision for learning a ground metric such that Sinkhorn divergences can be then used in two-sample tests on sliding windows to detect change points in an online manner. Our method can be used to learn a sparse metric which can be useful for both feature selection and interpretation in high-dimensional change point detection settings. Experiments on simulated as well as real world sequences show that our proposed method can substantially improve change point detection performance over existing unsupervised change point detection methods using only few labeled change point instances.
Drop, Swap, and Generate: A Self-Supervised Approach for Generating Neural Activity
Nov 03, 2021Abstract:Meaningful and simplified representations of neural activity can yield insights into how and what information is being processed within a neural circuit. However, without labels, finding representations that reveal the link between the brain and behavior can be challenging. Here, we introduce a novel unsupervised approach for learning disentangled representations of neural activity called Swap-VAE. Our approach combines a generative modeling framework with an instance-specific alignment loss that tries to maximize the representational similarity between transformed views of the input (brain state). These transformed (or augmented) views are created by dropping out neurons and jittering samples in time, which intuitively should lead the network to a representation that maintains both temporal consistency and invariance to the specific neurons used to represent the neural state. Through evaluations on both synthetic data and neural recordings from hundreds of neurons in different primate brains, we show that it is possible to build representations that disentangle neural datasets along relevant latent dimensions linked to behavior.
Neural Latents Benchmark '21: Evaluating latent variable models of neural population activity
Sep 10, 2021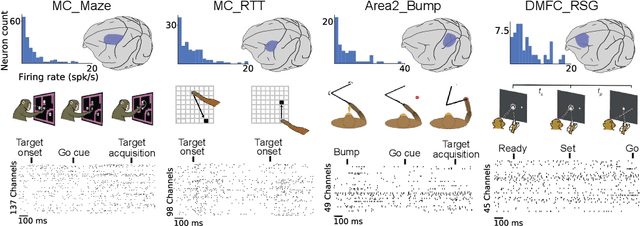
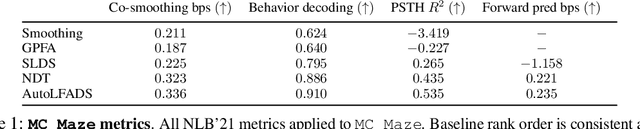

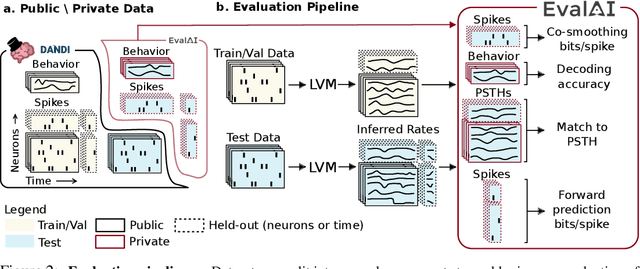
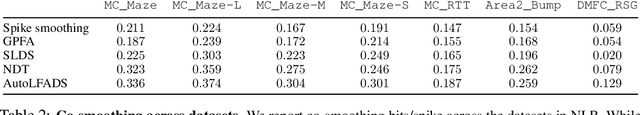
Abstract:Advances in neural recording present increasing opportunities to study neural activity in unprecedented detail. Latent variable models (LVMs) are promising tools for analyzing this rich activity across diverse neural systems and behaviors, as LVMs do not depend on known relationships between the activity and external experimental variables. However, progress in latent variable modeling is currently impeded by a lack of standardization, resulting in methods being developed and compared in an ad hoc manner. To coordinate these modeling efforts, we introduce a benchmark suite for latent variable modeling of neural population activity. We curate four datasets of neural spiking activity from cognitive, sensory, and motor areas to promote models that apply to the wide variety of activity seen across these areas. We identify unsupervised evaluation as a common framework for evaluating models across datasets, and apply several baselines that demonstrate benchmark diversity. We release this benchmark through EvalAI. http://neurallatents.github.io
Mine Your Own vieW: Self-Supervised Learning Through Across-Sample Prediction
Feb 19, 2021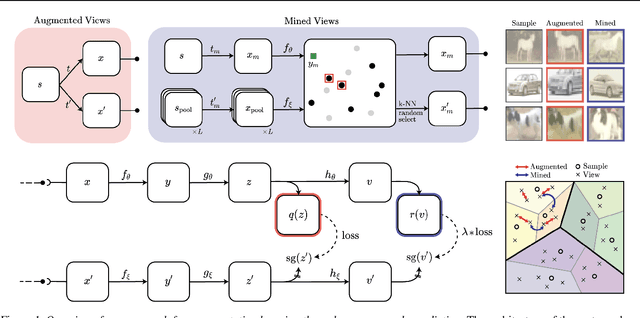
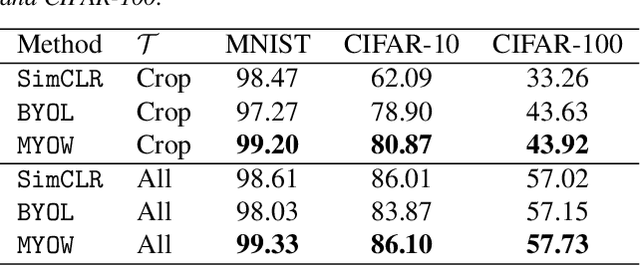
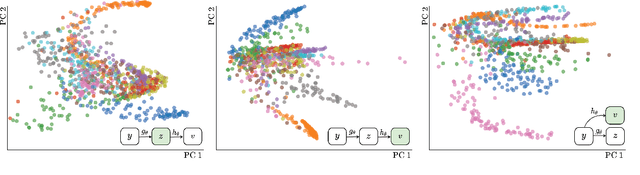
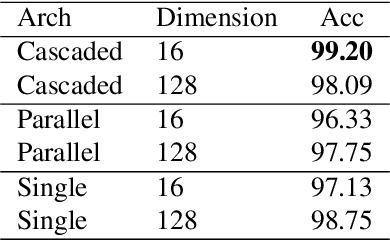
Abstract:State-of-the-art methods for self-supervised learning (SSL) build representations by maximizing the similarity between different augmented "views" of a sample. Because these approaches try to match views of the same sample, they can be too myopic and fail to produce meaningful results when augmentations are not sufficiently rich. This motivates the use of the dataset itself to find similar, yet distinct, samples to serve as views for one another. In this paper, we introduce Mine Your Own vieW (MYOW), a new approach for building across-sample prediction into SSL. The idea behind our approach is to actively mine views, finding samples that are close in the representation space of the network, and then predict, from one sample's latent representation, the representation of a nearby sample. In addition to showing the promise of MYOW on standard datasets used in computer vision, we highlight the power of this idea in a novel application in neuroscience where rich augmentations are not already established. When applied to neural datasets, MYOW outperforms other self-supervised approaches in all examples (in some cases by more than 10%), and surpasses the supervised baseline for most datasets. By learning to predict the latent representation of similar samples, we show that it is possible to learn good representations in new domains where augmentations are still limited.
Making transport more robust and interpretable by moving data through a small number of anchor points
Dec 21, 2020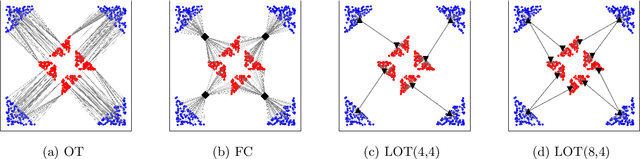
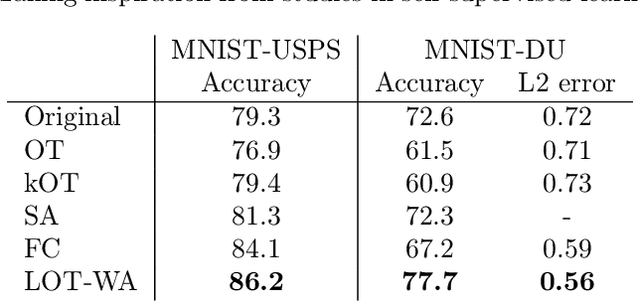
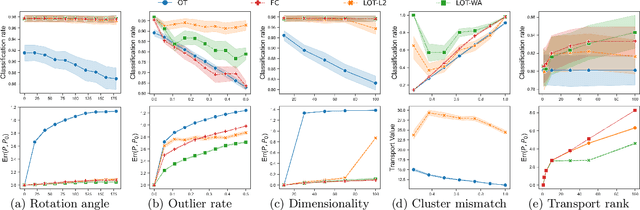
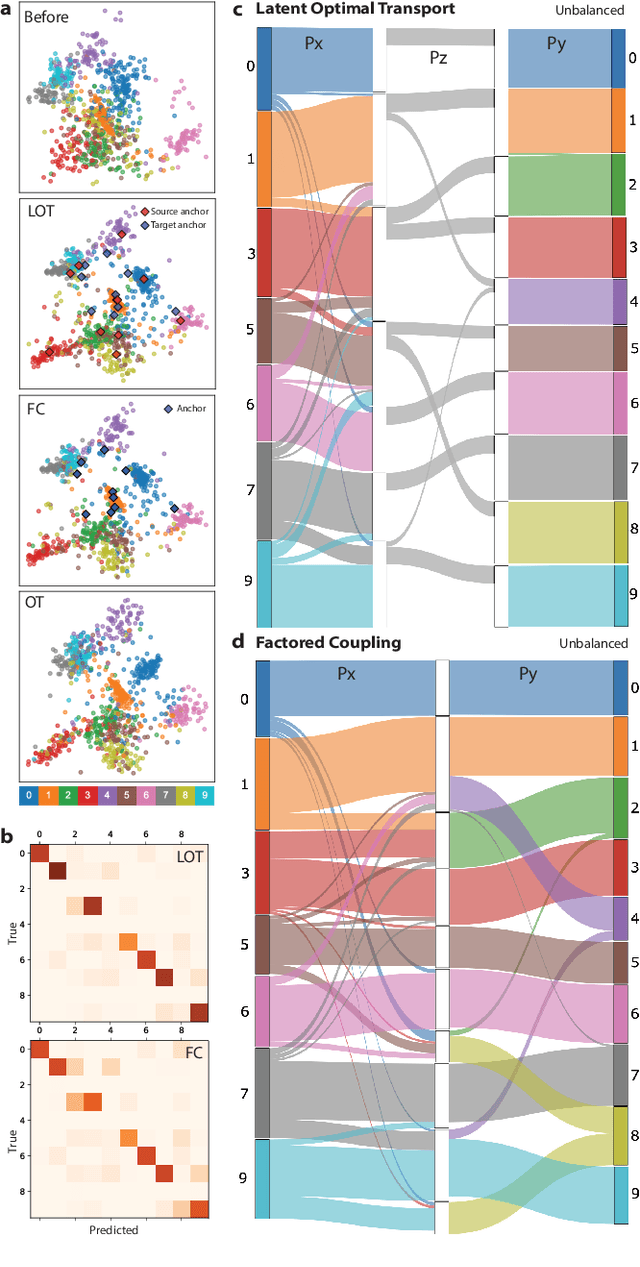
Abstract:Optimal transport (OT) is a widely used technique for distribution alignment, with applications throughout the machine learning, graphics, and vision communities. Without any additional structural assumptions on trans-port, however, OT can be fragile to outliers or noise, especially in high dimensions. Here, we introduce a new form of structured OT that simultaneously learns low-dimensional structure in data while leveraging this structure to solve the alignment task. Compared with OT, the resulting transport plan has better structural interpretability, highlighting the connections between individual data points and local geometry, and is more robust to noise and sampling. We apply the method to synthetic as well as real datasets, where we show that our method can facilitate alignment in noisy settings and can be used to both correct and interpret domain shift.
Bayesian optimization for modular black-box systems with switching costs
Jun 04, 2020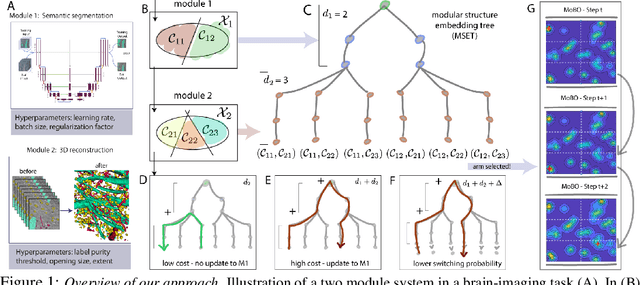
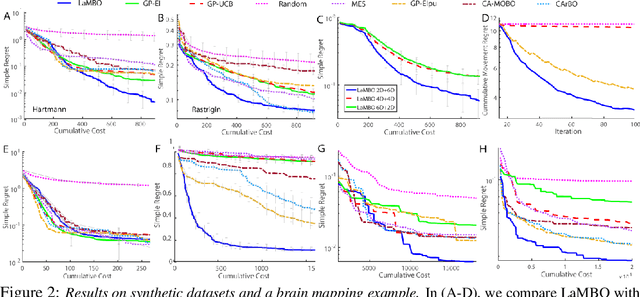
Abstract:Most existing black-box optimization methods assume that all variables in the system being optimized have equal cost and can change freely at each iteration. However, in many real world systems, inputs are passed through a sequence of different operations or modules, making variables in earlier stages of processing more costly to update. Such structure imposes a cost on switching variables in early parts of a data processing pipeline. In this work, we propose a new algorithm for switch cost-aware optimization called Lazy Modular Bayesian Optimization (LaMBO). This method efficiently identifies the global optimum while minimizing cost through a passive change of variables in early modules. The method is theoretical grounded and achieves vanishing regret when augmented with switching cost. We apply LaMBO to multiple synthetic functions and a three-stage image segmentation pipeline used in a neuroscience application, where we obtain promising improvements over prevailing cost-aware Bayesian optimization algorithms. Our results demonstrate that LaMBO is an effective strategy for black-box optimization that is capable of minimizing switching costs in modular systems.
Hierarchical Optimal Transport for Multimodal Distribution Alignment
Jun 27, 2019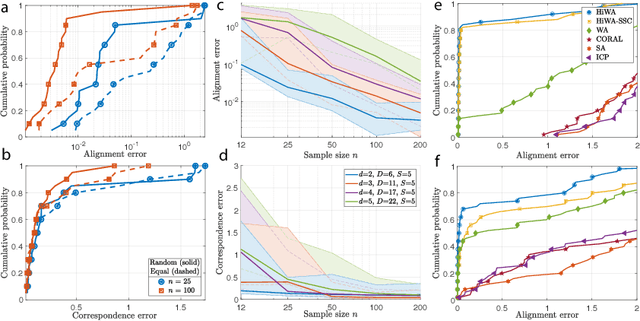
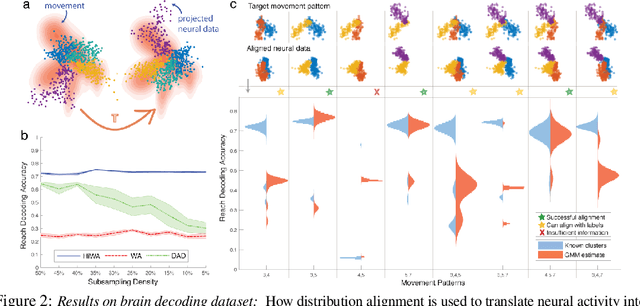
Abstract:In many machine learning applications, it is necessary to meaningfully aggregate, through alignment, different but related datasets. Optimal transport (OT)-based approaches pose alignment as a divergence minimization problem: the aim is to transform a source dataset to match a target dataset using the Wasserstein distance as a divergence measure. We introduce a hierarchical formulation of OT which leverages clustered structure in data to improve alignment in noisy, ambiguous, or multimodal settings. To solve this numerically, we propose a distributed ADMM algorithm that also exploits the Sinkhorn distance, thus it has an efficient computational complexity that scales quadratically with the size of the largest cluster. When the transformation between two datasets is unitary, we provide performance guarantees that describe when and how well aligned cluster correspondences can be recovered with our formulation, as well as provide worst-case dataset geometry for such a strategy. We apply this method to synthetic datasets that model data as mixtures of low-rank Gaussians and study the impact that different geometric properties of the data have on alignment. Next, we applied our approach to a neural decoding application where the goal is to predict movement directions and instantaneous velocities from populations of neurons in the macaque primary motor cortex. Our results demonstrate that when clustered structure exists in datasets, and is consistent across trials or time points, a hierarchical alignment strategy that leverages such structure can provide significant improvements in cross-domain alignment.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge