Eric Xing
Carnegie Mellon University
Stochastic Neural Networks with Infinite Width are Deterministic
Jan 30, 2022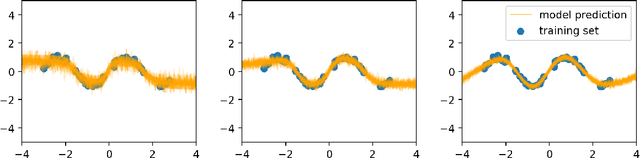
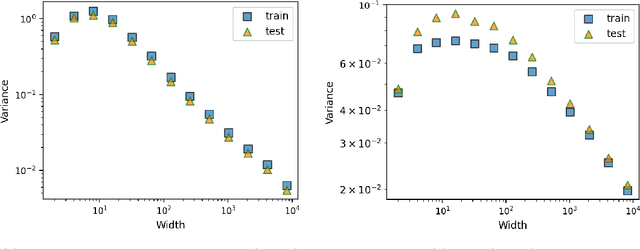
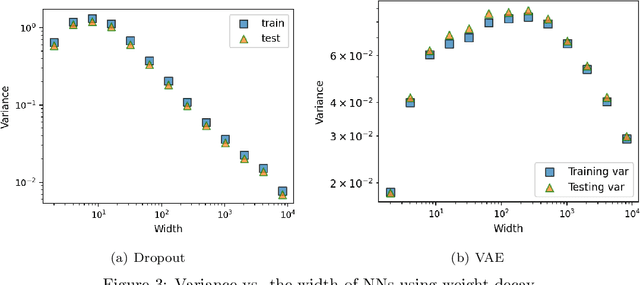
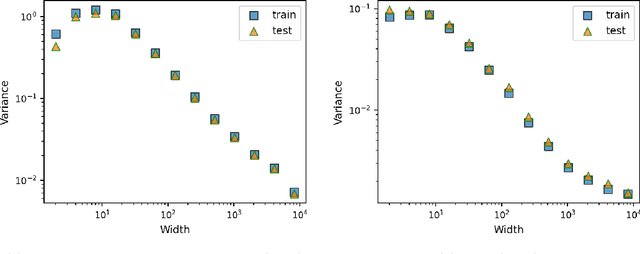
Abstract:This work theoretically studies stochastic neural networks, a main type of neural network in use. Specifically, we prove that as the width of an optimized stochastic neural network tends to infinity, its predictive variance on the training set decreases to zero. Two common examples that our theory applies to are neural networks with dropout and variational autoencoders. Our result helps better understand how stochasticity affects the learning of neural networks and thus design better architectures for practical problems.
Vision Transformer Slimming: Multi-Dimension Searching in Continuous Optimization Space
Jan 03, 2022
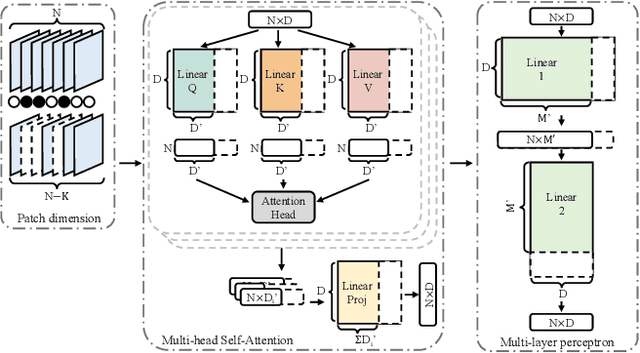
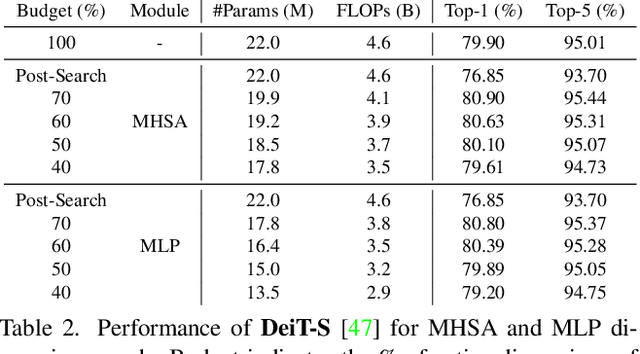
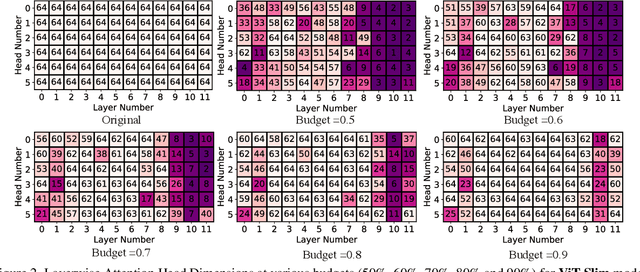
Abstract:This paper explores the feasibility of finding an optimal sub-model from a vision transformer and introduces a pure vision transformer slimming (ViT-Slim) framework that can search such a sub-structure from the original model end-to-end across multiple dimensions, including the input tokens, MHSA and MLP modules with state-of-the-art performance. Our method is based on a learnable and unified l1 sparsity constraint with pre-defined factors to reflect the global importance in the continuous searching space of different dimensions. The searching process is highly efficient through a single-shot training scheme. For instance, on DeiT-S, ViT-Slim only takes ~43 GPU hours for searching process, and the searched structure is flexible with diverse dimensionalities in different modules. Then, a budget threshold is employed according to the requirements of accuracy-FLOPs trade-off on running devices, and a re-training process is performed to obtain the final models. The extensive experiments show that our ViT-Slim can compress up to 40% of parameters and 40% FLOPs on various vision transformers while increasing the accuracy by ~0.6% on ImageNet. We also demonstrate the advantage of our searched models on several downstream datasets. Our source code will be publicly available.
Data-Free Neural Architecture Search via Recursive Label Calibration
Dec 03, 2021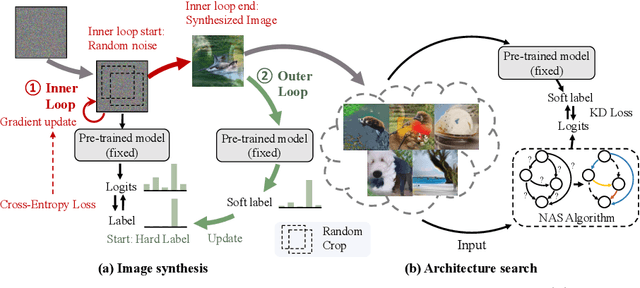
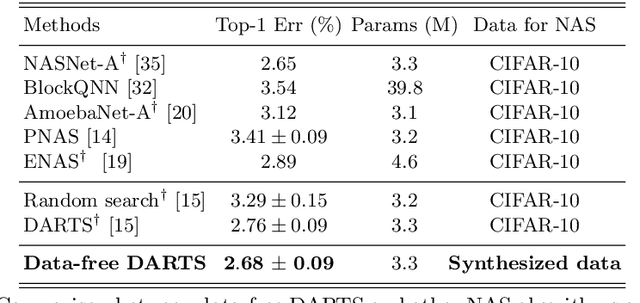
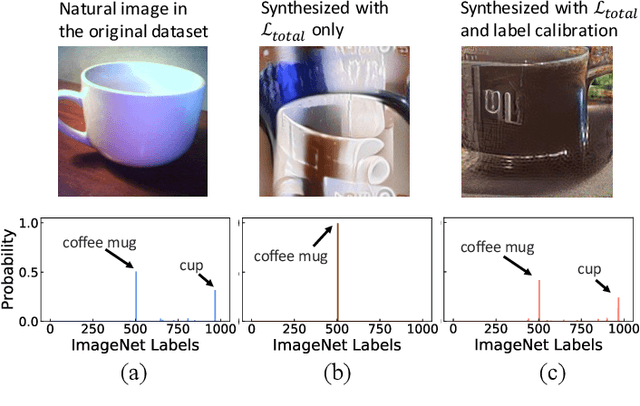
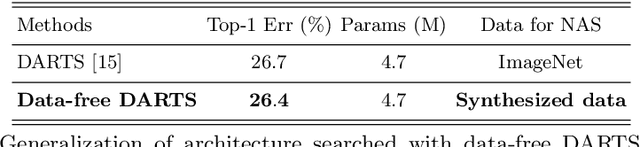
Abstract:This paper aims to explore the feasibility of neural architecture search (NAS) given only a pre-trained model without using any original training data. This is an important circumstance for privacy protection, bias avoidance, etc., in real-world scenarios. To achieve this, we start by synthesizing usable data through recovering the knowledge from a pre-trained deep neural network. Then we use the synthesized data and their predicted soft-labels to guide neural architecture search. We identify that the NAS task requires the synthesized data (we target at image domain here) with enough semantics, diversity, and a minimal domain gap from the natural images. For semantics, we propose recursive label calibration to produce more informative outputs. For diversity, we propose a regional update strategy to generate more diverse and semantically-enriched synthetic data. For minimal domain gap, we use input and feature-level regularization to mimic the original data distribution in latent space. We instantiate our proposed framework with three popular NAS algorithms: DARTS, ProxylessNAS and SPOS. Surprisingly, our results demonstrate that the architectures discovered by searching with our synthetic data achieve accuracy that is comparable to, or even higher than, architectures discovered by searching from the original ones, for the first time, deriving the conclusion that NAS can be done effectively with no need of access to the original or called natural data if the synthesis method is well designed. Our code will be publicly available.
A Fast Knowledge Distillation Framework for Visual Recognition
Dec 02, 2021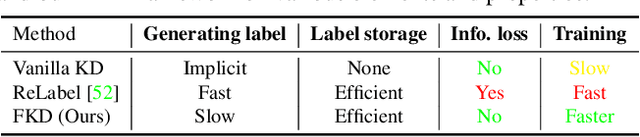
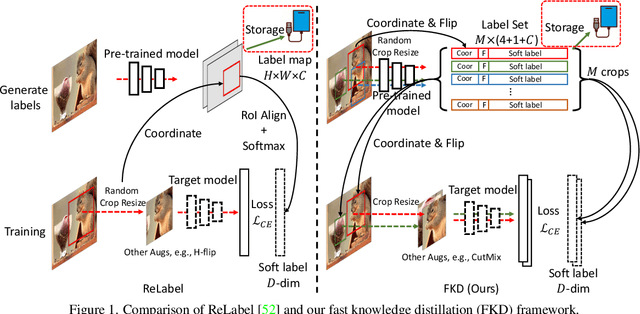
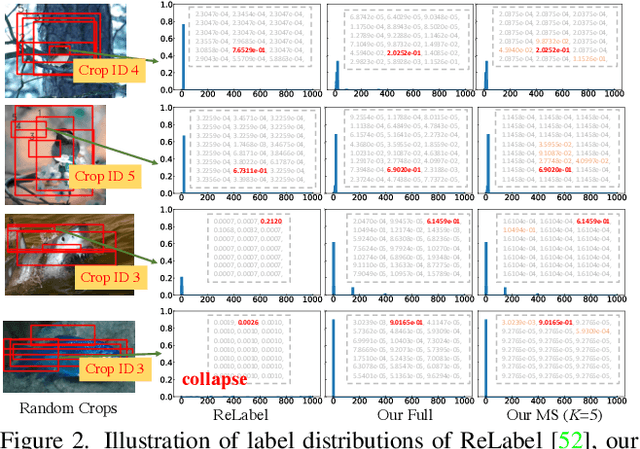
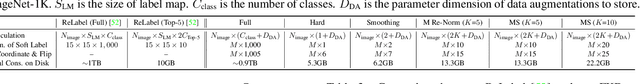
Abstract:While Knowledge Distillation (KD) has been recognized as a useful tool in many visual tasks, such as supervised classification and self-supervised representation learning, the main drawback of a vanilla KD framework is its mechanism, which consumes the majority of the computational overhead on forwarding through the giant teacher networks, making the entire learning procedure inefficient and costly. ReLabel, a recently proposed solution, suggests creating a label map for the entire image. During training, it receives the cropped region-level label by RoI aligning on a pre-generated entire label map, allowing for efficient supervision generation without having to pass through the teachers many times. However, as the KD teachers are from conventional multi-crop training, there are various mismatches between the global label-map and region-level label in this technique, resulting in performance deterioration. In this study, we present a Fast Knowledge Distillation (FKD) framework that replicates the distillation training phase and generates soft labels using the multi-crop KD approach, while training faster than ReLabel since no post-processes such as RoI align and softmax operations are used. When conducting multi-crop in the same image for data loading, our FKD is even more efficient than the traditional image classification framework. On ImageNet-1K, we obtain 79.8% with ResNet-50, outperforming ReLabel by ~1.0% while being faster. On the self-supervised learning task, we also show that FKD has an efficiency advantage. Our project page: http://zhiqiangshen.com/projects/FKD/index.html, source code and models are available at: https://github.com/szq0214/FKD.
Nonuniform-to-Uniform Quantization: Towards Accurate Quantization via Generalized Straight-Through Estimation
Nov 29, 2021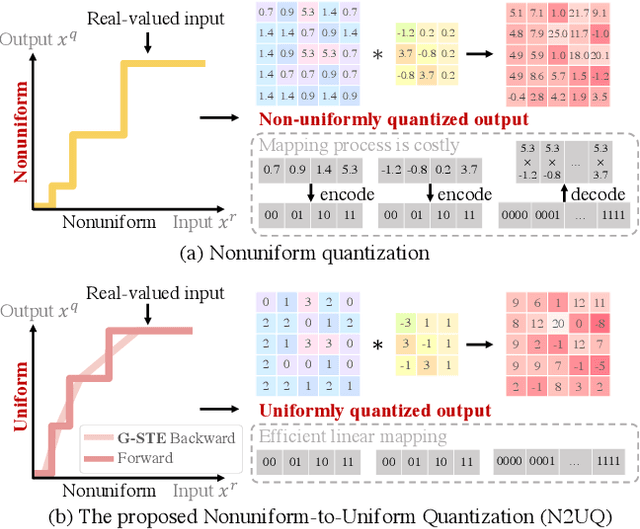
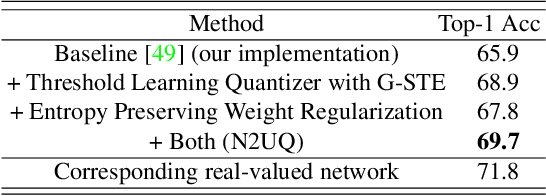
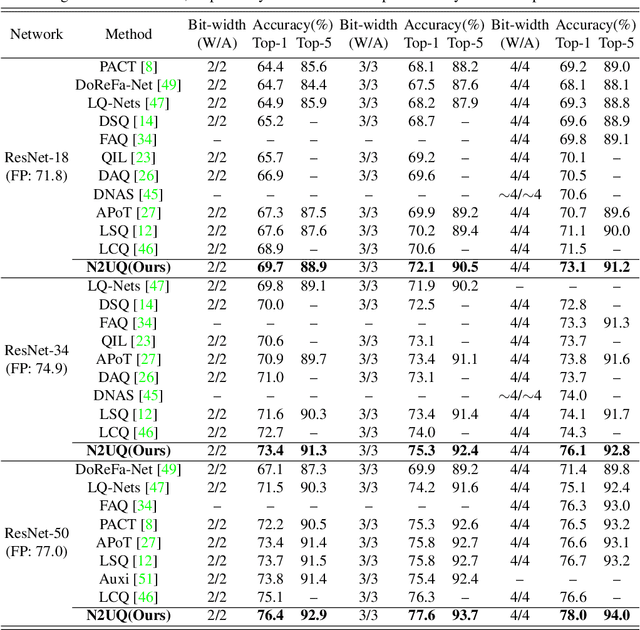
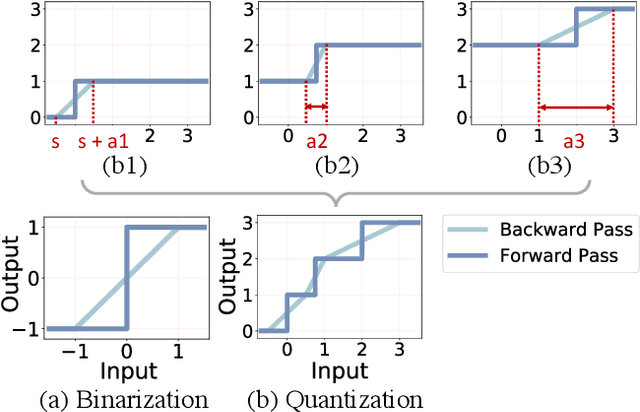

Abstract:The nonuniform quantization strategy for compressing neural networks usually achieves better performance than its counterpart, i.e., uniform strategy, due to its superior representational capacity. However, many nonuniform quantization methods overlook the complicated projection process in implementing the nonuniformly quantized weights/activations, which incurs non-negligible time and space overhead in hardware deployment. In this study, we propose Nonuniform-to-Uniform Quantization (N2UQ), a method that can maintain the strong representation ability of nonuniform methods while being hardware-friendly and efficient as the uniform quantization for model inference. We achieve this through learning the flexible in-equidistant input thresholds to better fit the underlying distribution while quantizing these real-valued inputs into equidistant output levels. To train the quantized network with learnable input thresholds, we introduce a generalized straight-through estimator (G-STE) for intractable backward derivative calculation w.r.t. threshold parameters. Additionally, we consider entropy preserving regularization to further reduce information loss in weight quantization. Even under this adverse constraint of imposing uniformly quantized weights and activations, our N2UQ outperforms state-of-the-art nonuniform quantization methods by 0.7~1.8% on ImageNet, demonstrating the contribution of N2UQ design. Code will be made publicly available.
Learning from Mistakes -- A Framework for Neural Architecture Search
Nov 11, 2021



Abstract:Learning from one's mistakes is an effective human learning technique where the learners focus more on the topics where mistakes were made, so as to deepen their understanding. In this paper, we investigate if this human learning strategy can be applied in machine learning. We propose a novel machine learning method called Learning From Mistakes (LFM), wherein the learner improves its ability to learn by focusing more on the mistakes during revision. We formulate LFM as a three-stage optimization problem: 1) learner learns; 2) learner re-learns focusing on the mistakes, and; 3) learner validates its learning. We develop an efficient algorithm to solve the LFM problem. We apply the LFM framework to neural architecture search on CIFAR-10, CIFAR-100, and Imagenet. Experimental results strongly demonstrate the effectiveness of our model.
Sliced Recursive Transformer
Nov 09, 2021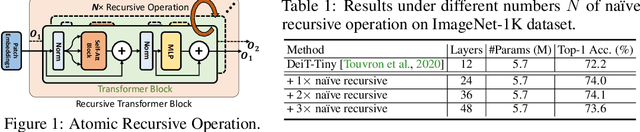
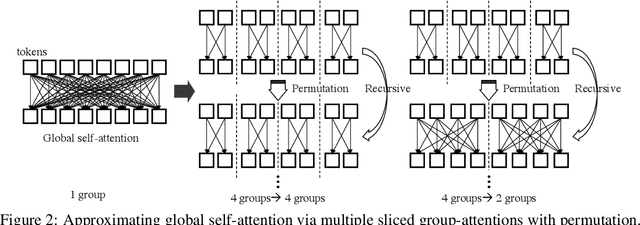
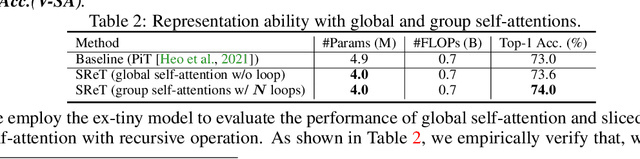
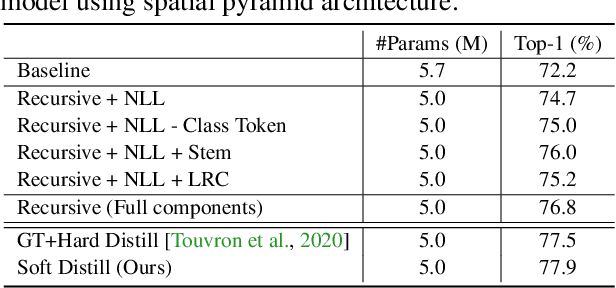
Abstract:We present a neat yet effective recursive operation on vision transformers that can improve parameter utilization without involving additional parameters. This is achieved by sharing weights across depth of transformer networks. The proposed method can obtain a substantial gain (~2%) simply using na\"ive recursive operation, requires no special or sophisticated knowledge for designing principles of networks, and introduces minimum computational overhead to the training procedure. To reduce the additional computation caused by recursive operation while maintaining the superior accuracy, we propose an approximating method through multiple sliced group self-attentions across recursive layers which can reduce the cost consumption by 10~30% with minimal performance loss. We call our model Sliced Recursive Transformer (SReT), which is compatible with a broad range of other designs for efficient vision transformers. Our best model establishes significant improvement on ImageNet over state-of-the-art methods while containing fewer parameters. The proposed sliced recursive operation allows us to build a transformer with more than 100 or even 1000 layers effortlessly under a still small size (13~15M), to avoid difficulties in optimization when the model size is too large. The flexible scalability has shown great potential for scaling up and constructing extremely deep and large dimensionality vision transformers. Our code and models are available at https://github.com/szq0214/SReT.
Toward Learning Human-aligned Cross-domain Robust Models by Countering Misaligned Features
Nov 05, 2021
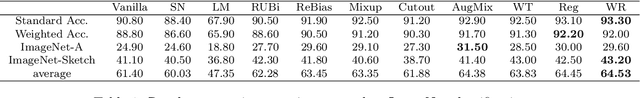
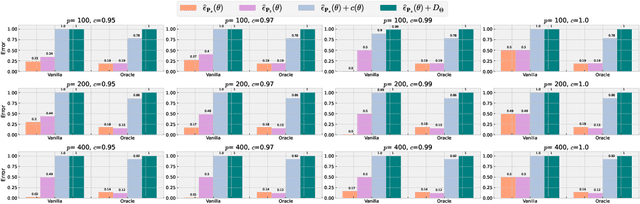
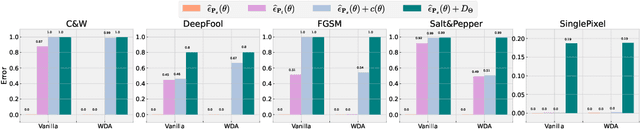
Abstract:Machine learning has demonstrated remarkable prediction accuracy over i.i.d data, but the accuracy often drops when tested with data from another distribution. In this paper, we aim to offer another view of this problem in a perspective assuming the reason behind this accuracy drop is the reliance of models on the features that are not aligned well with how a data annotator considers similar across these two datasets. We refer to these features as misaligned features. We extend the conventional generalization error bound to a new one for this setup with the knowledge of how the misaligned features are associated with the label. Our analysis offers a set of techniques for this problem, and these techniques are naturally linked to many previous methods in robust machine learning literature. We also compared the empirical strength of these methods demonstrated the performance when these previous techniques are combined.
Tradeoffs of Linear Mixed Models in Genome-wide Association Studies
Nov 05, 2021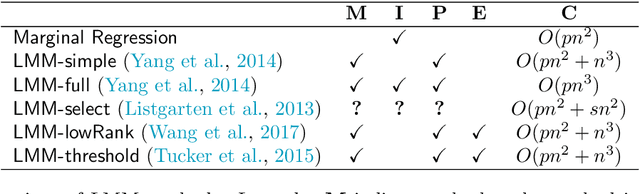
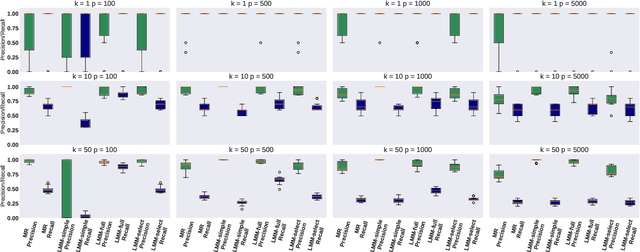
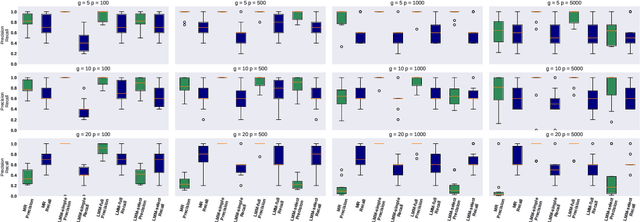
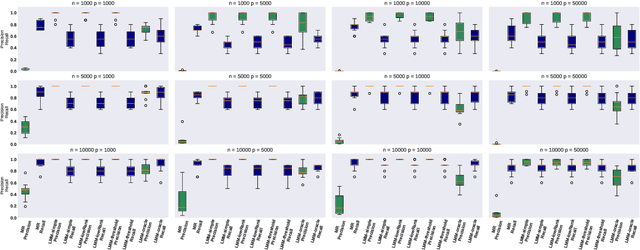
Abstract:Motivated by empirical arguments that are well-known from the genome-wide association studies (GWAS) literature, we study the statistical properties of linear mixed models (LMMs) applied to GWAS. First, we study the sensitivity of LMMs to the inclusion of a candidate SNP in the kinship matrix, which is often done in practice to speed up computations. Our results shed light on the size of the error incurred by including a candidate SNP, providing a justification to this technique in order to trade-off velocity against veracity. Second, we investigate how mixed models can correct confounders in GWAS, which is widely accepted as an advantage of LMMs over traditional methods. We consider two sources of confounding factors, population stratification and environmental confounding factors, and study how different methods that are commonly used in practice trade-off these two confounding factors differently.
Multi-task Learning of Order-Consistent Causal Graphs
Nov 03, 2021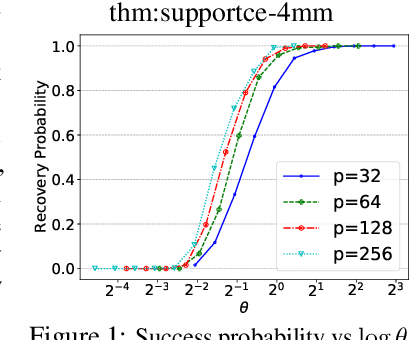
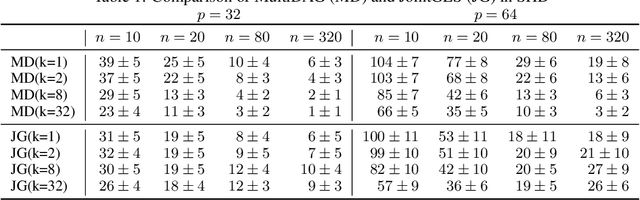
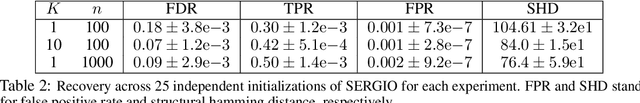

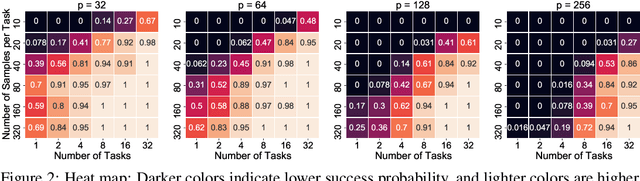

Abstract:We consider the problem of discovering $K$ related Gaussian directed acyclic graphs (DAGs), where the involved graph structures share a consistent causal order and sparse unions of supports. Under the multi-task learning setting, we propose a $l_1/l_2$-regularized maximum likelihood estimator (MLE) for learning $K$ linear structural equation models. We theoretically show that the joint estimator, by leveraging data across related tasks, can achieve a better sample complexity for recovering the causal order (or topological order) than separate estimations. Moreover, the joint estimator is able to recover non-identifiable DAGs, by estimating them together with some identifiable DAGs. Lastly, our analysis also shows the consistency of union support recovery of the structures. To allow practical implementation, we design a continuous optimization problem whose optimizer is the same as the joint estimator and can be approximated efficiently by an iterative algorithm. We validate the theoretical analysis and the effectiveness of the joint estimator in experiments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge