Ender Konukoglu
for the Alzheimer's Disease Neuroimaging Initiative
Gradient flow encoding with distance optimization adaptive step size
May 11, 2021
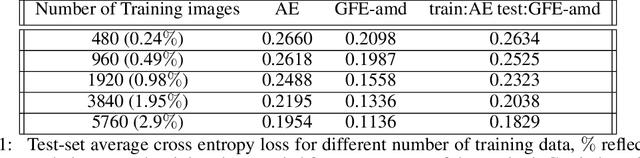
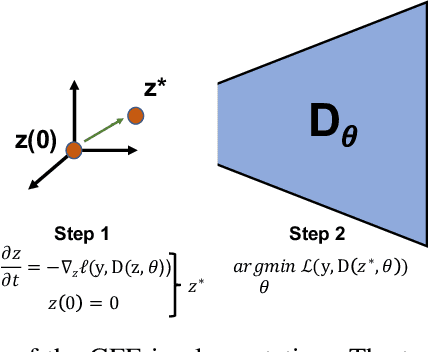
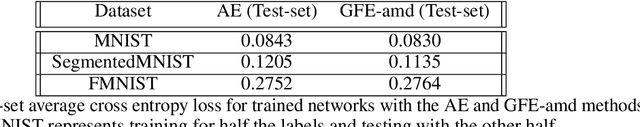
Abstract:The autoencoder model uses an encoder to map data samples to a lower dimensional latent space and then a decoder to map the latent space representations back to the data space. Implicitly, it relies on the encoder to approximate the inverse of the decoder network, so that samples can be mapped to and back from the latent space faithfully. This approximation may lead to sub-optimal latent space representations. In this work, we investigate a decoder-only method that uses gradient flow to encode data samples in the latent space. The gradient flow is defined based on a given decoder and aims to find the optimal latent space representation for any given sample through optimisation, eliminating the need of an approximate inversion through an encoder. Implementing gradient flow through ordinary differential equations (ODE), we leverage the adjoint method to train a given decoder. We further show empirically that the costly integrals in the adjoint method may not be entirely necessary. Additionally, we propose a $2^{nd}$ order ODE variant to the method, which approximates Nesterov's accelerated gradient descent, with faster convergence per iteration. Commonly used ODE solvers can be quite sensitive to the integration step-size depending on the stiffness of the ODE. To overcome the sensitivity for gradient flow encoding, we use an adaptive solver that prioritises minimising loss at each integration step. We assess the proposed method in comparison to the autoencoding model. In our experiments, GFE showed a much higher data-efficiency than the autoencoding model, which can be crucial for data scarce applications.
Contrastive Learning of Single-Cell Phenotypic Representations for Treatment Classification
Mar 30, 2021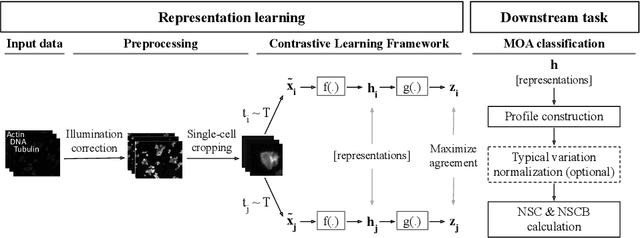
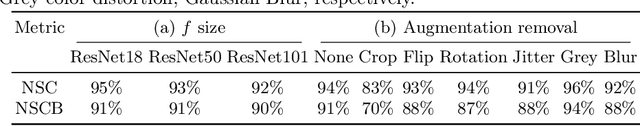

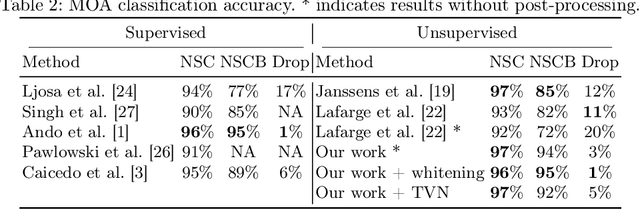
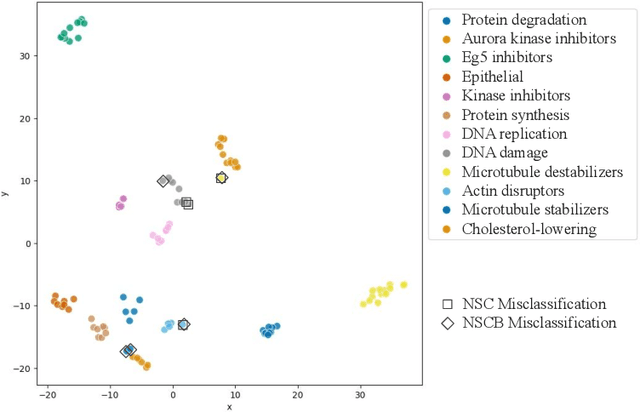
Abstract:Learning robust representations to discriminate cell phenotypes based on microscopy images is important for drug discovery. Drug development efforts typically analyse thousands of cell images to screen for potential treatments. Early works focus on creating hand-engineered features from these images or learn such features with deep neural networks in a fully or weakly-supervised framework. Both require prior knowledge or labelled datasets. Therefore, subsequent works propose unsupervised approaches based on generative models to learn these representations. Recently, representations learned with self-supervised contrastive loss-based methods have yielded state-of-the-art results on various imaging tasks compared to earlier unsupervised approaches. In this work, we leverage a contrastive learning framework to learn appropriate representations from single-cell fluorescent microscopy images for the task of Mechanism-of-Action classification. The proposed work is evaluated on the annotated BBBC021 dataset, and we obtain state-of-the-art results in NSC, NCSB and drop metrics for an unsupervised approach. We observe an improvement of 10% in NCSB accuracy and 11% in NSC-NSCB drop over the previously best unsupervised method. Moreover, the performance of our unsupervised approach ties with the best supervised approach. Additionally, we observe that our framework performs well even without post-processing, unlike earlier methods. With this, we conclude that one can learn robust cell representations with contrastive learning.
Constrained Optimization for Training Deep Neural Networks Under Class Imbalance
Feb 21, 2021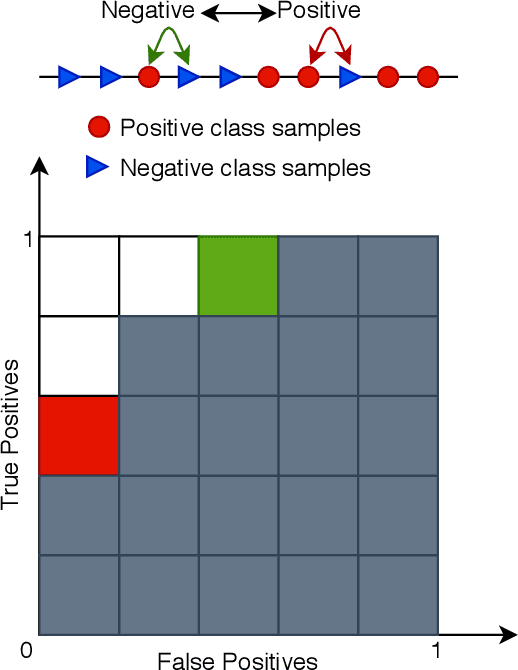
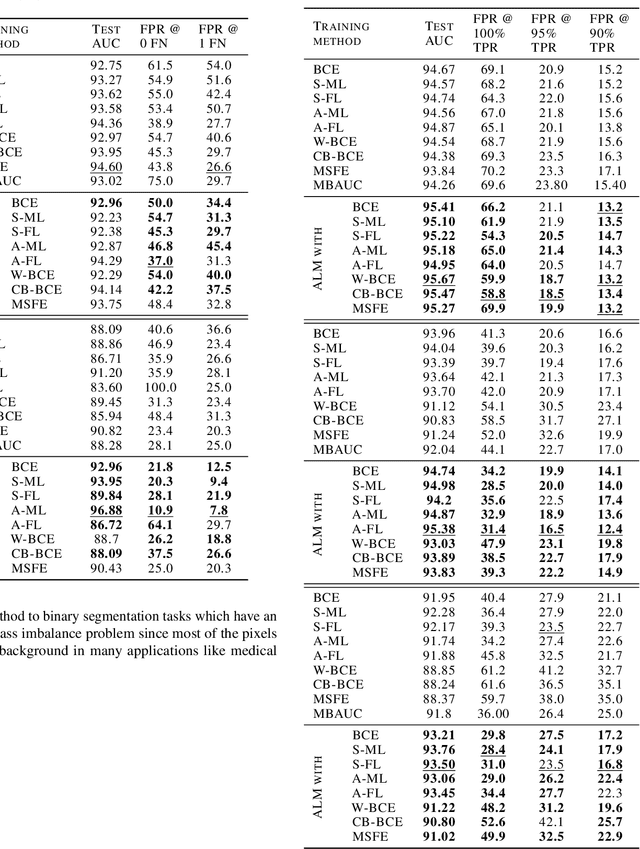
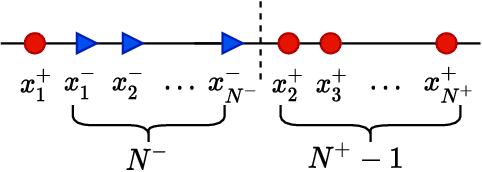
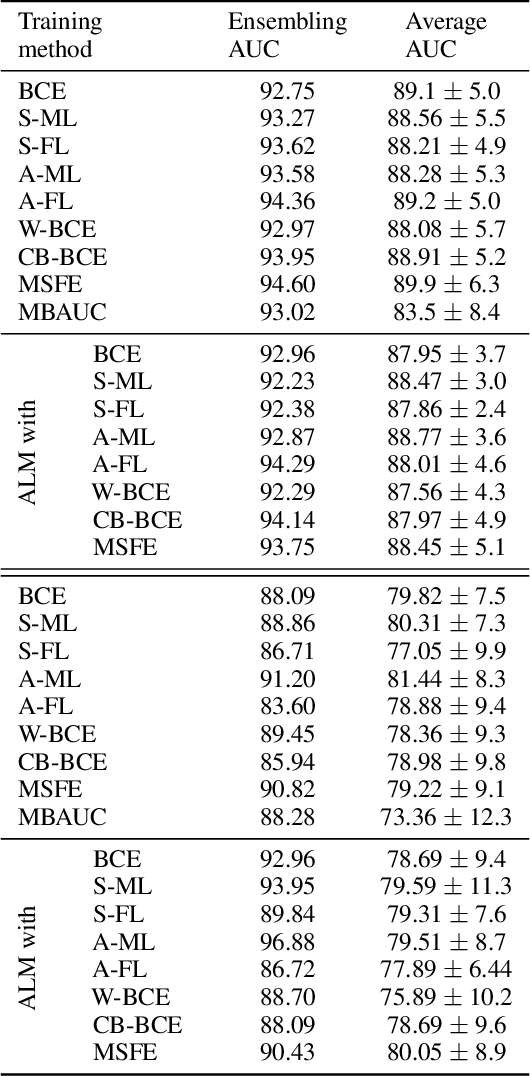
Abstract:Deep neural networks (DNNs) are notorious for making more mistakes for the classes that have substantially fewer samples than the others during training. Such class imbalance is ubiquitous in clinical applications and very crucial to handle because the classes with fewer samples most often correspond to critical cases (e.g., cancer) where misclassifications can have severe consequences. Not to miss such cases, binary classifiers need to be operated at high True Positive Rates (TPR) by setting a higher threshold but this comes at the cost of very high False Positive Rates (FPR) for problems with class imbalance. Existing methods for learning under class imbalance most often do not take this into account. We argue that prediction accuracy should be improved by emphasizing reducing FPRs at high TPRs for problems where misclassification of the positive samples are associated with higher cost. To this end, we pose the training of a DNN for binary classification as a constrained optimization problem and introduce a novel constraint that can be used with existing loss functions to enforce maximal area under the ROC curve (AUC). We solve the resulting constrained optimization problem using an Augmented Lagrangian method (ALM), where the constraint emphasizes reduction of FPR at high TPR. We present experimental results for image-based classification applications using the CIFAR10 and an in-house medical imaging dataset. Our results demonstrate that the proposed method almost always improves the loss functions it is used with by attaining lower FPR at high TPR and higher or equal AUC.
Exploring Cross-Image Pixel Contrast for Semantic Segmentation
Feb 11, 2021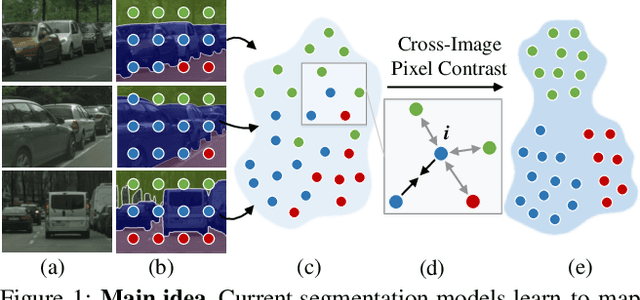
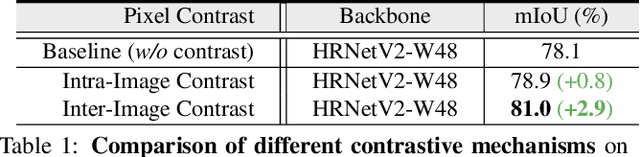
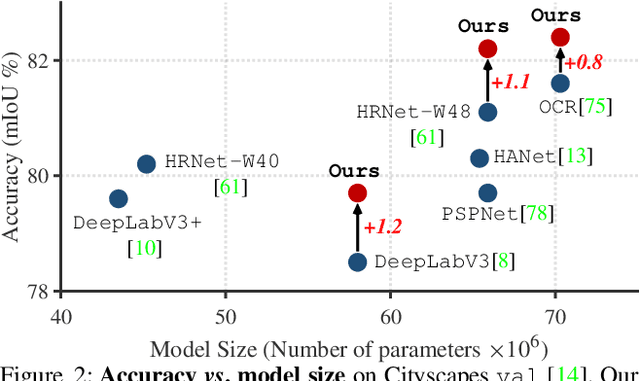

Abstract:Current semantic segmentation methods focus only on mining "local" context, i.e., dependencies between pixels within individual images, by context-aggregation modules (e.g., dilated convolution, neural attention) or structure-aware optimization criteria (e.g., IoU-like loss). However, they ignore "global" context of the training data, i.e., rich semantic relations between pixels across different images. Inspired by the recent advance in unsupervised contrastive representation learning, we propose a pixel-wise contrastive framework for semantic segmentation in the fully supervised setting. The core idea is to enforce pixel embeddings belonging to a same semantic class to be more similar than embeddings from different classes. It raises a pixel-wise metric learning paradigm for semantic segmentation, by explicitly exploring the structures of labeled pixels, which were rarely explored before. Our method can be effortlessly incorporated into existing segmentation frameworks without extra overhead during testing. We experimentally show that, with famous segmentation models (i.e., DeepLabV3, HRNet, OCR) and backbones (i.e., ResNet, HR-Net), our method brings consistent performance improvements across diverse datasets (i.e., Cityscapes, PASCAL-Context, COCO-Stuff). We expect this work will encourage our community to rethink the current de facto training paradigm in fully supervised semantic segmentation.
Hyperspectral Image Super-Resolution with Spectral Mixup and Heterogeneous Datasets
Jan 19, 2021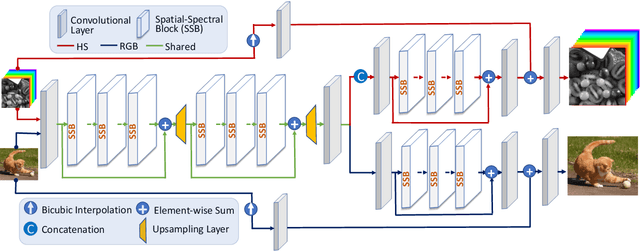
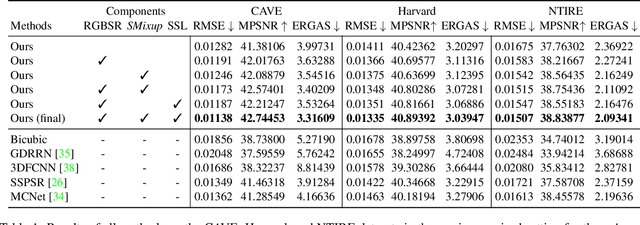
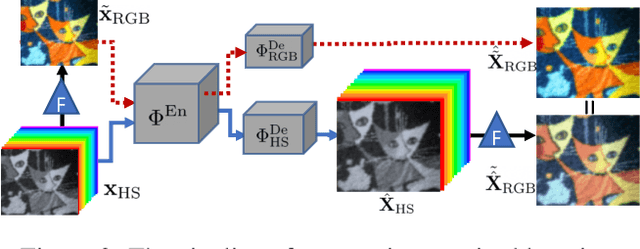
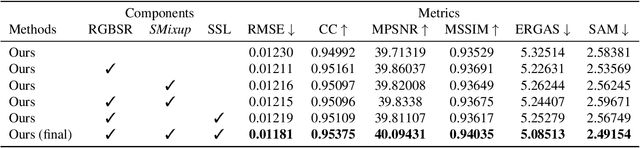
Abstract:This work studies Hyperspectral image (HSI) super-resolution (SR). HSI SR is characterized by high-dimensional data and a limited amount of training examples. This exacerbates the undesirable behaviors of neural networks such as memorization and sensitivity to out-of-distribution samples. This work addresses these issues with three contributions. First, we propose a simple, yet effective data augmentation routine, termed Spectral Mixup, to construct effective virtual training samples. Second, we observe that HSI SR and RGB image SR are correlated and develop a novel multi-tasking network to train them jointly so that the auxiliary task RGB image SR can provide additional supervision. Finally, we extend the network to a semi-supervised setting so that it can learn from datasets containing low-resolution HSIs only. With these contributions, our method is able to learn from heterogeneous datasets and lift the requirement for having a large amount of HD HSI training samples. Extensive experiments on four datasets show that our method outperforms existing methods significantly and underpin the relevance of our contributions. The code of this work will be released soon.
Probabilistic 3D surface reconstruction from sparse MRI information
Oct 05, 2020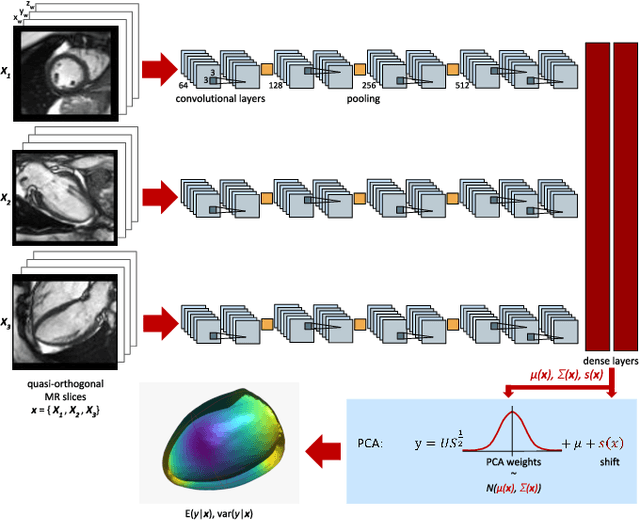
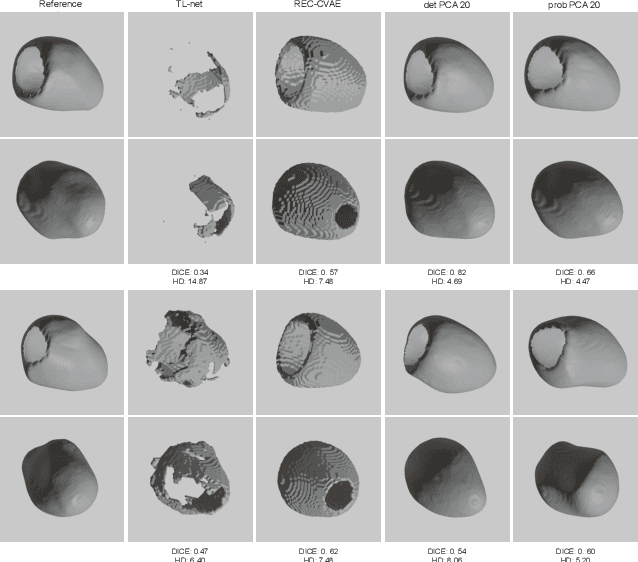
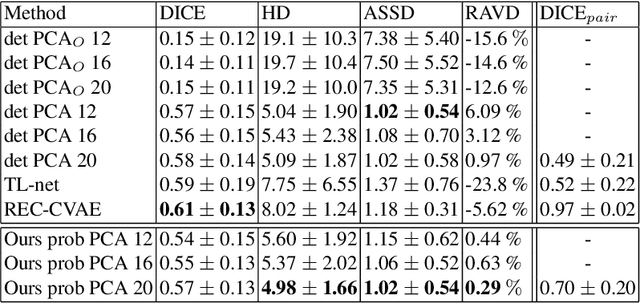

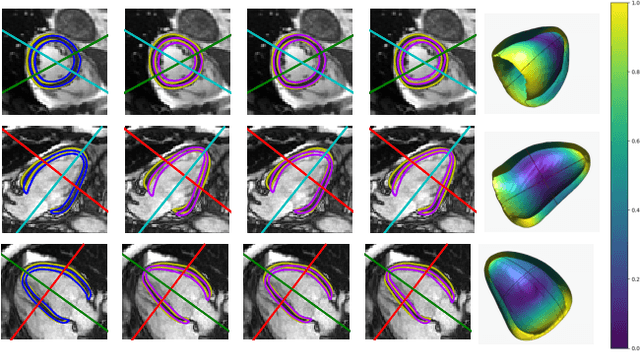
Abstract:Surface reconstruction from magnetic resonance (MR) imaging data is indispensable in medical image analysis and clinical research. A reliable and effective reconstruction tool should: be fast in prediction of accurate well localised and high resolution models, evaluate prediction uncertainty, work with as little input data as possible. Current deep learning state of the art (SOTA) 3D reconstruction methods, however, often only produce shapes of limited variability positioned in a canonical position or lack uncertainty evaluation. In this paper, we present a novel probabilistic deep learning approach for concurrent 3D surface reconstruction from sparse 2D MR image data and aleatoric uncertainty prediction. Our method is capable of reconstructing large surface meshes from three quasi-orthogonal MR imaging slices from limited training sets whilst modelling the location of each mesh vertex through a Gaussian distribution. Prior shape information is encoded using a built-in linear principal component analysis (PCA) model. Extensive experiments on cardiac MR data show that our probabilistic approach successfully assesses prediction uncertainty while at the same time qualitatively and quantitatively outperforms SOTA methods in shape prediction. Compared to SOTA, we are capable of properly localising and orientating the prediction via the use of a spatially aware neural network.
Sampling possible reconstructions of undersampled acquisitions in MR imaging
Sep 30, 2020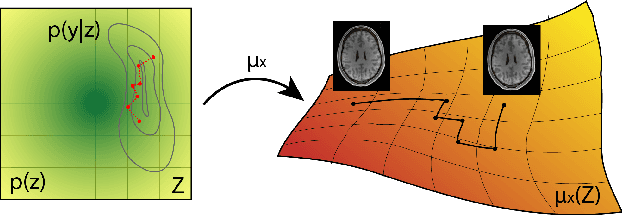
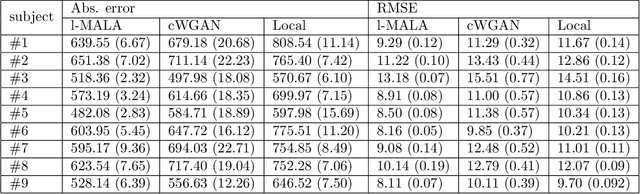
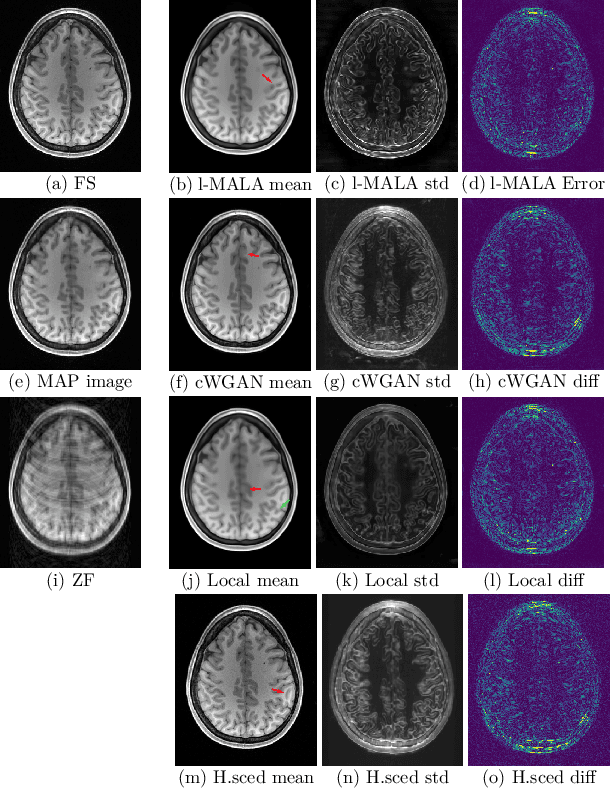
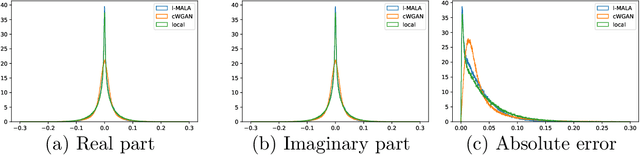
Abstract:Undersampling the k-space during MR acquisitions saves time, however results in an ill-posed inversion problem, leading to an infinite set of images as possible solutions. Traditionally, this is tackled as a reconstruction problem by searching for a single "best" image out of this solution set according to some chosen regularization or prior. This approach, however, misses the possibility of other solutions and hence ignores the uncertainty in the inversion process. In this paper, we propose a method that instead returns multiple images which are possible under the acquisition model and the chosen prior. To this end, we introduce a low dimensional latent space and model the posterior distribution of the latent vectors given the acquisition data in k-space, from which we can sample in the latent space and obtain the corresponding images. We use a variational autoencoder for the latent model and the Metropolis adjusted Langevin algorithm for the sampling. This approach allows us to obtain multiple possible images and capture the uncertainty in the inversion process under the used prior. We evaluate our method on images from the Human Connectome Project dataset as well as in-house measured multi-coil images and compare to two different methods. The results indicate that the proposed method is capable of producing images that match the ground truth in regions where acquired k-space data is informative and construct different possible reconstructions, which show realistic structural variations, in regions where acquired k-space data is not informative. Keywords: Magnetic Resonance image reconstruction, uncertainty estimation, inverse problems, sampling, MCMC, deep learning, unsupervised learning.
RevPHiSeg: A Memory-Efficient Neural Network for Uncertainty Quantification in Medical Image Segmentation
Aug 18, 2020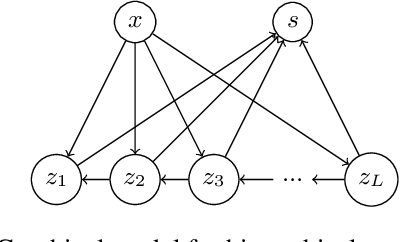
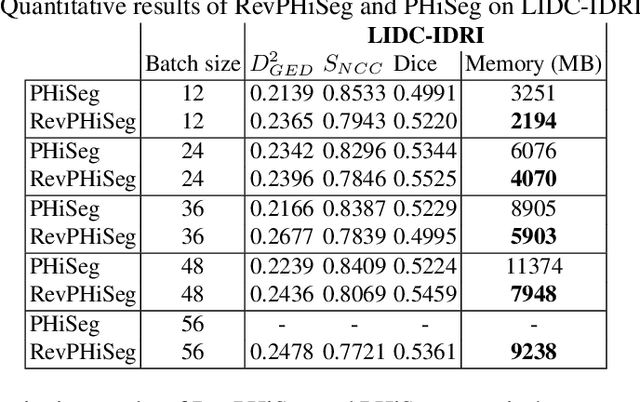
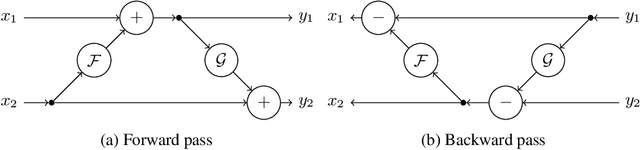
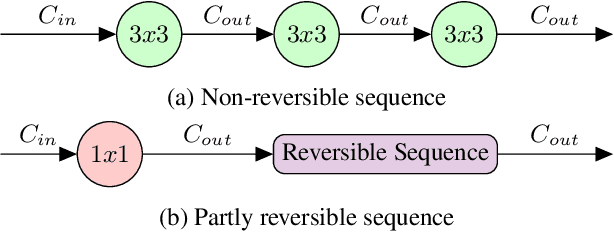
Abstract:Quantifying segmentation uncertainty has become an important issue in medical image analysis due to the inherent ambiguity of anatomical structures and its pathologies. Recently, neural network-based uncertainty quantification methods have been successfully applied to various problems. One of the main limitations of the existing techniques is the high memory requirement during training; which limits their application to processing smaller field-of-views (FOVs) and/or using shallower architectures. In this paper, we investigate the effect of using reversible blocks for building memory-efficient neural network architectures for quantification of segmentation uncertainty. The reversible architecture achieves memory saving by exactly computing the activations from the outputs of the subsequent layers during backpropagation instead of storing the activations for each layer. We incorporate the reversible blocks into a recently proposed architecture called PHiSeg that is developed for uncertainty quantification in medical image segmentation. The reversible architecture, RevPHiSeg, allows training neural networks for quantifying segmentation uncertainty on GPUs with limited memory and processing larger FOVs. We perform experiments on the LIDC-IDRI dataset and an in-house prostate dataset, and present comparisons with PHiSeg. The results demonstrate that RevPHiSeg consumes ~30% less memory compared to PHiSeg while achieving very similar segmentation accuracy.
Joint reconstruction and bias field correction for undersampled MR imaging
Jul 26, 2020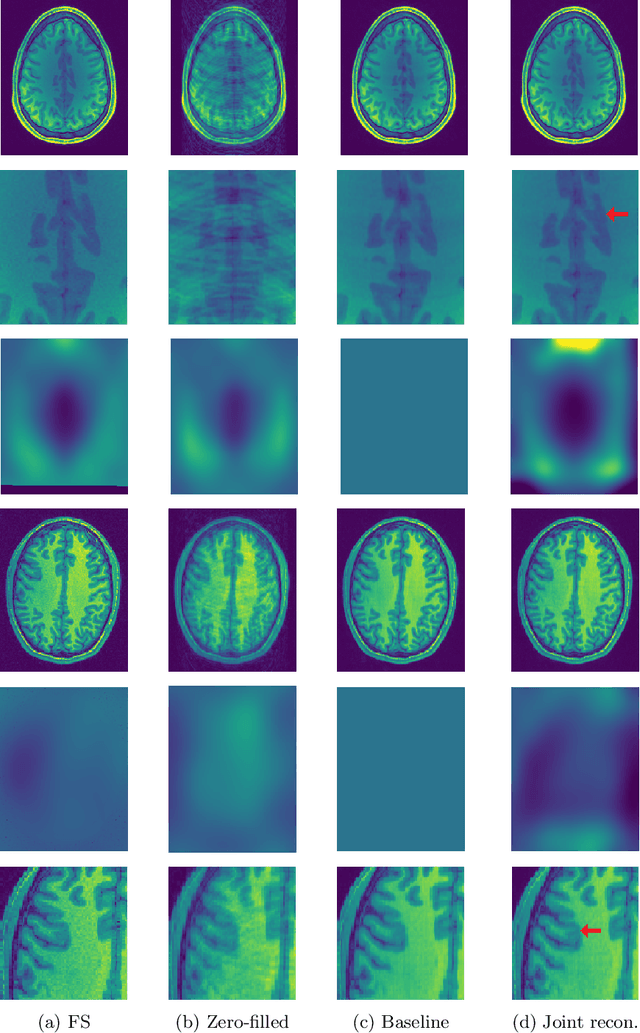
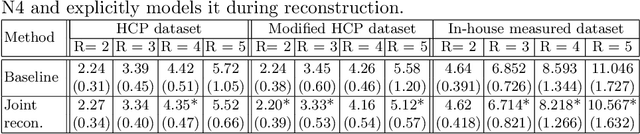
Abstract:Undersampling the k-space in MRI allows saving precious acquisition time, yet results in an ill-posed inversion problem. Recently, many deep learning techniques have been developed, addressing this issue of recovering the fully sampled MR image from the undersampled data. However, these learning based schemes are susceptible to differences between the training data and the image to be reconstructed at test time. One such difference can be attributed to the bias field present in MR images, caused by field inhomogeneities and coil sensitivities. In this work, we address the sensitivity of the reconstruction problem to the bias field and propose to model it explicitly in the reconstruction, in order to decrease this sensitivity. To this end, we use an unsupervised learning based reconstruction algorithm as our basis and combine it with a N4-based bias field estimation method, in a joint optimization scheme. We use the HCP dataset as well as in-house measured images for the evaluations. We show that the proposed method improves the reconstruction quality, both visually and in terms of RMSE.
Semi-supervised Task-driven Data Augmentation for Medical Image Segmentation
Jul 09, 2020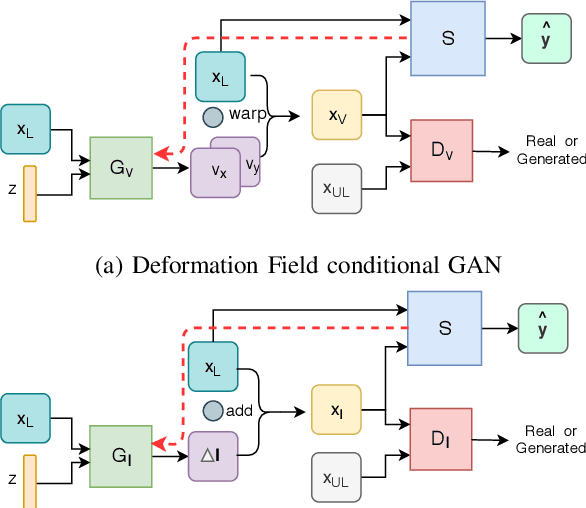
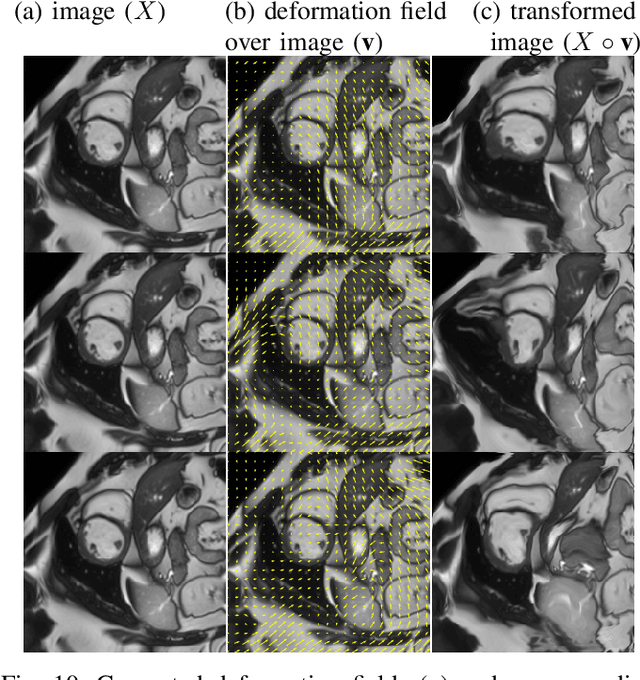
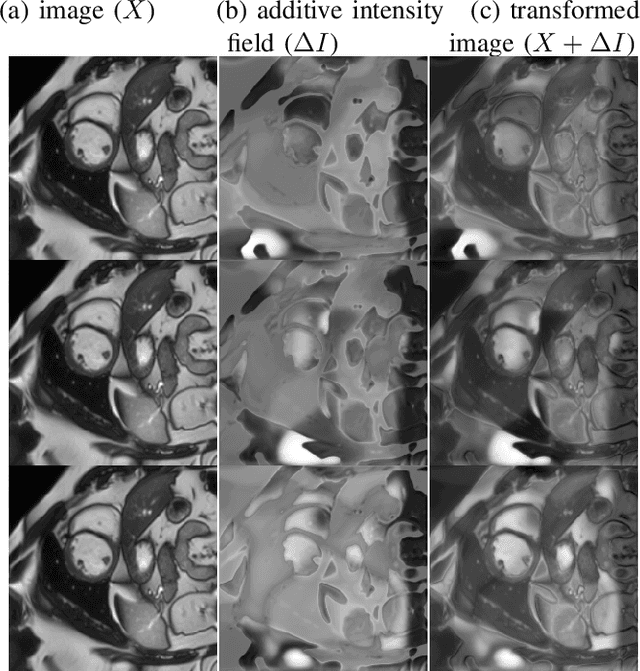
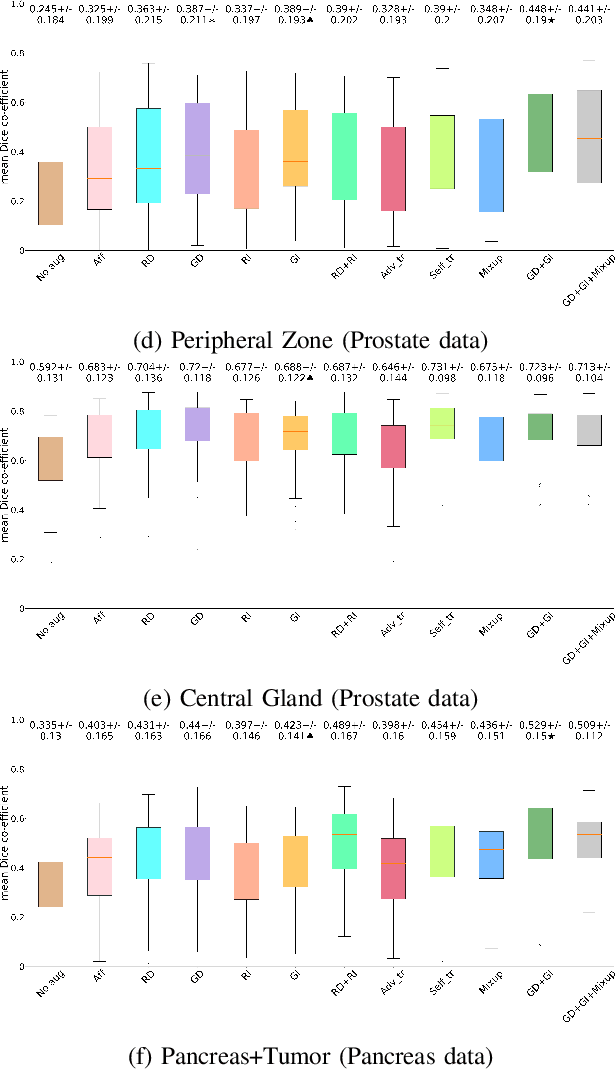
Abstract:Supervised learning-based segmentation methods typically require a large number of annotated training data to generalize well at test time. In medical applications, curating such datasets is not a favourable option because acquiring a large number of annotated samples from experts is time-consuming and expensive. Consequently, numerous methods have been proposed in the literature for learning with limited annotated examples. Unfortunately, the proposed approaches in the literature have not yet yielded significant gains over random data augmentation for image segmentation, where random augmentations themselves do not yield high accuracy. In this work, we propose a novel task-driven data augmentation method for learning with limited labeled data where the synthetic data generator, is optimized for the segmentation task. The generator of the proposed method models intensity and shape variations using two sets of transformations, as additive intensity transformations and deformation fields. Both transformations are optimized using labeled as well as unlabeled examples in a semi-supervised framework. Our experiments on three medical datasets, namely cardic, prostate and pancreas, show that the proposed approach significantly outperforms standard augmentation and semi-supervised approaches for image segmentation in the limited annotation setting. The code is made publicly available at https://github.com/krishnabits001/task$\_$driven$\_$data$\_$augmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge