Dominique van Midden
DALPHIN: Benchmarking Digital Pathology AI Copilots Against Pathologists on an Open Multicentric Dataset
May 05, 2026Abstract:Foundation models with visual question answering capabilities for digital pathology are emerging. Such unprecedented technology requires independent benchmarking to assess its potential in assisting pathologists in routine diagnostics. We created DALPHIN, the first multicentric open benchmark for pathology AI copilots, comprising 1236 images from 300 cases, spanning 130 rare to common diagnoses, 6 countries, and 14 subspecialties. The DALPHIN design and dataset are introduced alongside a human performance benchmark of 31 pathologists from 10 countries with varying expertise. We report results for two general-purpose (GPT-5, Gemini 2.5 Pro) and one pathology-specific copilot (PathChat+) for sequential and independent answer generation. We observed no statistically significant difference from expert-level performance in four of six tasks for PathChat, 2/6 tasks for Gemini, and 1/6 tasks for GPT. DALPHIN is publicly released with sequestered, indirectly accessible ground truth to foster robust and enduring benchmarking. Data, methods, and the evaluation platform are accessible through dalphin.grand-challenge.org.
Prototype-Based Multiple Instance Learning for Gigapixel Whole Slide Image Classification
Mar 11, 2025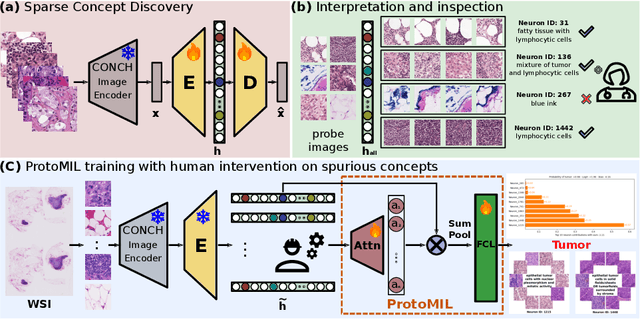
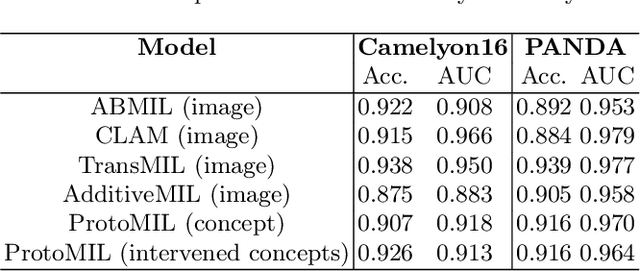
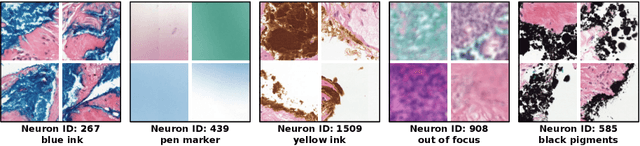
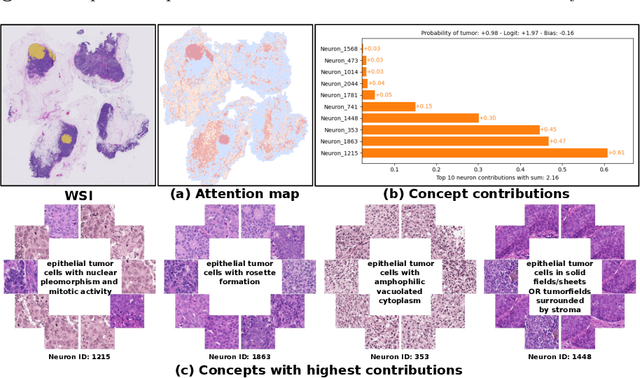
Abstract:Multiple Instance Learning (MIL) methods have succeeded remarkably in histopathology whole slide image (WSI) analysis. However, most MIL models only offer attention-based explanations that do not faithfully capture the model's decision mechanism and do not allow human-model interaction. To address these limitations, we introduce ProtoMIL, an inherently interpretable MIL model for WSI analysis that offers user-friendly explanations and supports human intervention. Our approach employs a sparse autoencoder to discover human-interpretable concepts from the image feature space, which are then used to train ProtoMIL. The model represents predictions as linear combinations of concepts, making the decision process transparent. Furthermore, ProtoMIL allows users to perform model interventions by altering the input concepts. Experiments on two widely used pathology datasets demonstrate that ProtoMIL achieves a classification performance comparable to state-of-the-art MIL models while offering intuitively understandable explanations. Moreover, we demonstrate that our method can eliminate reliance on diagnostically irrelevant information via human intervention, guiding the model toward being right for the right reason. Code will be publicly available at https://github.com/ss-sun/ProtoMIL.
Label-free Concept Based Multiple Instance Learning for Gigapixel Histopathology
Jan 06, 2025



Abstract:Multiple Instance Learning (MIL) methods allow for gigapixel Whole-Slide Image (WSI) analysis with only slide-level annotations. Interpretability is crucial for safely deploying such algorithms in high-stakes medical domains. Traditional MIL methods offer explanations by highlighting salient regions. However, such spatial heatmaps provide limited insights for end users. To address this, we propose a novel inherently interpretable WSI-classification approach that uses human-understandable pathology concepts to generate explanations. Our proposed Concept MIL model leverages recent advances in vision-language models to directly predict pathology concepts based on image features. The model's predictions are obtained through a linear combination of the concepts identified on the top-K patches of a WSI, enabling inherent explanations by tracing each concept's influence on the prediction. In contrast to traditional concept-based interpretable models, our approach eliminates the need for costly human annotations by leveraging the vision-language model. We validate our method on two widely used pathology datasets: Camelyon16 and PANDA. On both datasets, Concept MIL achieves AUC and accuracy scores over 0.9, putting it on par with state-of-the-art models. We further find that 87.1\% (Camelyon16) and 85.3\% (PANDA) of the top 20 patches fall within the tumor region. A user study shows that the concepts identified by our model align with the concepts used by pathologists, making it a promising strategy for human-interpretable WSI classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge