Deborah L. McGuinness
Explanation Ontology: A Model of Explanations for User-Centered AI
Oct 04, 2020
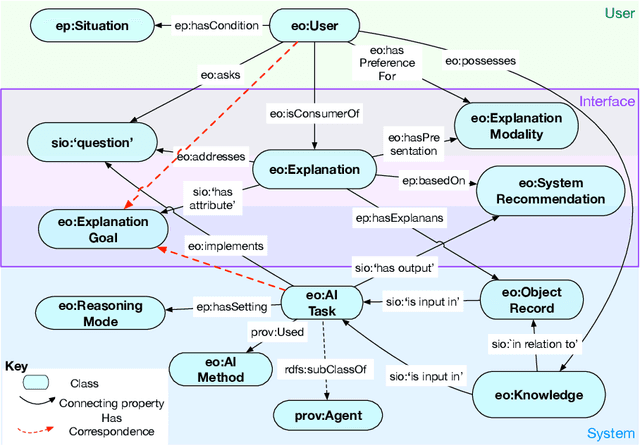
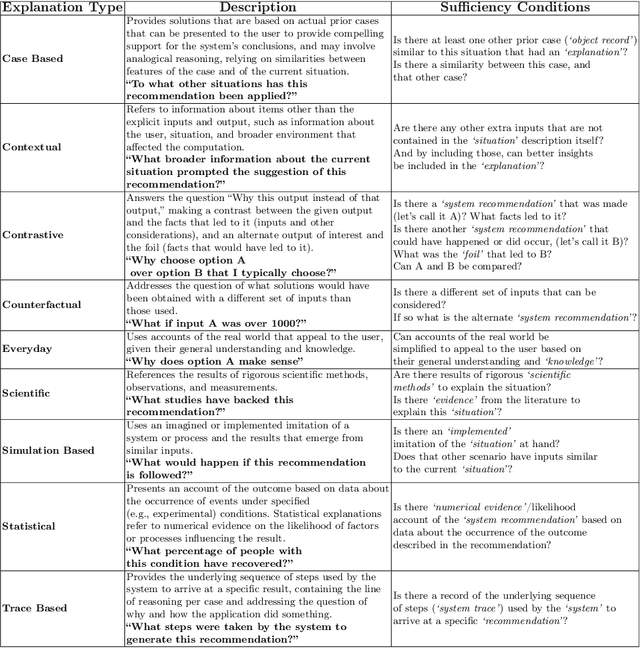
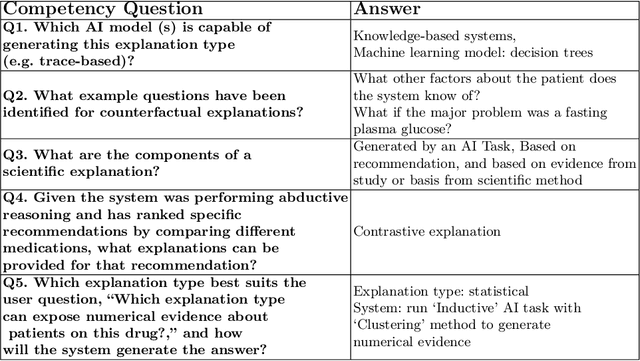
Abstract:Explainability has been a goal for Artificial Intelligence (AI) systems since their conception, with the need for explainability growing as more complex AI models are increasingly used in critical, high-stakes settings such as healthcare. Explanations have often added to an AI system in a non-principled, post-hoc manner. With greater adoption of these systems and emphasis on user-centric explainability, there is a need for a structured representation that treats explainability as a primary consideration, mapping end user needs to specific explanation types and the system's AI capabilities. We design an explanation ontology to model both the role of explanations, accounting for the system and user attributes in the process, and the range of different literature-derived explanation types. We indicate how the ontology can support user requirements for explanations in the domain of healthcare. We evaluate our ontology with a set of competency questions geared towards a system designer who might use our ontology to decide which explanation types to include, given a combination of users' needs and a system's capabilities, both in system design settings and in real-time operations. Through the use of this ontology, system designers will be able to make informed choices on which explanations AI systems can and should provide.
* 16 pages (but 1 reference over on arxiv), 5 tables, 3 code listings, 1 figure
Explanation Ontology in Action: A Clinical Use-Case
Oct 04, 2020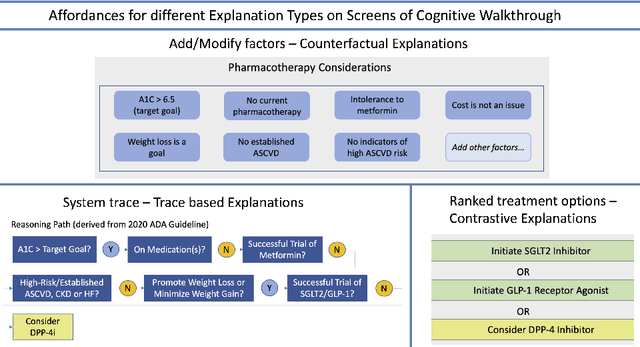
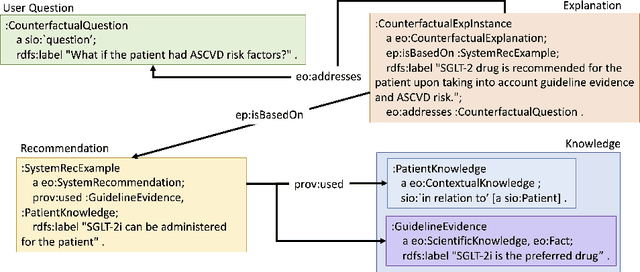
Abstract:We addressed the problem of a lack of semantic representation for user-centric explanations and different explanation types in our Explanation Ontology (https://purl.org/heals/eo). Such a representation is increasingly necessary as explainability has become an important problem in Artificial Intelligence with the emergence of complex methods and an uptake in high-precision and user-facing settings. In this submission, we provide step-by-step guidance for system designers to utilize our ontology, introduced in our resource track paper, to plan and model for explanations during the design of their Artificial Intelligence systems. We also provide a detailed example with our utilization of this guidance in a clinical setting.
* 5 pages, 2 figures, 1 protocol
Directions for Explainable Knowledge-Enabled Systems
Mar 17, 2020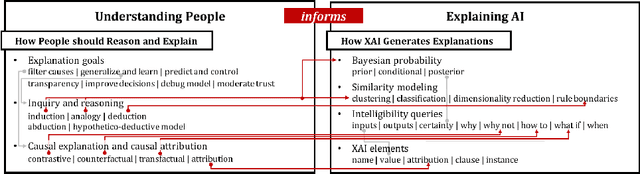
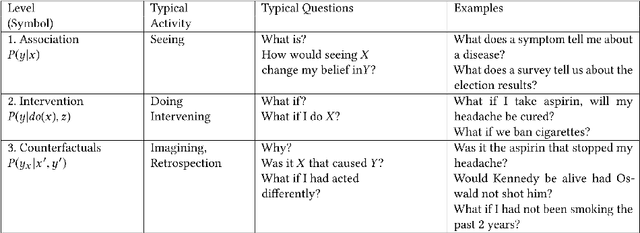
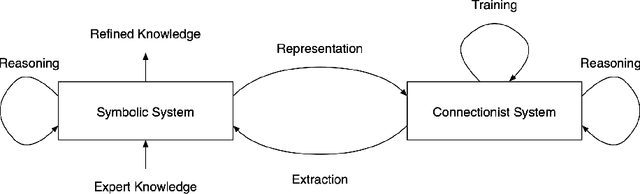
Abstract:Interest in the field of Explainable Artificial Intelligence has been growing for decades and has accelerated recently. As Artificial Intelligence models have become more complex, and often more opaque, with the incorporation of complex machine learning techniques, explainability has become more critical. Recently, researchers have been investigating and tackling explainability with a user-centric focus, looking for explanations to consider trustworthiness, comprehensibility, explicit provenance, and context-awareness. In this chapter, we leverage our survey of explanation literature in Artificial Intelligence and closely related fields and use these past efforts to generate a set of explanation types that we feel reflect the expanded needs of explanation for today's artificial intelligence applications. We define each type and provide an example question that would motivate the need for this style of explanation. We believe this set of explanation types will help future system designers in their generation and prioritization of requirements and further help generate explanations that are better aligned to users' and situational needs.
Foundations of Explainable Knowledge-Enabled Systems
Mar 17, 2020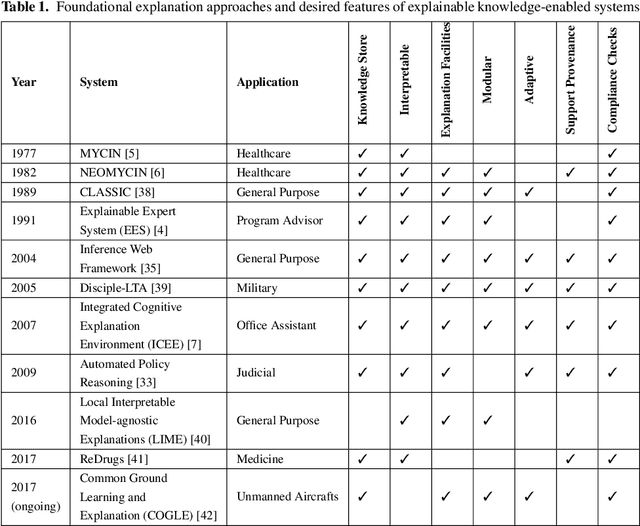
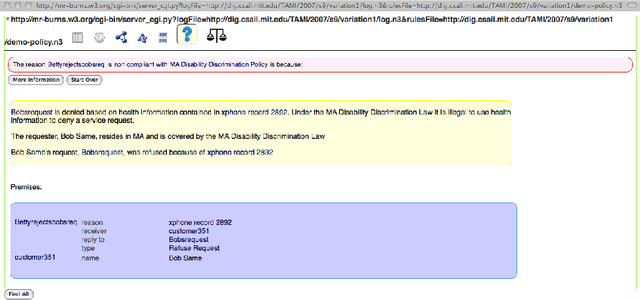
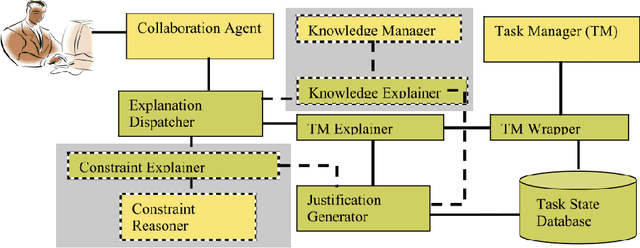
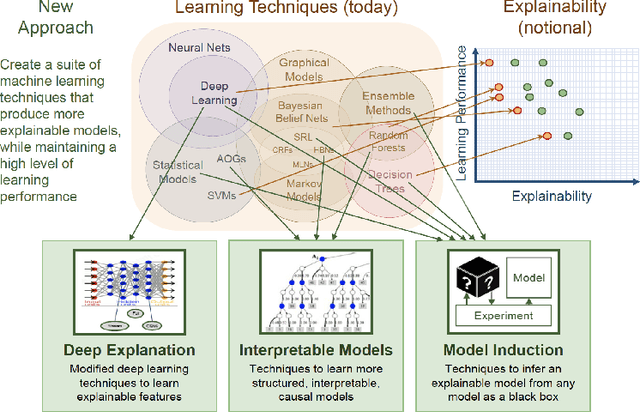
Abstract:Explainability has been an important goal since the early days of Artificial Intelligence. Several approaches for producing explanations have been developed. However, many of these approaches were tightly coupled with the capabilities of the artificial intelligence systems at the time. With the proliferation of AI-enabled systems in sometimes critical settings, there is a need for them to be explainable to end-users and decision-makers. We present a historical overview of explainable artificial intelligence systems, with a focus on knowledge-enabled systems, spanning the expert systems, cognitive assistants, semantic applications, and machine learning domains. Additionally, borrowing from the strengths of past approaches and identifying gaps needed to make explanations user- and context-focused, we propose new definitions for explanations and explainable knowledge-enabled systems.
Making Study Populations Visible through Knowledge Graphs
Jul 09, 2019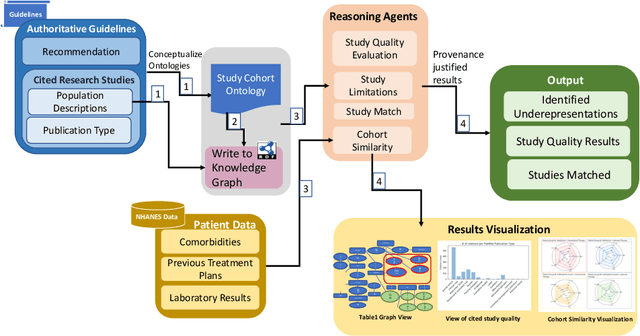

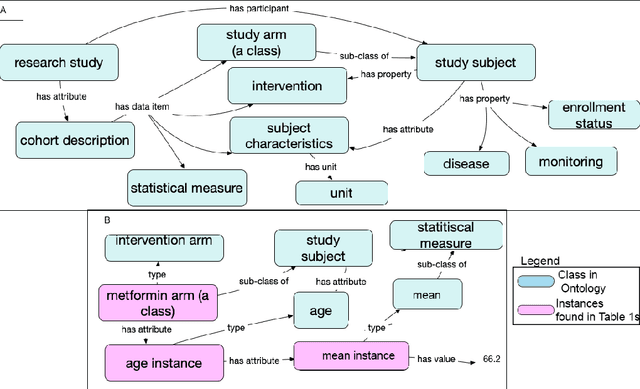
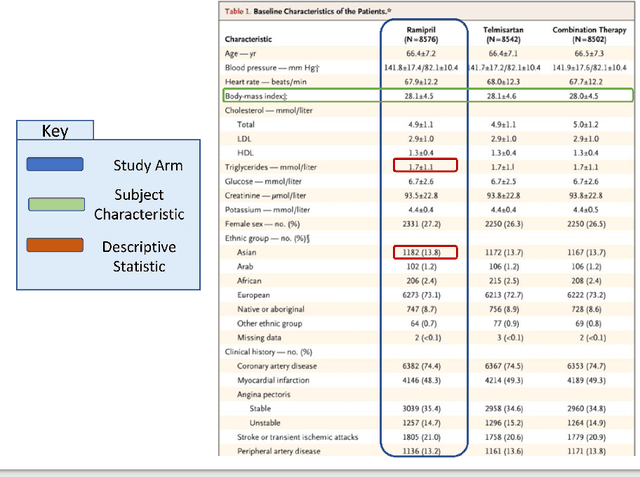
Abstract:Treatment recommendations within Clinical Practice Guidelines (CPGs) are largely based on findings from clinical trials and case studies, referred to here as research studies, that are often based on highly selective clinical populations, referred to here as study cohorts. When medical practitioners apply CPG recommendations, they need to understand how well their patient population matches the characteristics of those in the study cohort, and thus are confronted with the challenges of locating the study cohort information and making an analytic comparison. To address these challenges, we develop an ontology-enabled prototype system, which exposes the population descriptions in research studies in a declarative manner, with the ultimate goal of allowing medical practitioners to better understand the applicability and generalizability of treatment recommendations. We build a Study Cohort Ontology (SCO) to encode the vocabulary of study population descriptions, that are often reported in the first table in the published work, thus they are often referred to as Table 1. We leverage the well-used Semanticscience Integrated Ontology (SIO) for defining property associations between classes. Further, we model the key components of Table 1s, i.e., collections of study subjects, subject characteristics, and statistical measures in RDF knowledge graphs. We design scenarios for medical practitioners to perform population analysis, and generate cohort similarity visualizations to determine the applicability of a study population to the clinical population of interest. Our semantic approach to make study populations visible, by standardized representations of Table 1s, allows users to quickly derive clinically relevant inferences about study populations.
Semantically-aware population health risk analyses
Nov 27, 2018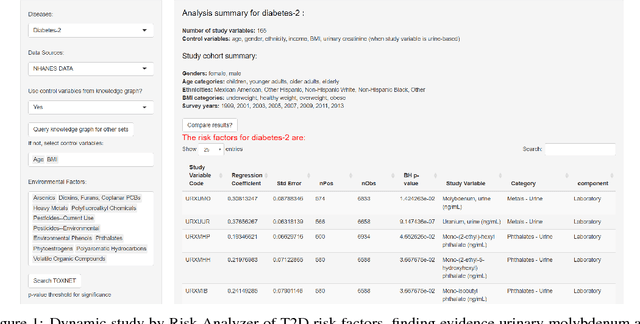

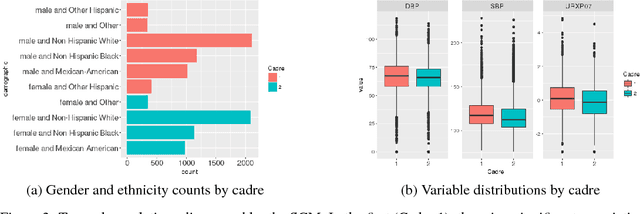
Abstract:One primary task of population health analysis is the identification of risk factors that, for some subpopulation, have a significant association with some health condition. Examples include finding lifestyle factors associated with chronic diseases and finding genetic mutations associated with diseases in precision health. We develop a combined semantic and machine learning system that uses a health risk ontology and knowledge graph (KG) to dynamically discover risk factors and their associated subpopulations. Semantics and the novel supervised cadre model make our system explainable. Future population health studies are easily performed and documented with provenance by specifying additional input and output KG cartridges.
Seq2RDF: An end-to-end application for deriving Triples from Natural Language Text
Aug 08, 2018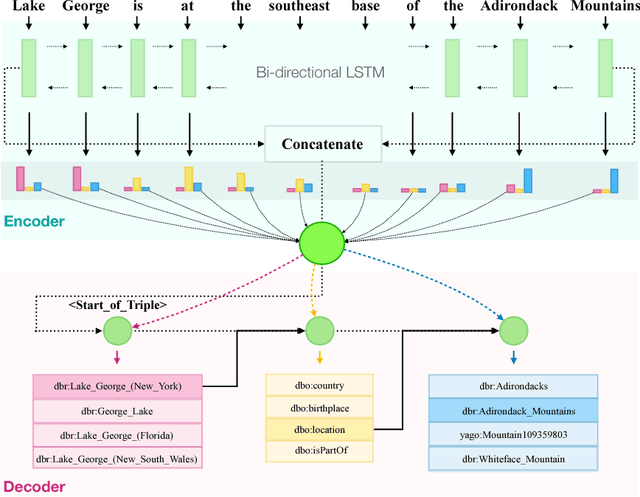
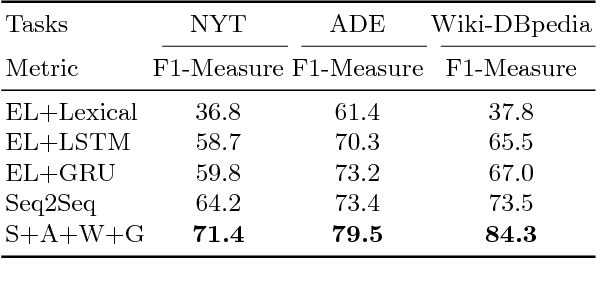
Abstract:We present an end-to-end approach that takes unstructured textual input and generates structured output compliant with a given vocabulary. Inspired by recent successes in neural machine translation, we treat the triples within a given knowledge graph as an independent graph language and propose an encoder-decoder framework with an attention mechanism that leverages knowledge graph embeddings. Our model learns the mapping from natural language text to triple representation in the form of subject-predicate-object using the selected knowledge graph vocabulary. Experiments on three different data sets show that we achieve competitive F1-Measures over the baselines using our simple yet effective approach. A demo video is included.
Knowledge Integration for Disease Characterization: A Breast Cancer Example
Jul 20, 2018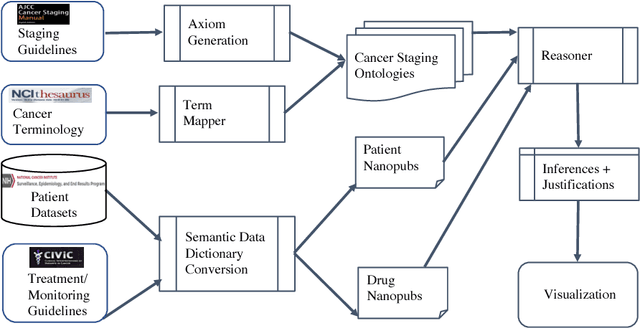
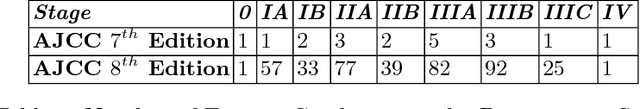
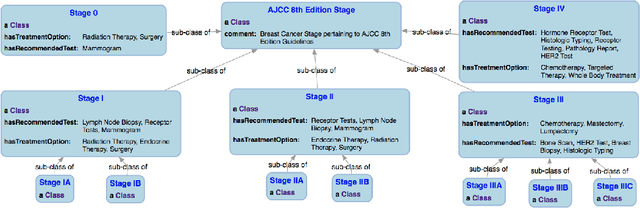
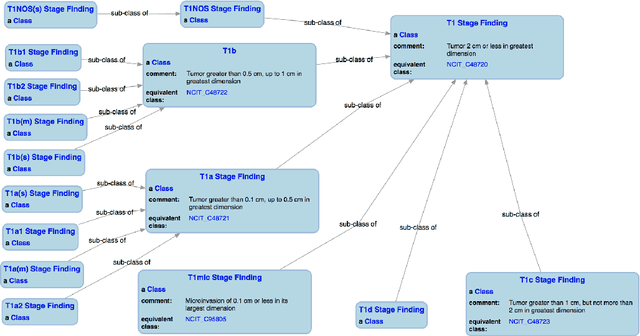
Abstract:With the rapid advancements in cancer research, the information that is useful for characterizing disease, staging tumors, and creating treatment and survivorship plans has been changing at a pace that creates challenges when physicians try to remain current. One example involves increasing usage of biomarkers when characterizing the pathologic prognostic stage of a breast tumor. We present our semantic technology approach to support cancer characterization and demonstrate it in our end-to-end prototype system that collects the newest breast cancer staging criteria from authoritative oncology manuals to construct an ontology for breast cancer. Using a tool we developed that utilizes this ontology, physician-facing applications can be used to quickly stage a new patient to support identifying risks, treatment options, and monitoring plans based on authoritative and best practice guidelines. Physicians can also re-stage existing patients or patient populations, allowing them to find patients whose stage has changed in a given patient cohort. As new guidelines emerge, using our proposed mechanism, which is grounded by semantic technologies for ingesting new data from staging manuals, we have created an enriched cancer staging ontology that integrates relevant data from several sources with very little human intervention.
Exploiting Task-Oriented Resources to Learn Word Embeddings for Clinical Abbreviation Expansion
Apr 11, 2018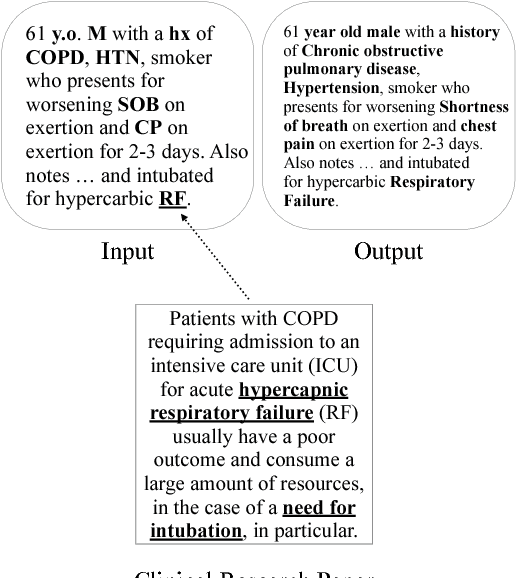
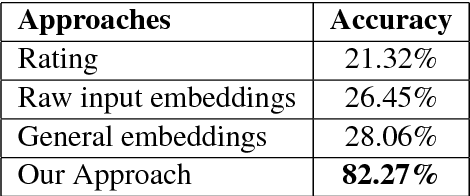
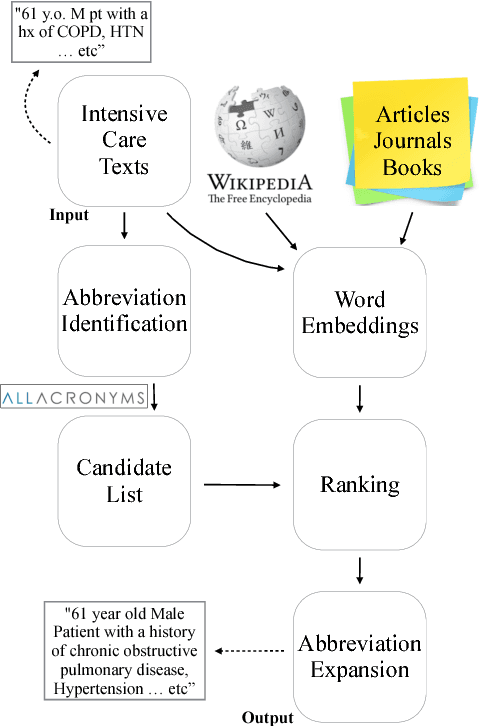
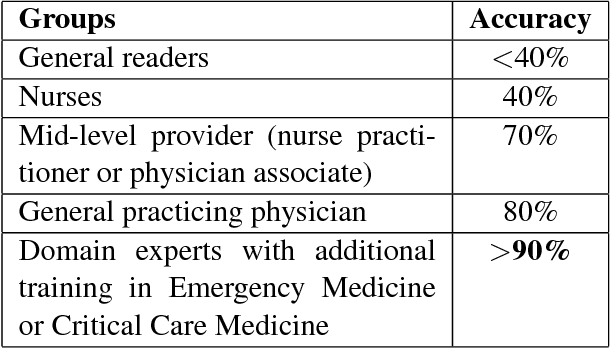
Abstract:In the medical domain, identifying and expanding abbreviations in clinical texts is a vital task for both better human and machine understanding. It is a challenging task because many abbreviations are ambiguous especially for intensive care medicine texts, in which phrase abbreviations are frequently used. Besides the fact that there is no universal dictionary of clinical abbreviations and no universal rules for abbreviation writing, such texts are difficult to acquire, expensive to annotate and even sometimes, confusing to domain experts. This paper proposes a novel and effective approach - exploiting task-oriented resources to learn word embeddings for expanding abbreviations in clinical notes. We achieved 82.27% accuracy, close to expert human performance.
From Data to City Indicators: A Knowledge Graph for Supporting Automatic Generation of Dashboards
Apr 06, 2017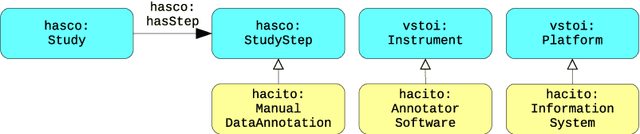
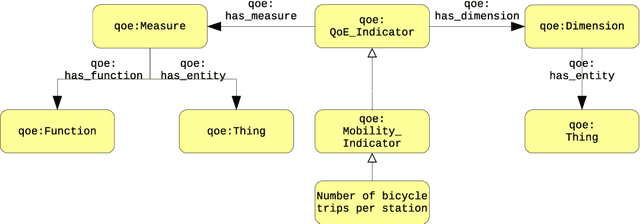
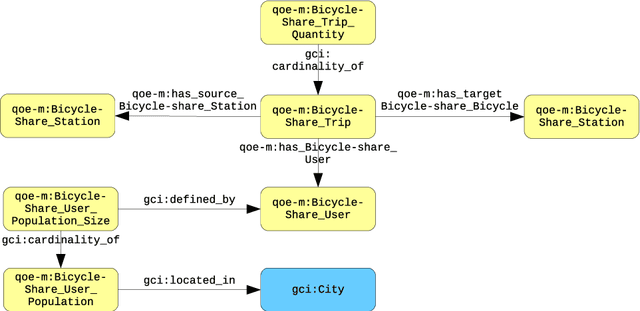
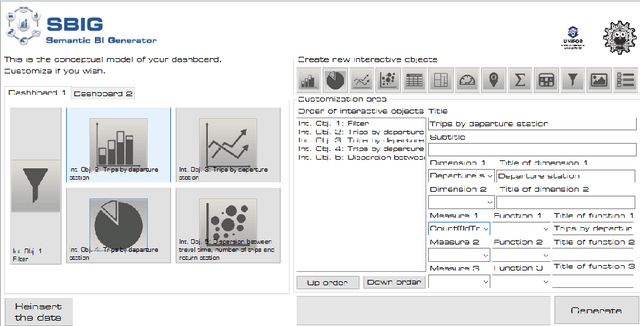
Abstract:In the context of Smart Cities, indicator definitions have been used to calculate values that enable the comparison among different cities. The calculation of an indicator values has challenges as the calculation may need to combine some aspects of quality while addressing different levels of abstraction. Knowledge graphs (KGs) have been used successfully to support flexible representation, which can support improved understanding and data analysis in similar settings. This paper presents an operational description for a city KG, an indicator ontology that support indicator discovery and data visualization and an application capable of performing metadata analysis to automatically build and display dashboards according to discovered indicators. We describe our implementation in an urban mobility setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge