David A. Klindt
Understanding Neural Coding on Latent Manifolds by Sharing Features and Dividing Ensembles
Oct 06, 2022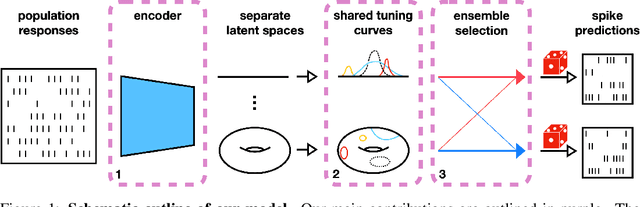
Abstract:Systems neuroscience relies on two complementary views of neural data, characterized by single neuron tuning curves and analysis of population activity. These two perspectives combine elegantly in neural latent variable models that constrain the relationship between latent variables and neural activity, modeled by simple tuning curve functions. This has recently been demonstrated using Gaussian processes, with applications to realistic and topologically relevant latent manifolds. Those and previous models, however, missed crucial shared coding properties of neural populations. We propose feature sharing across neural tuning curves, which significantly improves performance and leads to better-behaved optimization. We also propose a solution to the problem of ensemble detection, whereby different groups of neurons, i.e., ensembles, can be modulated by different latent manifolds. This is achieved through a soft clustering of neurons during training, thus allowing for the separation of mixed neural populations in an unsupervised manner. These innovations lead to more interpretable models of neural population activity that train well and perform better even on mixtures of complex latent manifolds. Finally, we apply our method on a recently published grid cell dataset, recovering distinct ensembles, inferring toroidal latents and predicting neural tuning curves all in a single integrated modeling framework.
Score-Based Generative Classifiers
Oct 01, 2021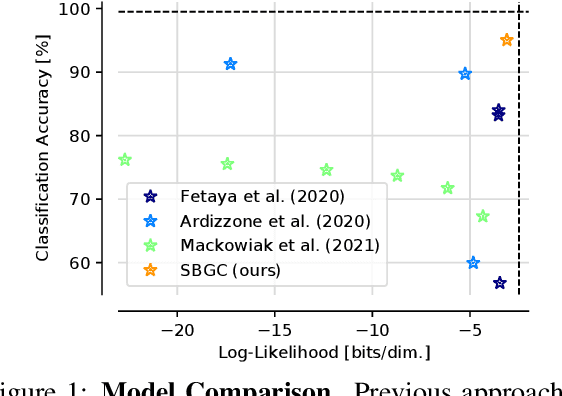
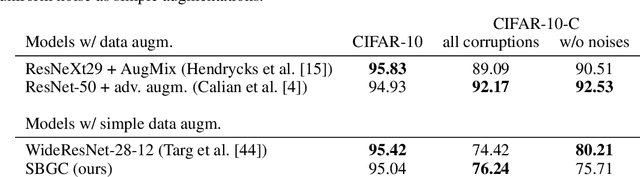
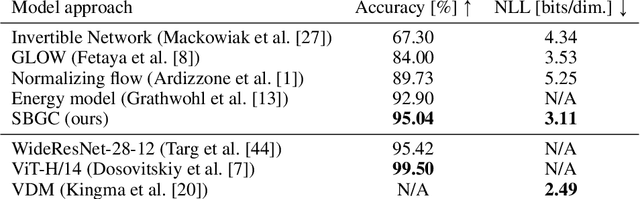

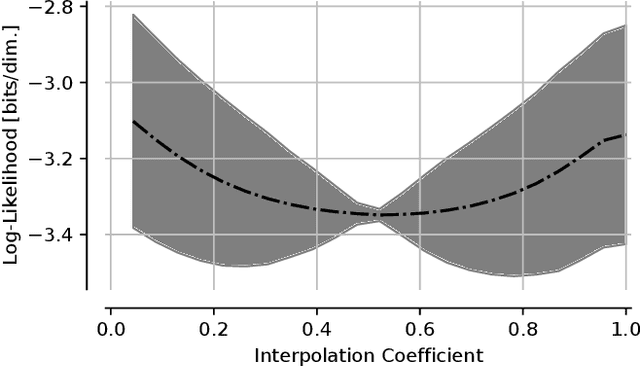
Abstract:The tremendous success of generative models in recent years raises the question whether they can also be used to perform classification. Generative models have been used as adversarially robust classifiers on simple datasets such as MNIST, but this robustness has not been observed on more complex datasets like CIFAR-10. Additionally, on natural image datasets, previous results have suggested a trade-off between the likelihood of the data and classification accuracy. In this work, we investigate score-based generative models as classifiers for natural images. We show that these models not only obtain competitive likelihood values but simultaneously achieve state-of-the-art classification accuracy for generative classifiers on CIFAR-10. Nevertheless, we find that these models are only slightly, if at all, more robust than discriminative baseline models on out-of-distribution tasks based on common image corruptions. Similarly and contrary to prior results, we find that score-based are prone to worst-case distribution shifts in the form of adversarial perturbations. Our work highlights that score-based generative models are closing the gap in classification accuracy compared to standard discriminative models. While they do not yet deliver on the promise of adversarial and out-of-domain robustness, they provide a different approach to classification that warrants further research.
Neural system identification for large populations separating "what" and "where"
Jan 29, 2018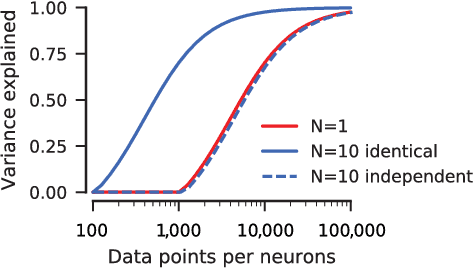
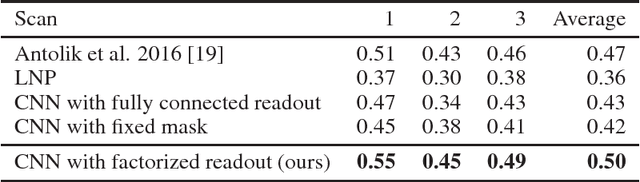
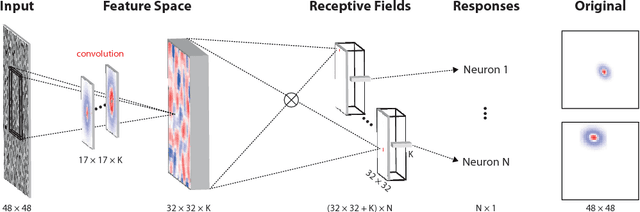
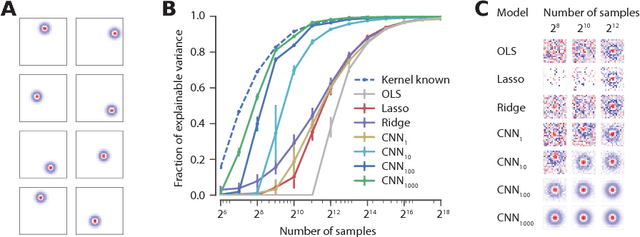
Abstract:Neuroscientists classify neurons into different types that perform similar computations at different locations in the visual field. Traditional methods for neural system identification do not capitalize on this separation of 'what' and 'where'. Learning deep convolutional feature spaces that are shared among many neurons provides an exciting path forward, but the architectural design needs to account for data limitations: While new experimental techniques enable recordings from thousands of neurons, experimental time is limited so that one can sample only a small fraction of each neuron's response space. Here, we show that a major bottleneck for fitting convolutional neural networks (CNNs) to neural data is the estimation of the individual receptive field locations, a problem that has been scratched only at the surface thus far. We propose a CNN architecture with a sparse readout layer factorizing the spatial (where) and feature (what) dimensions. Our network scales well to thousands of neurons and short recordings and can be trained end-to-end. We evaluate this architecture on ground-truth data to explore the challenges and limitations of CNN-based system identification. Moreover, we show that our network model outperforms current state-of-the art system identification models of mouse primary visual cortex.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge