Daphne Koller
Stanford University
Learning Factor Graphs in Polynomial Time & Sample Complexity
Jul 04, 2012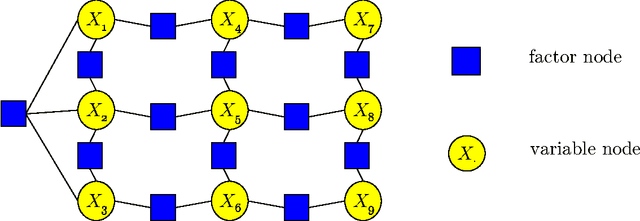
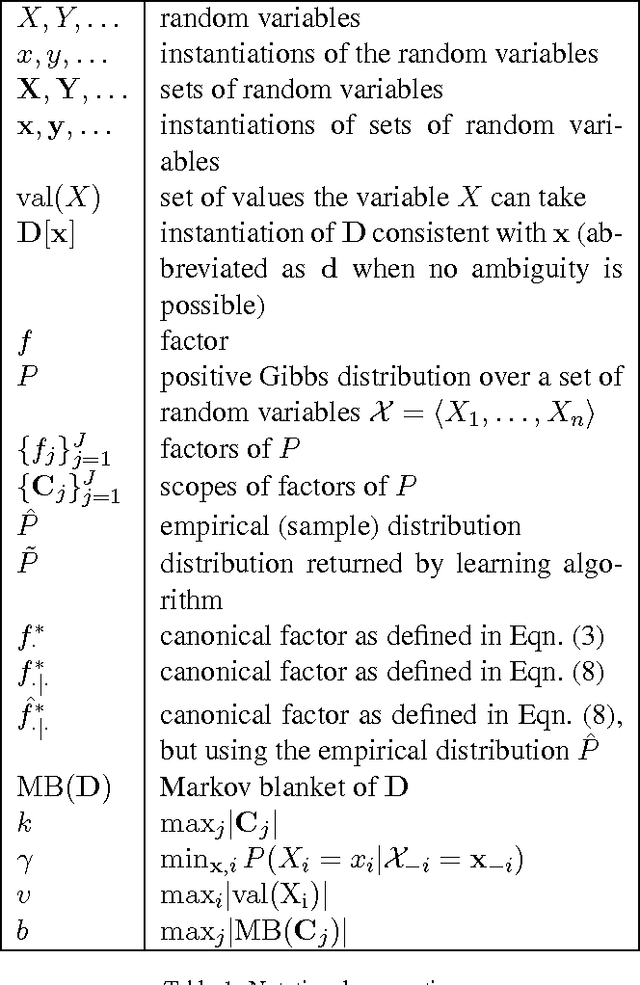

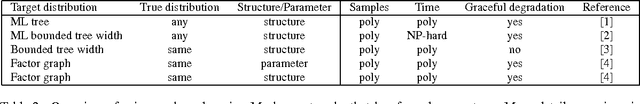
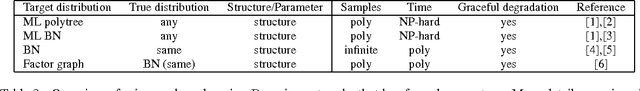
Abstract:We study computational and sample complexity of parameter and structure learning in graphical models. Our main result shows that the class of factor graphs with bounded factor size and bounded connectivity can be learned in polynomial time and polynomial number of samples, assuming that the data is generated by a network in this class. This result covers both parameter estimation for a known network structure and structure learning. It implies as a corollary that we can learn factor graphs for both Bayesian networks and Markov networks of bounded degree, in polynomial time and sample complexity. Unlike maximum likelihood estimation, our method does not require inference in the underlying network, and so applies to networks where inference is intractable. We also show that the error of our learned model degrades gracefully when the generating distribution is not a member of the target class of networks.
Continuous Time Markov Networks
Jun 27, 2012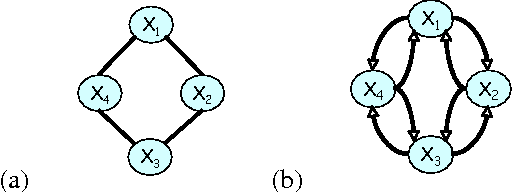
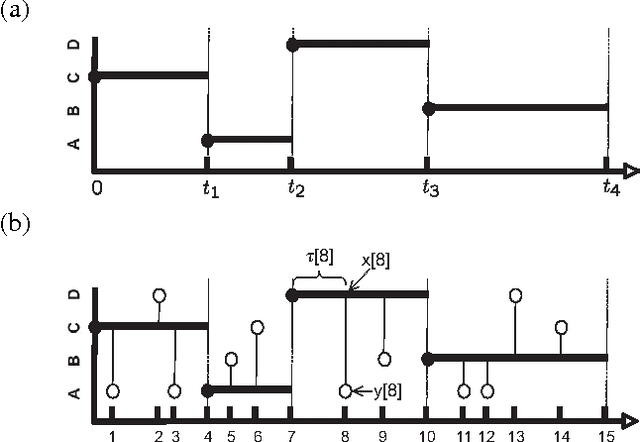
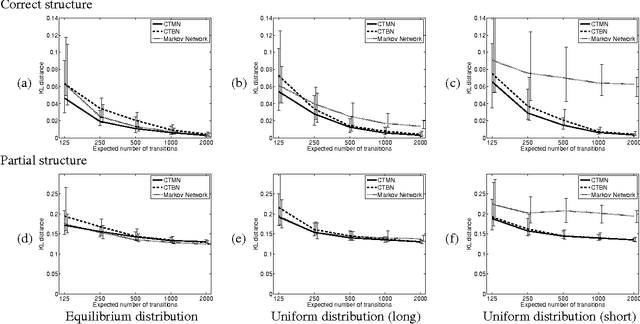
Abstract:A central task in many applications is reasoning about processes that change in a continuous time. The mathematical framework of Continuous Time Markov Processes provides the basic foundations for modeling such systems. Recently, Nodelman et al introduced continuous time Bayesian networks (CTBNs), which allow a compact representation of continuous-time processes over a factored state space. In this paper, we introduce continuous time Markov networks (CTMNs), an alternative representation language that represents a different type of continuous-time dynamics. In many real life processes, such as biological and chemical systems, the dynamics of the process can be naturally described as an interplay between two forces - the tendency of each entity to change its state, and the overall fitness or energy function of the entire system. In our model, the first force is described by a continuous-time proposal process that suggests possible local changes to the state of the system at different rates. The second force is represented by a Markov network that encodes the fitness, or desirability, of different states; a proposed local change is then accepted with a probability that is a function of the change in the fitness distribution. We show that the fitness distribution is also the stationary distribution of the Markov process, so that this representation provides a characterization of a temporal process whose stationary distribution has a compact graphical representation. This allows us to naturally capture a different type of structure in complex dynamical processes, such as evolving biological sequences. We describe the semantics of the representation, its basic properties, and how it compares to CTBNs. We also provide algorithms for learning such models from data, and discuss its applicability to biological sequence evolution.
Residual Belief Propagation: Informed Scheduling for Asynchronous Message Passing
Jun 27, 2012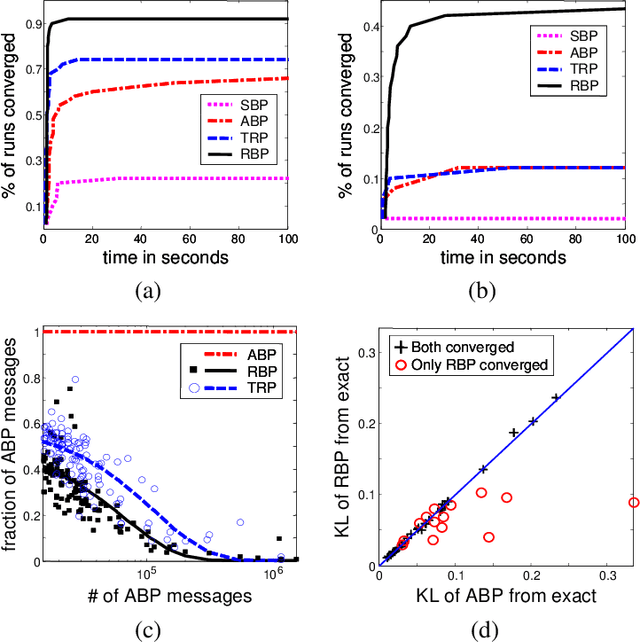
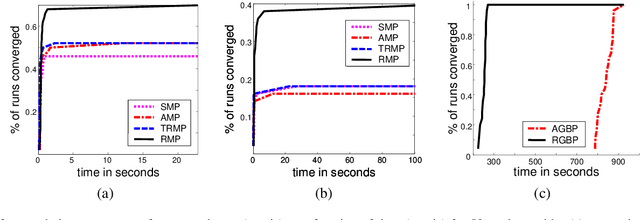
Abstract:Inference for probabilistic graphical models is still very much a practical challenge in large domains. The commonly used and effective belief propagation (BP) algorithm and its generalizations often do not converge when applied to hard, real-life inference tasks. While it is widely recognized that the scheduling of messages in these algorithms may have significant consequences, this issue remains largely unexplored. In this work, we address the question of how to schedule messages for asynchronous propagation so that a fixed point is reached faster and more often. We first show that any reasonable asynchronous BP converges to a unique fixed point under conditions similar to those that guarantee convergence of synchronous BP. In addition, we show that the convergence rate of a simple round-robin schedule is at least as good as that of synchronous propagation. We then propose residual belief propagation (RBP), a novel, easy-to-implement, asynchronous propagation algorithm that schedules messages in an informed way, that pushes down a bound on the distance from the fixed point. Finally, we demonstrate the superiority of RBP over state-of-the-art methods for a variety of challenging synthetic and real-life problems: RBP converges significantly more often than other methods; and it significantly reduces running time until convergence, even when other methods converge.
Reasoning at the Right Time Granularity
Jun 20, 2012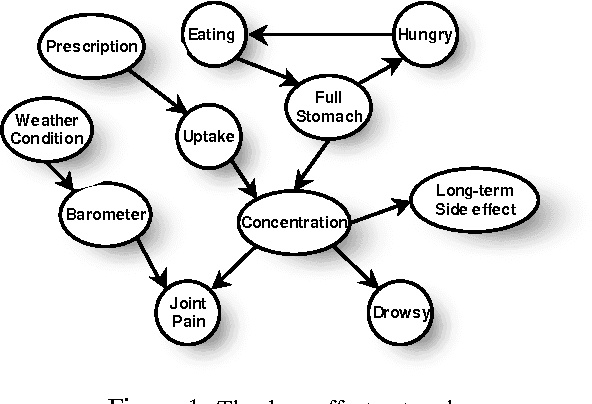
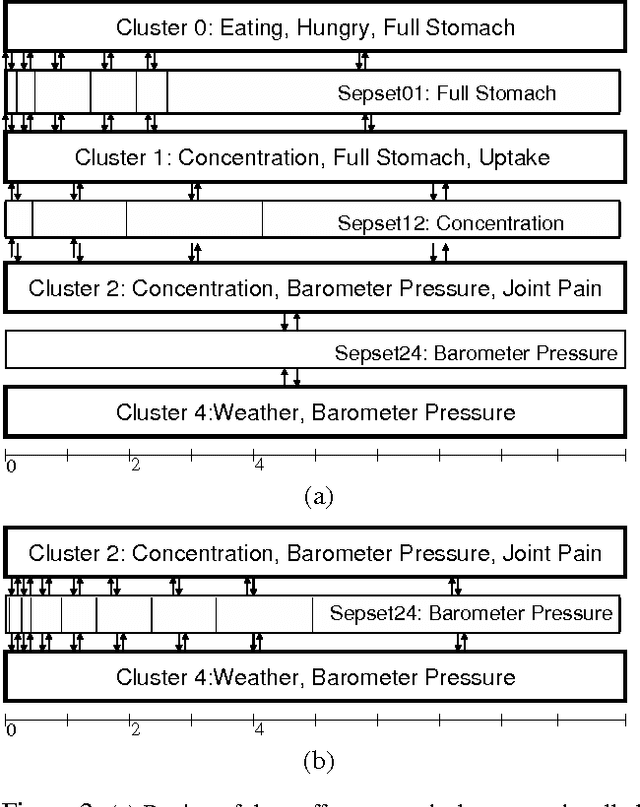
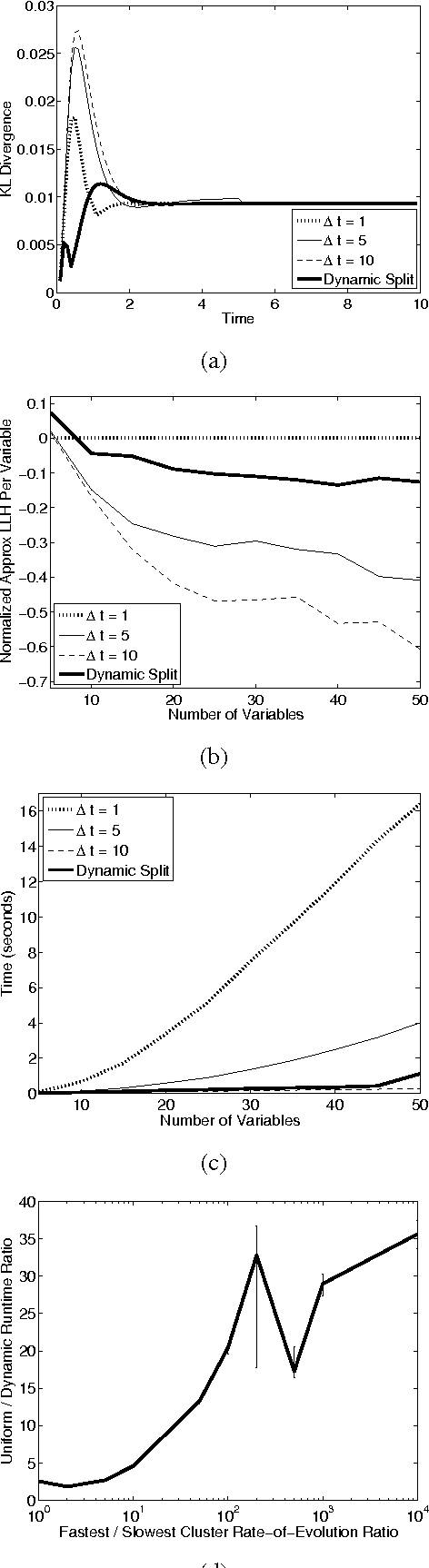
Abstract:Most real-world dynamic systems are composed of different components that often evolve at very different rates. In traditional temporal graphical models, such as dynamic Bayesian networks, time is modeled at a fixed granularity, generally selected based on the rate at which the fastest component evolves. Inference must then be performed at this fastest granularity, potentially at significant computational cost. Continuous Time Bayesian Networks (CTBNs) avoid time-slicing in the representation by modeling the system as evolving continuously over time. The expectation-propagation (EP) inference algorithm of Nodelman et al. (2005) can then vary the inference granularity over time, but the granularity is uniform across all parts of the system, and must be selected in advance. In this paper, we provide a new EP algorithm that utilizes a general cluster graph architecture where clusters contain distributions that can overlap in both space (set of variables) and time. This architecture allows different parts of the system to be modeled at very different time granularities, according to their current rate of evolution. We also provide an information-theoretic criterion for dynamically re-partitioning the clusters during inference to tune the level of approximation to the current rate of evolution. This avoids the need to hand-select the appropriate granularity, and allows the granularity to adapt as information is transmitted across the network. We present experiments demonstrating that this approach can result in significant computational savings.
Modeling Latent Variable Uncertainty for Loss-based Learning
Jun 18, 2012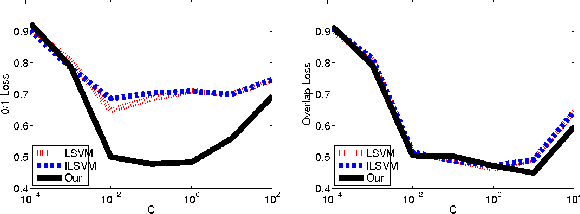
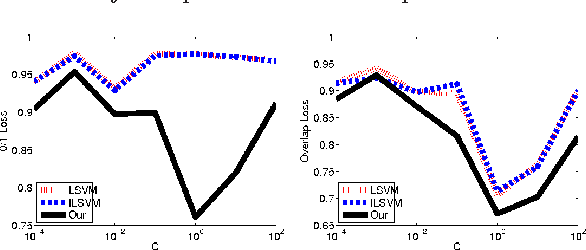
Abstract:We consider the problem of parameter estimation using weakly supervised datasets, where a training sample consists of the input and a partially specified annotation, which we refer to as the output. The missing information in the annotation is modeled using latent variables. Previous methods overburden a single distribution with two separate tasks: (i) modeling the uncertainty in the latent variables during training; and (ii) making accurate predictions for the output and the latent variables during testing. We propose a novel framework that separates the demands of the two tasks using two distributions: (i) a conditional distribution to model the uncertainty of the latent variables for a given input-output pair; and (ii) a delta distribution to predict the output and the latent variables for a given input. During learning, we encourage agreement between the two distributions by minimizing a loss-based dissimilarity coefficient. Our approach generalizes latent SVM in two important ways: (i) it models the uncertainty over latent variables instead of relying on a pointwise estimate; and (ii) it allows the use of loss functions that depend on latent variables, which greatly increases its applicability. We demonstrate the efficacy of our approach on two challenging problems---object detection and action detection---using publicly available datasets.
Constrained Approximate Maximum Entropy Learning of Markov Random Fields
Jun 13, 2012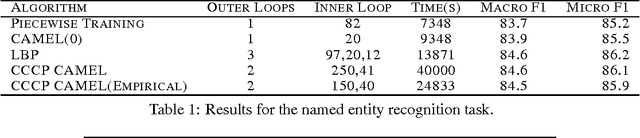
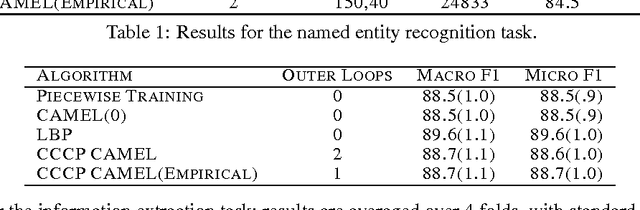

Abstract:Parameter estimation in Markov random fields (MRFs) is a difficult task, in which inference over the network is run in the inner loop of a gradient descent procedure. Replacing exact inference with approximate methods such as loopy belief propagation (LBP) can suffer from poor convergence. In this paper, we provide a different approach for combining MRF learning and Bethe approximation. We consider the dual of maximum likelihood Markov network learning - maximizing entropy with moment matching constraints - and then approximate both the objective and the constraints in the resulting optimization problem. Unlike previous work along these lines (Teh & Welling, 2003), our formulation allows parameter sharing between features in a general log-linear model, parameter regularization and conditional training. We show that piecewise training (Sutton & McCallum, 2005) is a very restricted special case of this formulation. We study two optimization strategies: one based on a single convex approximation and one that uses repeated convex approximations. We show results on several real-world networks that demonstrate that these algorithms can significantly outperform learning with loopy and piecewise. Our results also provide a framework for analyzing the trade-offs of different relaxations of the entropy objective and of the constraints.
Convex Point Estimation using Undirected Bayesian Transfer Hierarchies
Jun 13, 2012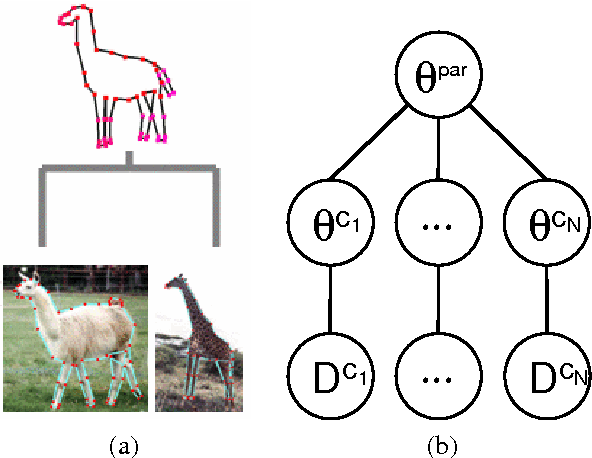
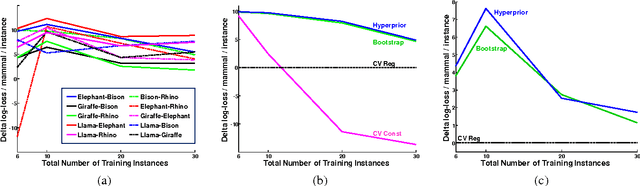
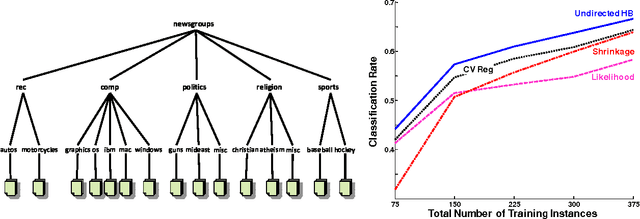
Abstract:When related learning tasks are naturally arranged in a hierarchy, an appealing approach for coping with scarcity of instances is that of transfer learning using a hierarchical Bayes framework. As fully Bayesian computations can be difficult and computationally demanding, it is often desirable to use posterior point estimates that facilitate (relatively) efficient prediction. However, the hierarchical Bayes framework does not always lend itself naturally to this maximum aposteriori goal. In this work we propose an undirected reformulation of hierarchical Bayes that relies on priors in the form of similarity measures. We introduce the notion of "degree of transfer" weights on components of these similarity measures, and show how they can be automatically learned within a joint probabilistic framework. Importantly, our reformulation results in a convex objective for many learning problems, thus facilitating optimal posterior point estimation using standard optimization techniques. In addition, we no longer require proper priors, allowing for flexible and straightforward specification of joint distributions over transfer hierarchies. We show that our framework is effective for learning models that are part of transfer hierarchies for two real-life tasks: object shape modeling using Gaussian density estimation and document classification.
Projected Subgradient Methods for Learning Sparse Gaussians
Jun 13, 2012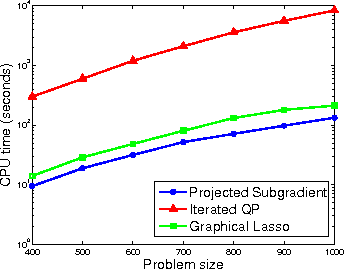
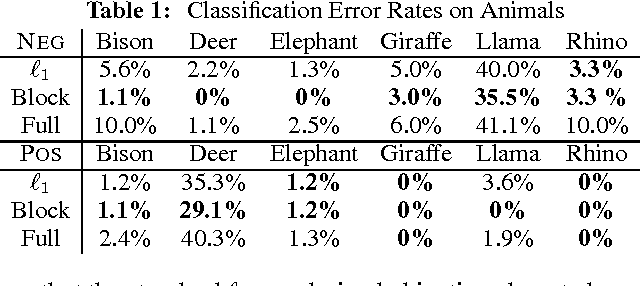
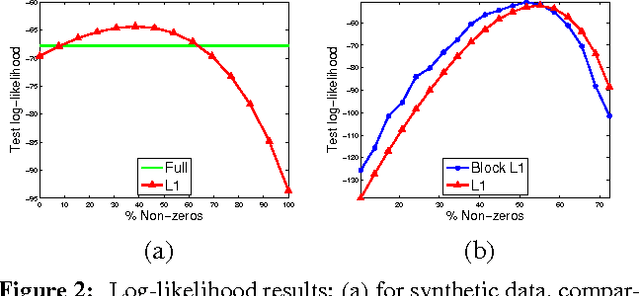
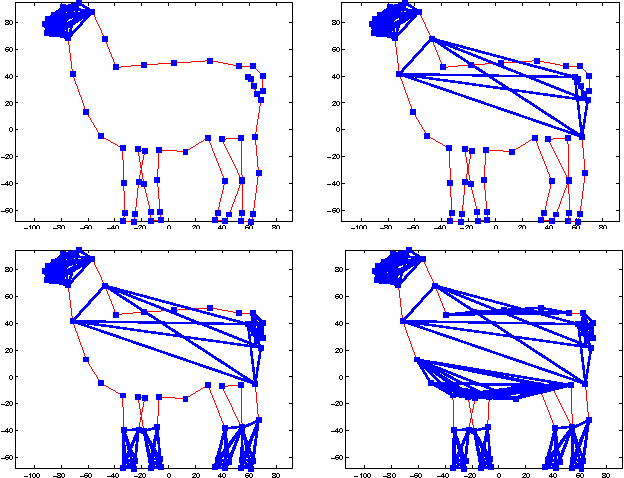
Abstract:Gaussian Markov random fields (GMRFs) are useful in a broad range of applications. In this paper we tackle the problem of learning a sparse GMRF in a high-dimensional space. Our approach uses the l1-norm as a regularization on the inverse covariance matrix. We utilize a novel projected gradient method, which is faster than previous methods in practice and equal to the best performing of these in asymptotic complexity. We also extend the l1-regularized objective to the problem of sparsifying entire blocks within the inverse covariance matrix. Our methods generalize fairly easily to this case, while other methods do not. We demonstrate that our extensions give better generalization performance on two real domains--biological network analysis and a 2D-shape modeling image task.
MAP Estimation of Semi-Metric MRFs via Hierarchical Graph Cuts
May 09, 2012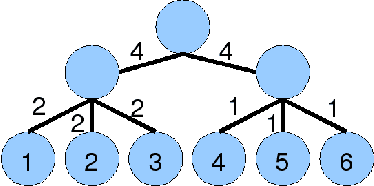
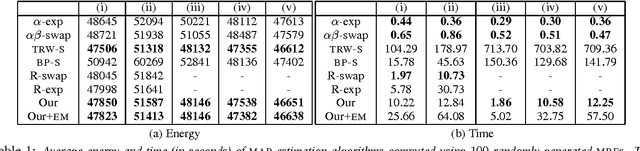


Abstract:We consider the task of obtaining the maximum a posteriori estimate of discrete pairwise random fields with arbitrary unary potentials and semimetric pairwise potentials. For this problem, we propose an accurate hierarchical move making strategy where each move is computed efficiently by solving an st-MINCUT problem. Unlike previous move making approaches, e.g. the widely used a-expansion algorithm, our method obtains the guarantees of the standard linear programming (LP) relaxation for the important special case of metric labeling. Unlike the existing LP relaxation solvers, e.g. interior-point algorithms or tree-reweighted message passing, our method is significantly faster as it uses only the efficient st-MINCUT algorithm in its design. Using both synthetic and real data experiments, we show that our technique outperforms several commonly used algorithms.
Discovering shared and individual latent structure in multiple time series
Aug 12, 2010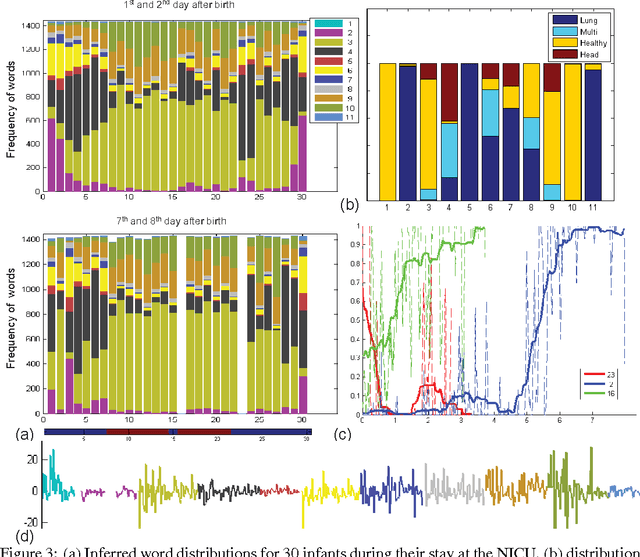
Abstract:This paper proposes a nonparametric Bayesian method for exploratory data analysis and feature construction in continuous time series. Our method focuses on understanding shared features in a set of time series that exhibit significant individual variability. Our method builds on the framework of latent Diricihlet allocation (LDA) and its extension to hierarchical Dirichlet processes, which allows us to characterize each series as switching between latent ``topics'', where each topic is characterized as a distribution over ``words'' that specify the series dynamics. However, unlike standard applications of LDA, we discover the words as we learn the model. We apply this model to the task of tracking the physiological signals of premature infants; our model obtains clinically significant insights as well as useful features for supervised learning tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge