Christian Marzahl
Dataset on Bi- and Multi-Nucleated Tumor Cells in Canine Cutaneous Mast Cell Tumors
Jan 05, 2021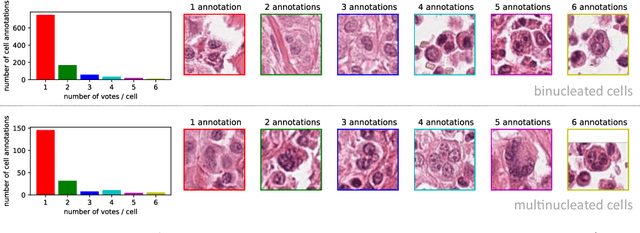
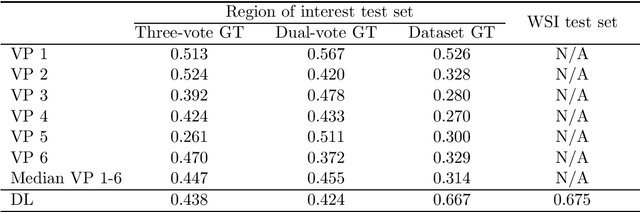
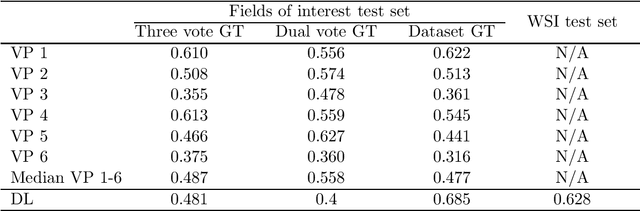
Abstract:Tumor cells with two nuclei (binucleated cells, BiNC) or more nuclei (multinucleated cells, MuNC) indicate an increased amount of cellular genetic material which is thought to facilitate oncogenesis, tumor progression and treatment resistance. In canine cutaneous mast cell tumors (ccMCT), binucleation and multinucleation are parameters used in cytologic and histologic grading schemes (respectively) which correlate with poor patient outcome. For this study, we created the first open source data-set with 19,983 annotations of BiNC and 1,416 annotations of MuNC in 32 histological whole slide images of ccMCT. Labels were created by a pathologist and an algorithmic-aided labeling approach with expert review of each generated candidate. A state-of-the-art deep learning-based model yielded an $F_1$ score of 0.675 for BiNC and 0.623 for MuNC on 11 test whole slide images. In regions of interest ($2.37 mm^2$) extracted from these test images, 6 pathologists had an object detection performance between 0.270 - 0.526 for BiNC and 0.316 - 0.622 for MuNC, while our model archived an $F_1$ score of 0.667 for BiNC and 0.685 for MuNC. This open dataset can facilitate development of automated image analysis for this task and may thereby help to promote standardization of this facet of histologic tumor prognostication.
Dogs as Model for Human Breast Cancer: A Completely Annotated Whole Slide Image Dataset
Aug 24, 2020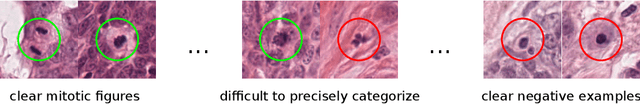
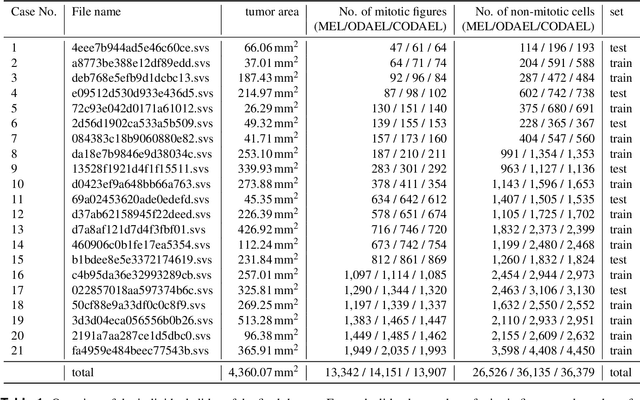
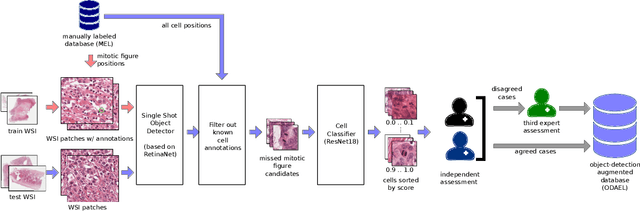
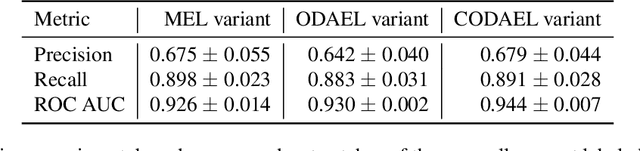
Abstract:Canine mammary carcinoma (CMC) has been used as a model to investigate the pathogenesis of human breast cancer and the same grading scheme is commonly used to assess tumor malignancy in both. One key component of this grading scheme is the density of mitotic figures (MF). Current publicly available datasets on human breast cancer only provide annotations for small subsets of whole slide images (WSIs). We present a novel dataset of 21 WSIs of CMC completely annotated for MF. For this, a pathologist screened all WSIs for potential MF and structures with a similar appearance. A second expert blindly assigned labels, and for non-matching labels, a third expert assigned the final labels. Additionally, we used machine learning to identify previously undetected MF. Finally, we performed representation learning and two-dimensional projection to further increase the consistency of the annotations. Our dataset consists of 13,907 MF and 36,379 hard negatives. We achieved a mean F1-score of 0.791 on the test set and of up to 0.696 on a human breast cancer dataset.
Are pathologist-defined labels reproducible? Comparison of the TUPAC16 mitotic figure dataset with an alternative set of labels
Jul 10, 2020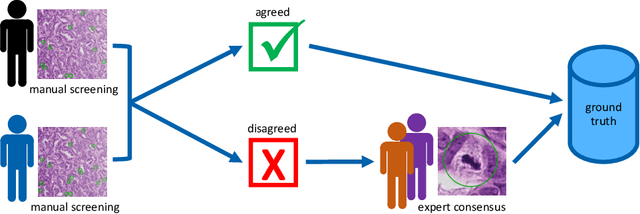
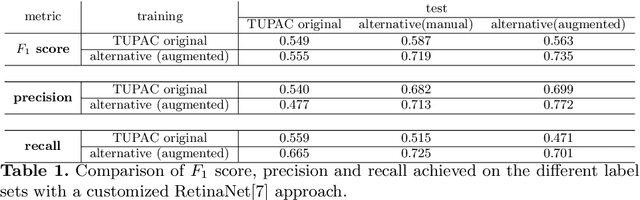
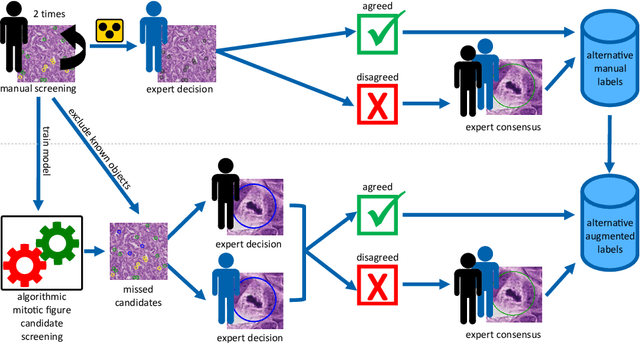
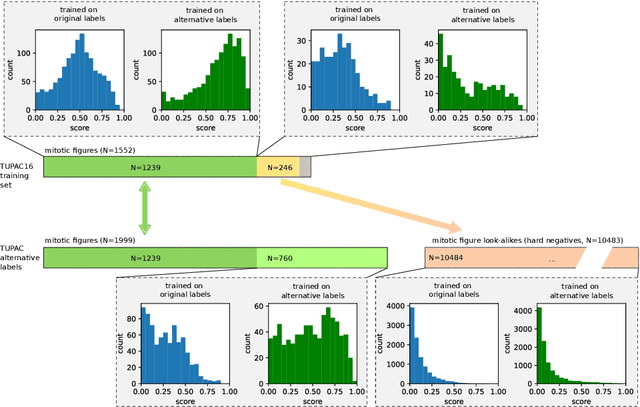
Abstract:Pathologist-defined labels are the gold standard for histopathological data sets, regardless of well-known limitations in consistency for some tasks. To date, some datasets on mitotic figures are available and were used for development of promising deep learning-based algorithms. In order to assess robustness of those algorithms and reproducibility of their methods it is necessary to test on several independent datasets. The influence of different labeling methods of these available datasets is currently unknown. To tackle this, we present an alternative set of labels for the images of the auxiliary mitosis dataset of the TUPAC16 challenge. Additional to manual mitotic figure screening, we used a novel, algorithm-aided labeling process, that allowed to minimize the risk of missing rare mitotic figures in the images. All potential mitotic figures were independently assessed by two pathologists. The novel, publicly available set of labels contains 1,999 mitotic figures (+28.80%) and additionally includes 10,483 labels of cells with high similarities to mitotic figures (hard examples). We found significant difference comparing F_1 scores between the original label set (0.549) and the new alternative label set (0.735) using a standard deep learning object detection architecture. The models trained on the alternative set showed higher overall confidence values, suggesting a higher overall label consistency. Findings of the present study show that pathologists-defined labels may vary significantly resulting in notable difference in the model performance. Comparison of deep learning-based algorithms between independent datasets with different labeling methods should be done with caution.
EXACT: A collaboration toolset for algorithm-aided annotation of almost everything
Apr 30, 2020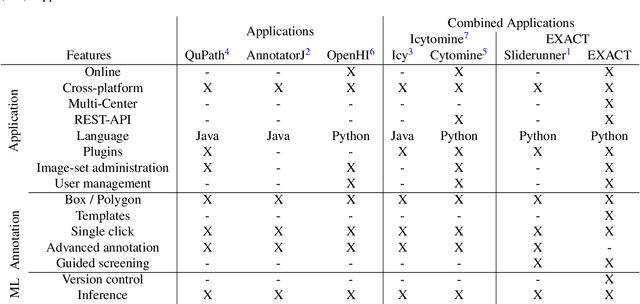
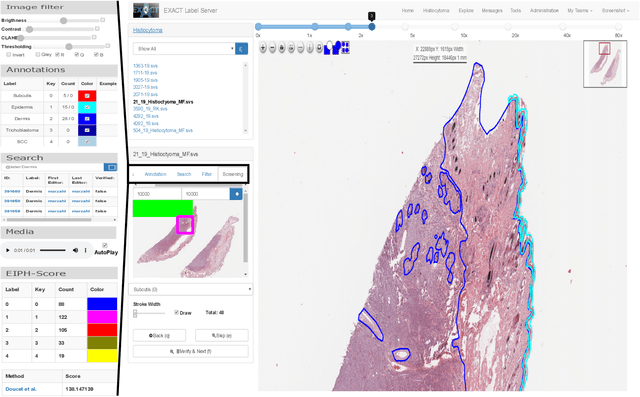
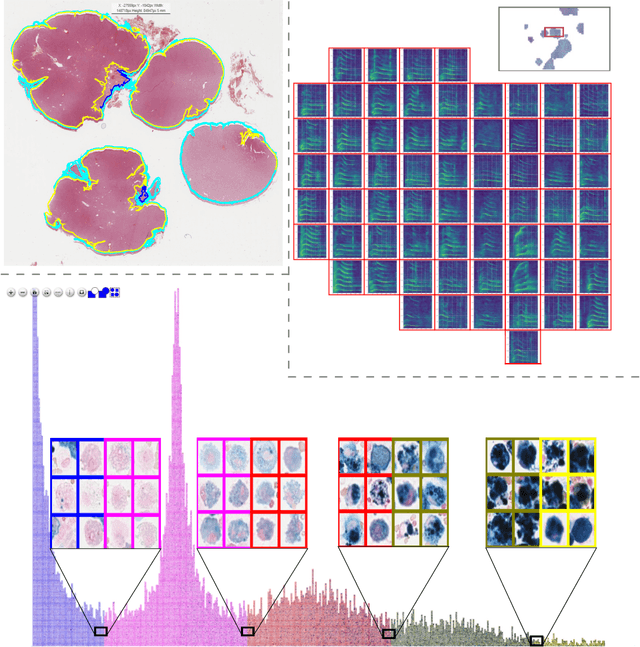
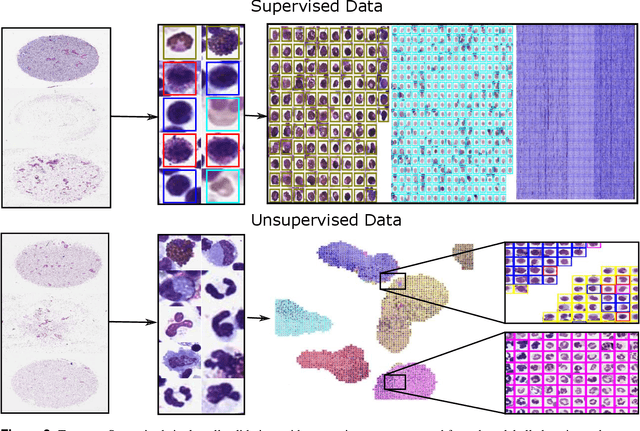
Abstract:In many research areas scientific progress is accelerated by multidisciplinary access to image data and their interdisciplinary annotation. However, keeping track of these annotations to ensure a high-quality multi purpose data set is a challenging and labour intensive task. We developed the open-source online platform EXACT (EXpert Algorithm Cooperation Tool) that enables the collaborative interdisciplinary analysis of images from different domains online and offline. EXACT supports multi-gigapixel whole slide medical images, as well as image series with thousands of images. The software utilises a flexible plugin system that can be adapted to diverse applications such as counting mitotic figures with the screening mode, finding false annotations on a novel validation view, or using the latest deep learning image analysis technologies. This is combined with a version control system which makes it possible to keep track of changes in data sets and, for example, to link the results of deep learning experiments to specific data set versions. EXACT is freely available and has been applied successfully to a broad range of annotation tasks already, including highly diverse applications like deep learning supported cytology grading, interdisciplinary multi-centre whole slide image tumour annotation, and highly specialised whale sound spectroscopy clustering.
Learning New Tricks from Old Dogs -- Inter-Species, Inter-Tissue Domain Adaptation for Mitotic Figure Assessment
Nov 25, 2019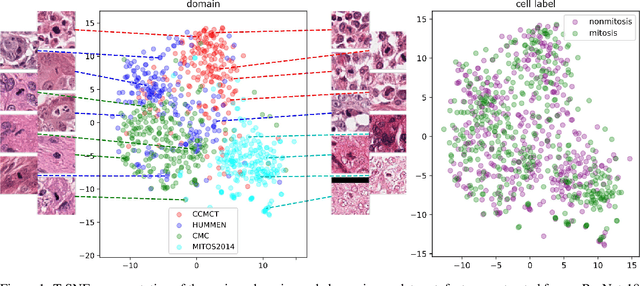
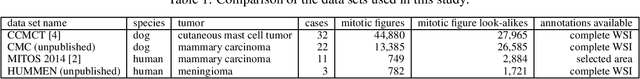
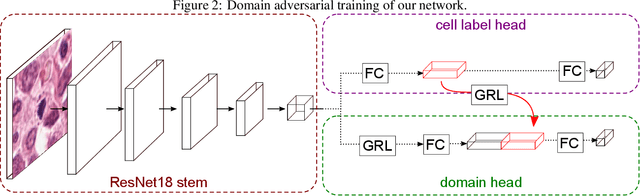
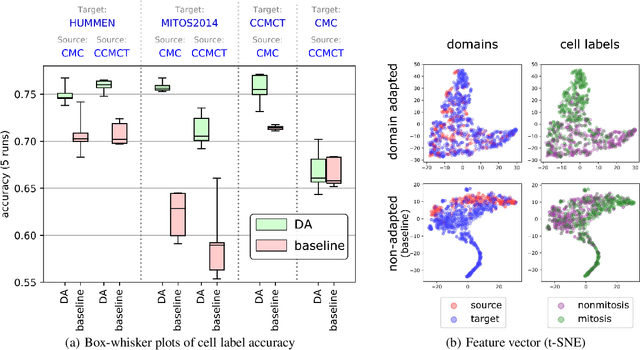
Abstract:For histopathological tumor assessment, the count of mitotic figures per area is an important part of prognostication. Algorithmic approaches - such as for mitotic figure identification - have significantly improved in recent times, potentially allowing for computer-augmented or fully automatic screening systems in the future. This trend is further supported by whole slide scanning microscopes becoming available in many pathology labs and could soon become a standard imaging tool. For an application in broader fields of such algorithms, the availability of mitotic figure data sets of sufficient size for the respective tissue type and species is an important precondition, that is, however, rarely met. While algorithmic performance climbed steadily for e.g. human mammary carcinoma, thanks to several challenges held in the field, for most tumor types, data sets are not available. In this work, we assess domain transfer of mitotic figure recognition using domain adversarial training on four data sets, two from dogs and two from humans. We were able to show that domain adversarial training considerably improves accuracy when applying mitotic figure classification learned from the canine on the human data sets (up to +12.8% in accuracy) and is thus a helpful method to transfer knowledge from existing data sets to new tissue types and species.
Deep Learning-Based Quantification of Pulmonary Hemosiderophages in Cytology Slides
Aug 12, 2019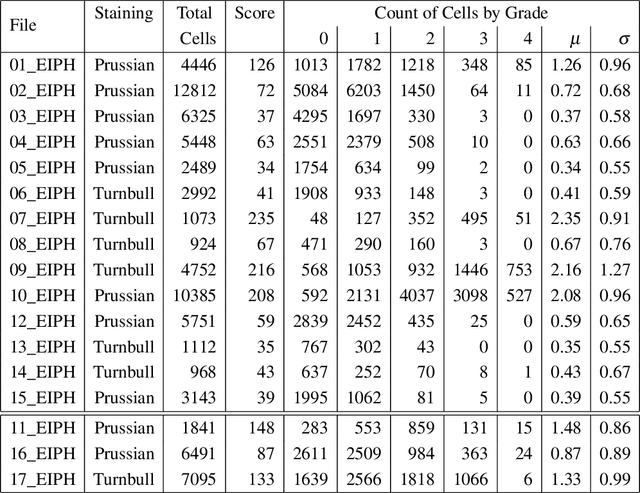
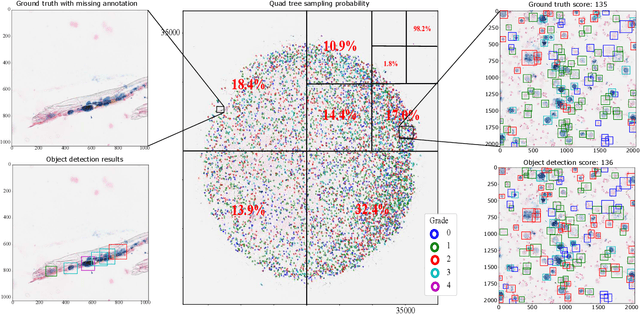
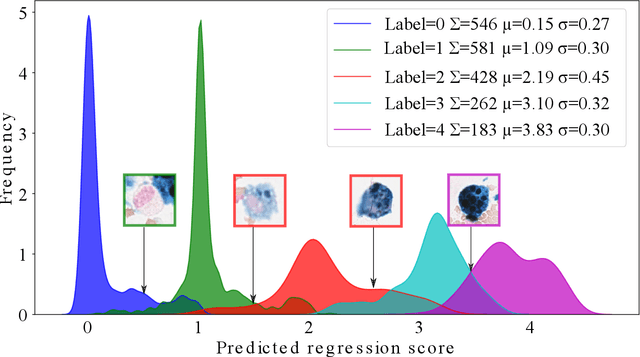
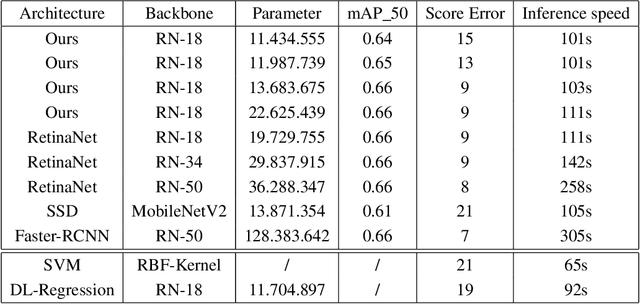
Abstract:Purpose: Exercise-induced pulmonary hemorrhage (EIPH) is a common syndrome in sport horses with negative impact on performance. Cytology of bronchoalveolar lavage fluid by use of a scoring system is considered the most sensitive diagnostic method. Macrophages are classified depending on the degree of cytoplasmic hemosiderin content. The current gold standard is manual grading, which is however monotonous and time-consuming. Methods: We evaluated state-of-the-art deep learning-based methods for single cell macrophage classification and compared them against the performance of nine cytology experts and evaluated inter- and intra-observer variability. Additionally, we evaluated object detection methods on a novel data set of 17 completely annotated cytology whole slide images (WSI) containing 78,047 hemosiderophages. Resultsf: Our deep learning-based approach reached a concordance of 0.85, partially exceeding human expert concordance (0.68 to 0.86, $\mu$=0.73, $\sigma$ =0.04). Intra-observer variability was high (0.68 to 0.88) and inter-observer concordance was moderate (Fleiss kappa = 0.67). Our object detection approach has a mean average precision of 0.66 over the five classes from the whole slide gigapixel image and a computation time of below two minutes. Conclusion: To mitigate the high inter- and intra-rater variability, we propose our automated object detection pipeline, enabling accurate, reproducible and quick EIPH scoring in WSI.
Field of Interest Prediction for Computer-Aided Mitotic Count
Feb 12, 2019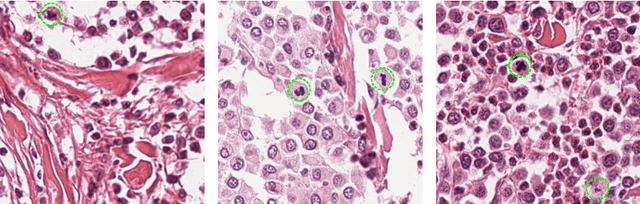
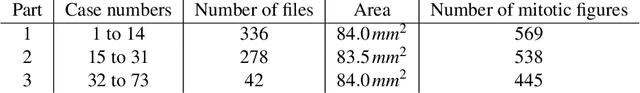
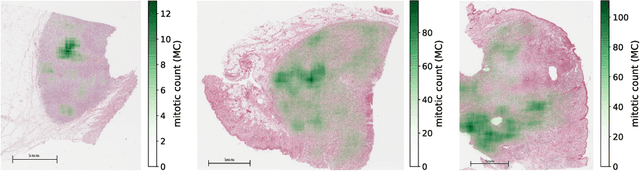
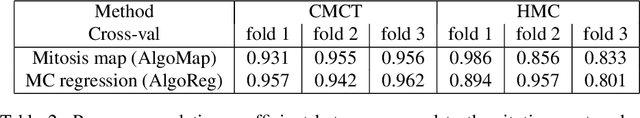
Abstract:Manual counts of mitotic figures, which are determined in the tumor region with the highest mitotic activity, are a key parameter of most tumor grading schemes. It is however strongly dependent on the area selection. To reduce potential variability of prognosis due to this, we propose to use an algorithmic field of interest prediction to assess the area of highest mitotic activity in a whole-slide image. Methods: We evaluated two state-of-the-art methods, all based on the use of deep convolutional neural networks on their ability to predict the mitotic count in digital histopathology slides. We evaluated them on a novel dataset of 32 completely annotated whole slide images from canine cutaneous mast cell tumors (CMCT) and one publicly available human mamma carcinoma (HMC) dataset. We first compared the mitotic counts (MC) predicted by the two models with the ground truth MC on both data sets. Second, for the CMCT data set, we compared the computationally predicted position and MC of the area of highest mitotic activity with size-equivalent areas selected by eight veterinary pathologists. Results: We found a high correlation between the mitotic count as predicted by the models (Pearson's correlation coefficient between 0.931 and 0.962 for the CMCT data set and between 0.801 and 0.986 for the HMC data set) on the slides. For the CMCT data set, this is also reflected in the predicted position representing mitotic counts in mostly the upper quartile of the slide's ground truth MC distribution. Further, we found strong differences between experts in position selection. Conclusion: While the mitotic counts in areas selected by the experts substantially varied, both algorithmic approaches were consistently able to generate a good estimate of the area of highest mitotic count. To achieve better inter-rater agreement, we propose to use computer-based area selection for manual mitotic count.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge