Chaitanya Kaul
IGAF: Incremental Guided Attention Fusion for Depth Super-Resolution
Jan 03, 2025Abstract:Accurate depth estimation is crucial for many fields, including robotics, navigation, and medical imaging. However, conventional depth sensors often produce low-resolution (LR) depth maps, making detailed scene perception challenging. To address this, enhancing LR depth maps to high-resolution (HR) ones has become essential, guided by HR-structured inputs like RGB or grayscale images. We propose a novel sensor fusion methodology for guided depth super-resolution (GDSR), a technique that combines LR depth maps with HR images to estimate detailed HR depth maps. Our key contribution is the Incremental guided attention fusion (IGAF) module, which effectively learns to fuse features from RGB images and LR depth maps, producing accurate HR depth maps. Using IGAF, we build a robust super-resolution model and evaluate it on multiple benchmark datasets. Our model achieves state-of-the-art results compared to all baseline models on the NYU v2 dataset for $\times 4$, $\times 8$, and $\times 16$ upsampling. It also outperforms all baselines in a zero-shot setting on the Middlebury, Lu, and RGB-D-D datasets. Code, environments, and models are available on GitHub.
HpEIS: Learning Hand Pose Embeddings for Multimedia Interactive Systems
Oct 11, 2024



Abstract:We present a novel Hand-pose Embedding Interactive System (HpEIS) as a virtual sensor, which maps users' flexible hand poses to a two-dimensional visual space using a Variational Autoencoder (VAE) trained on a variety of hand poses. HpEIS enables visually interpretable and guidable support for user explorations in multimedia collections, using only a camera as an external hand pose acquisition device. We identify general usability issues associated with system stability and smoothing requirements through pilot experiments with expert and inexperienced users. We then design stability and smoothing improvements, including hand-pose data augmentation, an anti-jitter regularisation term added to loss function, stabilising post-processing for movement turning points and smoothing post-processing based on One Euro Filters. In target selection experiments (n=12), we evaluate HpEIS by measures of task completion time and the final distance to target points, with and without the gesture guidance window condition. Experimental responses indicate that HpEIS provides users with a learnable, flexible, stable and smooth mid-air hand movement interaction experience.
AI-Enabled sensor fusion of time of flight imaging and mmwave for concealed metal detection
Aug 01, 2024



Abstract:In the field of detection and ranging, multiple complementary sensing modalities may be used to enrich the information obtained from a dynamic scene. One application of this sensor fusion is in public security and surveillance, whose efficacy and privacy protection measures must be continually evaluated. We present a novel deployment of sensor fusion for the discrete detection of concealed metal objects on persons whilst preserving their privacy. This is achieved by coupling off-the-shelf mmWave radar and depth camera technology with a novel neural network architecture that processes the radar signals using convolutional Long Short-term Memory (LSTM) blocks and the depth signal, using convolutional operations. The combined latent features are then magnified using a deep feature magnification to learn cross-modality dependencies in the data. We further propose a decoder, based on the feature extraction and embedding block, to learn an efficient upsampling of the latent space to learn the location of the concealed object in the spatial domain through radar feature guidance. We demonstrate the detection of presence and inference of 3D location of concealed metal objects with an accuracy of up to 95%, using a technique that is robust to multiple persons. This work provides a demonstration of the potential for cost effective and portable sensor fusion, with strong opportunities for further development.
Is One GPU Enough? Pushing Image Generation at Higher-Resolutions with Foundation Models
Jun 12, 2024



Abstract:In this work, we introduce Pixelsmith, a zero-shot text-to-image generative framework to sample images at higher resolutions with a single GPU. We are the first to show that it is possible to scale the output of a pre-trained diffusion model by a factor of 1000, opening the road for gigapixel image generation at no additional cost. Our cascading method uses the image generated at the lowest resolution as a baseline to sample at higher resolutions. For the guidance, we introduce the Slider, a tunable mechanism that fuses the overall structure contained in the first-generated image with enhanced fine details. At each inference step, we denoise patches rather than the entire latent space, minimizing memory demands such that a single GPU can handle the process, regardless of the image's resolution. Our experimental results show that Pixelsmith not only achieves higher quality and diversity compared to existing techniques, but also reduces sampling time and artifacts. The code for our work is available at https://github.com/Thanos-DB/Pixelsmith.
GLFNET: Global-Local (frequency) Filter Networks for efficient medical image segmentation
Mar 01, 2024



Abstract:We propose a novel transformer-style architecture called Global-Local Filter Network (GLFNet) for medical image segmentation and demonstrate its state-of-the-art performance. We replace the self-attention mechanism with a combination of global-local filter blocks to optimize model efficiency. The global filters extract features from the whole feature map whereas the local filters are being adaptively created as 4x4 patches of the same feature map and add restricted scale information. In particular, the feature extraction takes place in the frequency domain rather than the commonly used spatial (image) domain to facilitate faster computations. The fusion of information from both spatial and frequency spaces creates an efficient model with regards to complexity, required data and performance. We test GLFNet on three benchmark datasets achieving state-of-the-art performance on all of them while being almost twice as efficient in terms of GFLOP operations.
The legibility of the imaged human brain
Aug 23, 2023



Abstract:Our knowledge of the organisation of the human brain at the population-level is yet to translate into power to predict functional differences at the individual-level, limiting clinical applications, and casting doubt on the generalisability of inferred mechanisms. It remains unknown whether the difficulty arises from the absence of individuating biological patterns within the brain, or from limited power to access them with the models and compute at our disposal. Here we comprehensively investigate the resolvability of such patterns with data and compute at unprecedented scale. Across 23810 unique participants from UK Biobank, we systematically evaluate the predictability of 25 individual biological characteristics, from all available combinations of structural and functional neuroimaging data. Over 4526 GPU*hours of computation, we train, optimize, and evaluate out-of-sample 700 individual predictive models, including multilayer perceptrons of demographic, psychological, serological, chronic morbidity, and functional connectivity characteristics, and both uni- and multi-modal 3D convolutional neural network models of macro- and micro-structural brain imaging. We find a marked discrepancy between the high predictability of sex (balanced accuracy 99.7%), age (mean absolute error 2.048 years, R2 0.859), and weight (mean absolute error 2.609Kg, R2 0.625), for which we set new state-of-the-art performance, and the surprisingly low predictability of other characteristics. Neither structural nor functional imaging predicted individual psychology better than the coincidence of common chronic morbidity (p<0.05). Serology predicted common morbidity (p<0.05) and was best predicted by it (p<0.001), followed by structural neuroimaging (p<0.05). Our findings suggest either more informative imaging or more powerful models will be needed to decipher individual level characteristics from the brain.
mmSense: Detecting Concealed Weapons with a Miniature Radar Sensor
Feb 28, 2023



Abstract:For widespread adoption, public security and surveillance systems must be accurate, portable, compact, and real-time, without impeding the privacy of the individuals being observed. Current systems broadly fall into two categories -- image-based which are accurate, but lack privacy, and RF signal-based, which preserve privacy but lack portability, compactness and accuracy. Our paper proposes mmSense, an end-to-end portable miniaturised real-time system that can accurately detect the presence of concealed metallic objects on persons in a discrete, privacy-preserving modality. mmSense features millimeter wave radar technology, provided by Google's Soli sensor for its data acquisition, and TransDope, our real-time neural network, capable of processing a single radar data frame in 19 ms. mmSense achieves high recognition rates on a diverse set of challenging scenes while running on standard laptop hardware, demonstrating a significant advancement towards creating portable, cost-effective real-time radar based surveillance systems.
Optimizing Vision Transformers for Medical Image Segmentation and Few-Shot Domain Adaptation
Oct 14, 2022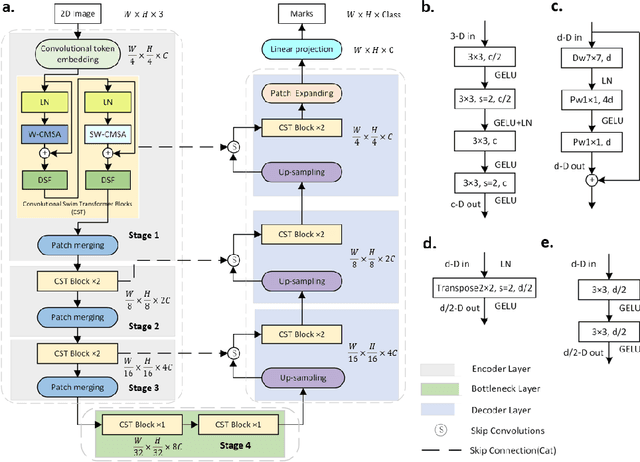
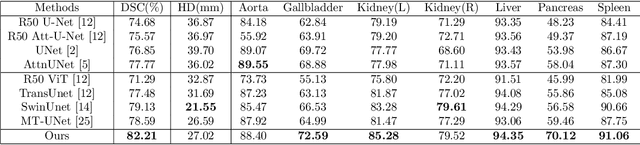
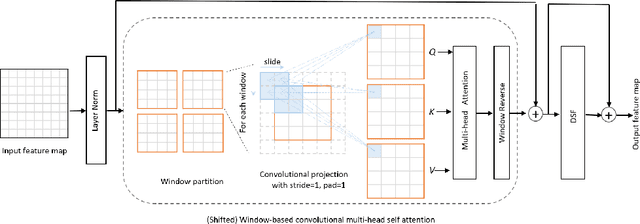
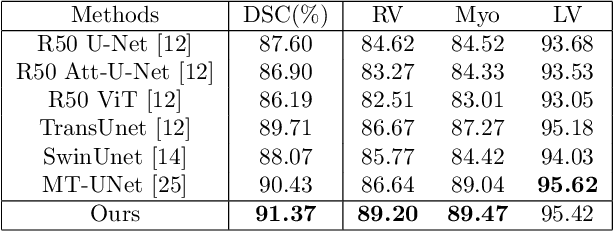
Abstract:The adaptation of transformers to computer vision is not straightforward because the modelling of image contextual information results in quadratic computational complexity with relation to the input features. Most of existing methods require extensive pre-training on massive datasets such as ImageNet and therefore their application to fields such as healthcare is less effective. CNNs are the dominant architecture in computer vision tasks because convolutional filters can effectively model local dependencies and reduce drastically the parameters required. However, convolutional filters cannot handle more complex interactions, which are beyond a small neighbour of pixels. Furthermore, their weights are fixed after training and thus they do not take into consideration changes in the visual input. Inspired by recent work on hybrid visual transformers with convolutions and hierarchical transformers, we propose Convolutional Swin-Unet (CS-Unet) transformer blocks and optimise their settings with relation to patch embedding, projection, the feed-forward network, up sampling and skip connections. CS-Unet can be trained from scratch and inherits the superiority of convolutions in each feature process phase. It helps to encode precise spatial information and produce hierarchical representations that contribute to object concepts at various scales. Experiments show that CS-Unet without pre-training surpasses other state-of-the-art counterparts by large margins on two medical CT and MRI datasets with fewer parameters. In addition, two domain-adaptation experiments on optic disc and polyp image segmentation further prove that our method is highly generalizable and effectively bridges the domain gap between images from different sources.
The Fully Convolutional Transformer for Medical Image Segmentation
Jun 01, 2022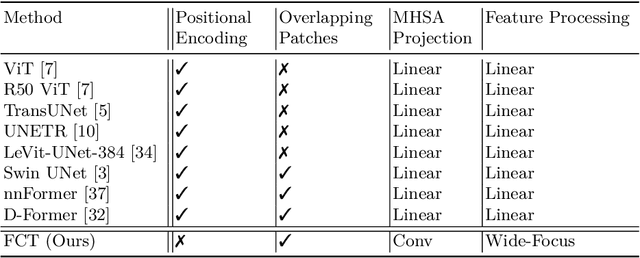
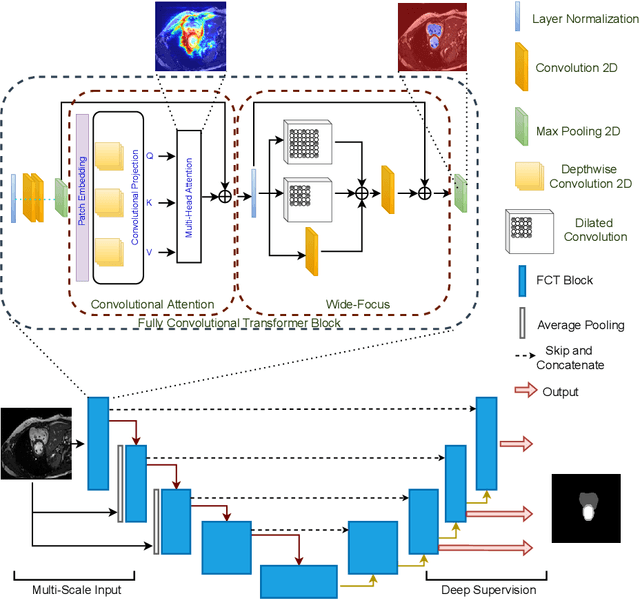
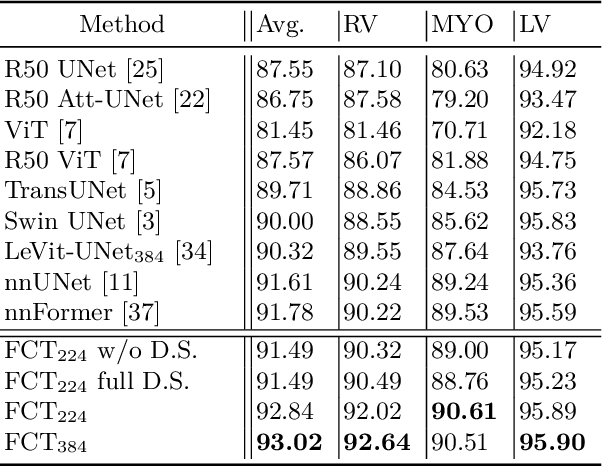
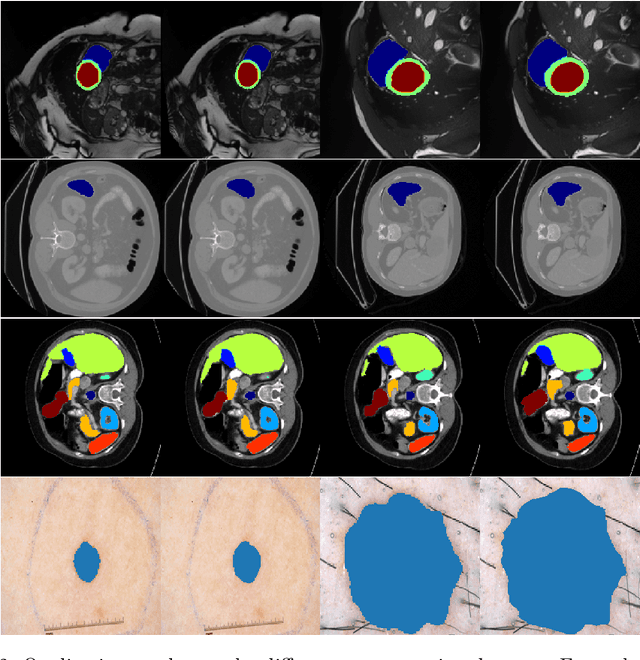
Abstract:We propose a novel transformer model, capable of segmenting medical images of varying modalities. Challenges posed by the fine grained nature of medical image analysis mean that the adaptation of the transformer for their analysis is still at nascent stages. The overwhelming success of the UNet lay in its ability to appreciate the fine-grained nature of the segmentation task, an ability which existing transformer based models do not currently posses. To address this shortcoming, we propose The Fully Convolutional Transformer (FCT), which builds on the proven ability of Convolutional Neural Networks to learn effective image representations, and combines them with the ability of Transformers to effectively capture long-term dependencies in its inputs. The FCT is the first fully convolutional Transformer model in medical imaging literature. It processes its input in two stages, where first, it learns to extract long range semantic dependencies from the input image, and then learns to capture hierarchical global attributes from the features. FCT is compact, accurate and robust. Our results show that it outperforms all existing transformer architectures by large margins across multiple medical image segmentation datasets of varying data modalities without the need for any pre-training. FCT outperforms its immediate competitor on the ACDC dataset by 1.3%, on the Synapse dataset by 4.4%, on the Spleen dataset by 1.2% and on ISIC 2017 dataset by 1.1% on the dice metric, with up to five times fewer parameters. Our code, environments and models will be available via GitHub.
Rotation Equivariant 3D Hand Mesh Generation from a Single RGB Image
Nov 25, 2021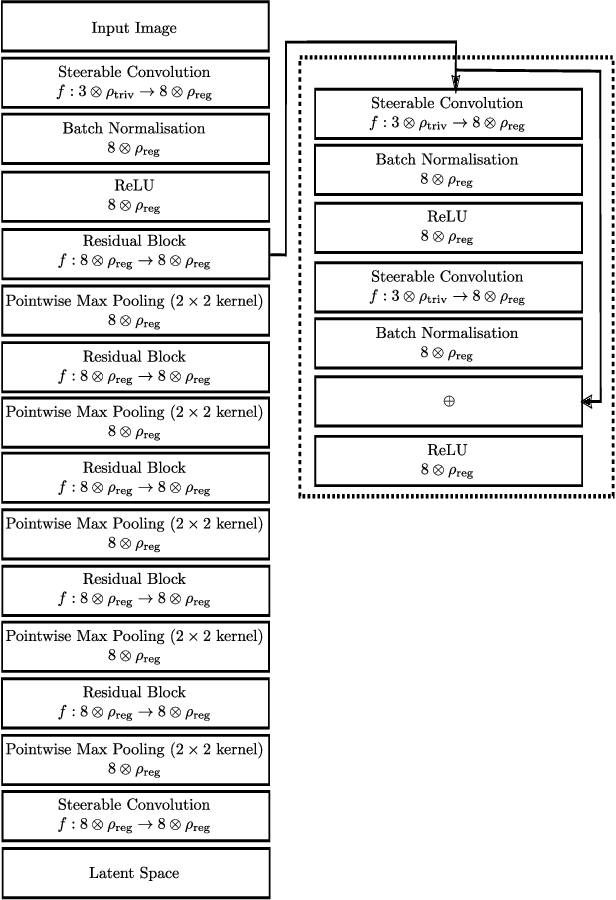
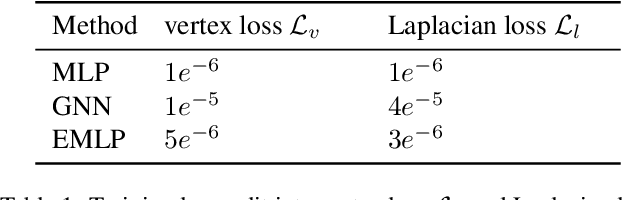
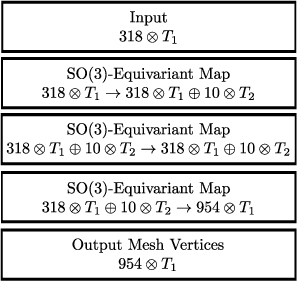
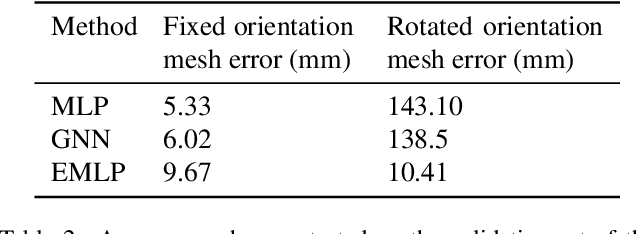
Abstract:We develop a rotation equivariant model for generating 3D hand meshes from 2D RGB images. This guarantees that as the input image of a hand is rotated the generated mesh undergoes a corresponding rotation. Furthermore, this removes undesirable deformations in the meshes often generated by methods without rotation equivariance. By building a rotation equivariant model, through considering symmetries in the problem, we reduce the need for training on very large datasets to achieve good mesh reconstruction. The encoder takes images defined on $\mathbb{Z}^{2}$ and maps these to latent functions defined on the group $C_{8}$. We introduce a novel vector mapping function to map the function defined on $C_{8}$ to a latent point cloud space defined on the group $\mathrm{SO}(2)$. Further, we introduce a 3D projection function that learns a 3D function from the $\mathrm{SO}(2)$ latent space. Finally, we use an $\mathrm{SO}(3)$ equivariant decoder to ensure rotation equivariance. Our rotation equivariant model outperforms state-of-the-art methods on a real-world dataset and we demonstrate that it accurately captures the shape and pose in the generated meshes under rotation of the input hand.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge