Carsten Eickhoff
Stable Anisotropic Regularization
May 30, 2023Abstract:Given the success of Large Language Models (LLMs), there has been considerable interest in studying the properties of model activations. The literature overwhelmingly agrees that LLM representations are dominated by a few ``outlier dimensions'' with exceedingly high variance and magnitude. Several studies in Natural Language Processing (NLP) have sought to mitigate the impact of such outlier dimensions and force LLMs to be isotropic (i.e., have uniform variance across all dimensions in embedding space). Isotropy is thought to be a desirable property for LLMs that improves model performance and more closely aligns textual representations with human intuition. However, many of the claims regarding isotropy in NLP have been based on the average cosine similarity of embeddings, which has recently been shown to be a flawed measure of isotropy. In this paper, we propose I-STAR: IsoScore$^{\star}$-based STable Anisotropic Regularization, a novel regularization method that can be used to increase or decrease levels of isotropy in embedding space during training. I-STAR uses IsoScore$^{\star}$, the first accurate measure of isotropy that is both differentiable and stable on mini-batch computations. In contrast to several previous works, we find that \textit{decreasing} isotropy in contextualized embeddings improves performance on the majority of tasks and models considered in this paper.
Language Models Implement Simple Word2Vec-style Vector Arithmetic
May 25, 2023



Abstract:A primary criticism towards language models (LMs) is their inscrutability. This paper presents evidence that, despite their size and complexity, LMs sometimes exploit a computational mechanism familiar from traditional word embeddings: the use of simple vector arithmetic in order to encode abstract relations (e.g., Poland:Warsaw::China:Beijing). We investigate a range of language model sizes (from 124M parameters to 176B parameters) in an in-context learning setting, and find that for a variety of tasks (involving capital cities, upper-casing, and past-tensing), a key part of the mechanism reduces to a simple linear update applied by the feedforward networks. We further show that this mechanism is specific to tasks that require retrieval from pretraining memory, rather than retrieval from local context. Our results contribute to a growing body of work on the mechanistic interpretability of LLMs, and offer reason to be optimistic that, despite the massive and non-linear nature of the models, the strategies they ultimately use to solve tasks can sometimes reduce to familiar and even intuitive algorithms.
Enhancing the Ranking Context of Dense Retrieval Methods through Reciprocal Nearest Neighbors
May 25, 2023Abstract:Sparse annotation poses persistent challenges to training dense retrieval models, such as the problem of false negatives, i.e. unlabeled relevant documents that are spuriously used as negatives in contrastive learning, distorting the training signal. To alleviate this problem, we introduce evidence-based label smoothing, a computationally efficient method that prevents penalizing the model for assigning high relevance to false negatives. To compute the target relevance distribution over candidate documents within the ranking context of a given query, candidates most similar to the ground truth are assigned a non-zero relevance probability based on the degree of their similarity to the ground-truth document(s). As a relevance estimate we leverage an improved similarity metric based on reciprocal nearest neighbors, which can also be used independently to rerank candidates in post-processing. Through extensive experiments on two large-scale ad hoc text retrieval datasets we demonstrate that both methods can improve the ranking effectiveness of dense retrieval models.
Neural Summarization of Electronic Health Records
May 24, 2023Abstract:Hospital discharge documentation is among the most essential, yet time-consuming documents written by medical practitioners. The objective of this study was to automatically generate hospital discharge summaries using neural network summarization models. We studied various data preparation and neural network training techniques that generate discharge summaries. Using nursing notes and discharge summaries from the MIMIC-III dataset, we studied the viability of the automatic generation of various sections of a discharge summary using four state-of-the-art neural network summarization models (BART, T5, Longformer and FLAN-T5). Our experiments indicated that training environments including nursing notes as the source, and discrete sections of the discharge summary as the target output (e.g. "History of Present Illness") improve language model efficiency and text quality. According to our findings, the fine-tuned BART model improved its ROUGE F1 score by 43.6% against its standard off-the-shelf version. We also found that fine-tuning the baseline BART model with other setups caused different degrees of improvement (up to 80% relative improvement). We also observed that a fine-tuned T5 generally achieves higher ROUGE F1 scores than other fine-tuned models and a fine-tuned FLAN-T5 achieves the highest ROUGE score overall, i.e., 45.6. For majority of the fine-tuned language models, summarizing discharge summary report sections separately outperformed the summarization the entire report quantitatively. On the other hand, fine-tuning language models that were previously instruction fine-tuned showed better performance in summarizing entire reports. This study concludes that a focused dataset designed for the automatic generation of discharge summaries by a language model can produce coherent Discharge Summary sections.
CroCoSum: A Benchmark Dataset for Cross-Lingual Code-Switched Summarization
Mar 07, 2023Abstract:Cross-lingual summarization (CLS) has attracted increasing interest in recent years due to the availability of large-scale web-mined datasets and the advancements of multilingual language models. However, given the rareness of naturally occurring CLS resources, the majority of datasets are forced to rely on translation which can contain overly literal artifacts. This restricts our ability to observe naturally occurring CLS pairs that capture organic diction, including instances of code-switching. This alteration between languages in mid-message is a common phenomenon in multilingual settings yet has been largely overlooked in cross-lingual contexts due to data scarcity. To address this gap, we introduce CroCoSum, a dataset of cross-lingual code-switched summarization of technology news. It consists of over 24,000 English source articles and 18,000 human-curated Chinese news summaries, with more than 92% of the summaries containing code-switched phrases. For reference, we evaluate the performance of existing approaches including pipeline, end-to-end, and zero-shot methods. We show that leveraging existing resources as a pretraining step does not improve performance on CroCoSum, indicating the limited generalizability of existing resources. Finally, we discuss the challenges of evaluating cross-lingual summarizers on code-switched generation through qualitative error analyses. Our collection and code can be accessed at https://github.com/RosenZhang/CroCoSum.
Parameter-efficient Modularised Bias Mitigation via AdapterFusion
Feb 13, 2023Abstract:Large pre-trained language models contain societal biases and carry along these biases to downstream tasks. Current in-processing bias mitigation approaches (like adversarial training) impose debiasing by updating a model's parameters, effectively transferring the model to a new, irreversible debiased state. In this work, we propose a novel approach to develop stand-alone debiasing functionalities separate from the model, which can be integrated into the model on-demand, while keeping the core model untouched. Drawing from the concept of AdapterFusion in multi-task learning, we introduce DAM (Debiasing with Adapter Modules) - a debiasing approach to first encapsulate arbitrary bias mitigation functionalities into separate adapters, and then add them to the model on-demand in order to deliver fairness qualities. We conduct a large set of experiments on three classification tasks with gender, race, and age as protected attributes. Our results show that DAM improves or maintains the effectiveness of bias mitigation, avoids catastrophic forgetting in a multi-attribute scenario, and maintains on-par task performance, while granting parameter-efficiency and easy switching between the original and debiased models.
Unsupervised Multivariate Time-Series Transformers for Seizure Identification on EEG
Jan 03, 2023



Abstract:Epilepsy is one of the most common neurological disorders, typically observed via seizure episodes. Epileptic seizures are commonly monitored through electroencephalogram (EEG) recordings due to their routine and low expense collection. The stochastic nature of EEG makes seizure identification via manual inspections performed by highly-trained experts a tedious endeavor, motivating the use of automated identification. The literature on automated identification focuses mostly on supervised learning methods requiring expert labels of EEG segments that contain seizures, which are difficult to obtain. Motivated by these observations, we pose seizure identification as an unsupervised anomaly detection problem. To this end, we employ the first unsupervised transformer-based model for seizure identification on raw EEG. We train an autoencoder involving a transformer encoder via an unsupervised loss function, incorporating a novel masking strategy uniquely designed for multivariate time-series data such as EEG. Training employs EEG recordings that do not contain any seizures, while seizures are identified with respect to reconstruction errors at inference time. We evaluate our method on three publicly available benchmark EEG datasets for distinguishing seizure vs. non-seizure windows. Our method leads to significantly better seizure identification performance than supervised learning counterparts, by up to 16% recall, 9% accuracy, and 9% Area under the Receiver Operating Characteristics Curve (AUC), establishing particular benefits on highly imbalanced data. Through accurate seizure identification, our method could facilitate widely accessible and early detection of epilepsy development, without needing expensive label collection or manual feature extraction.
Evaluating Search Explainability with Psychometrics and Crowdsourcing
Oct 17, 2022
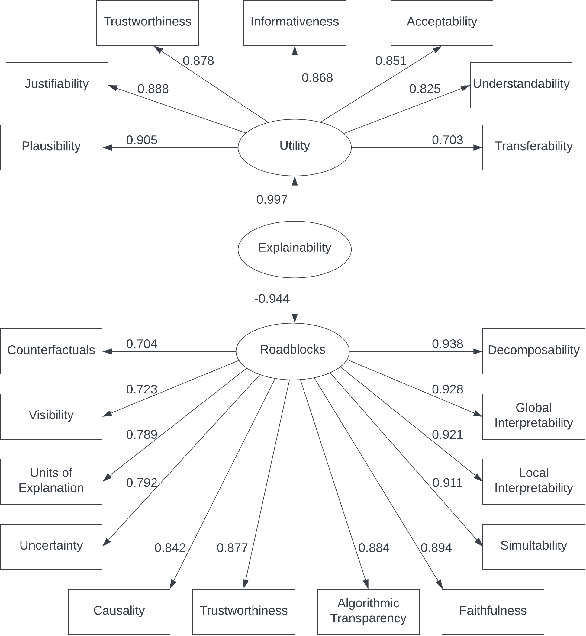
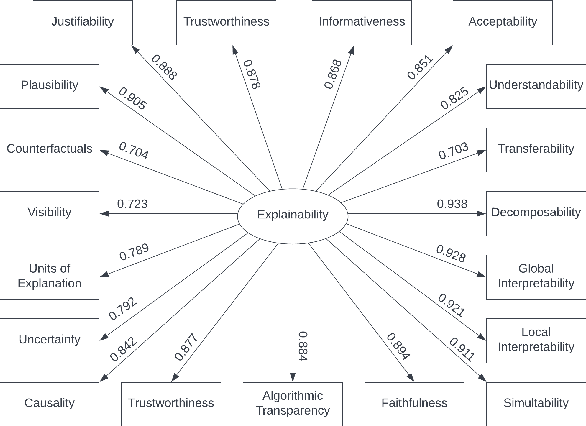
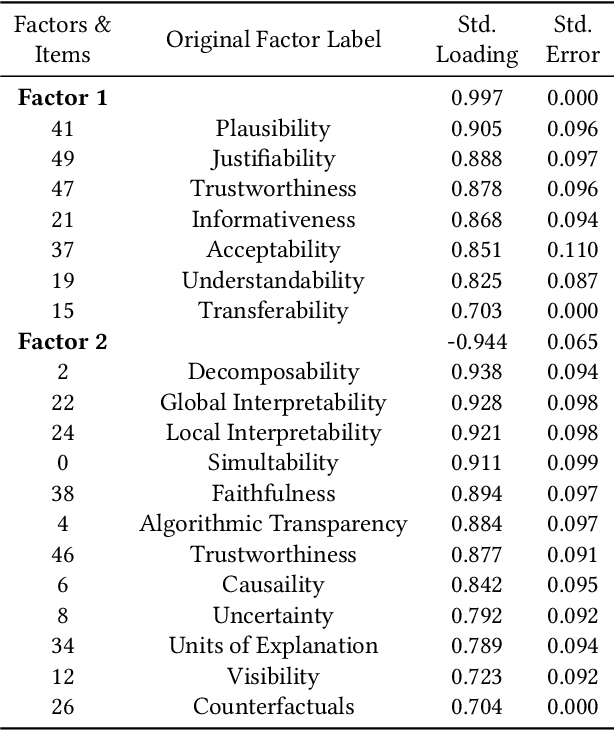
Abstract:Information retrieval (IR) systems have become an integral part of our everyday lives. As search engines, recommender systems, and conversational agents are employed across various domains from recreational search to clinical decision support, there is an increasing need for transparent and explainable systems to guarantee accountable, fair, and unbiased results. Despite many recent advances towards explainable AI and IR techniques, there is no consensus on what it means for a system to be explainable. Although a growing body of literature suggests that explainability is comprised of multiple subfactors, virtually all existing approaches treat it as a singular notion. In this paper, we examine explainability in Web search systems, leveraging psychometrics and crowdsourcing to identify human-centered factors of explainability. Based on these factors, we establish a continuous-scale evaluation instrument for explainable search systems that allows researchers and practitioners to trade-off performance in a more flexible manner than what was previously possible.
Linearly Mapping from Image to Text Space
Sep 30, 2022
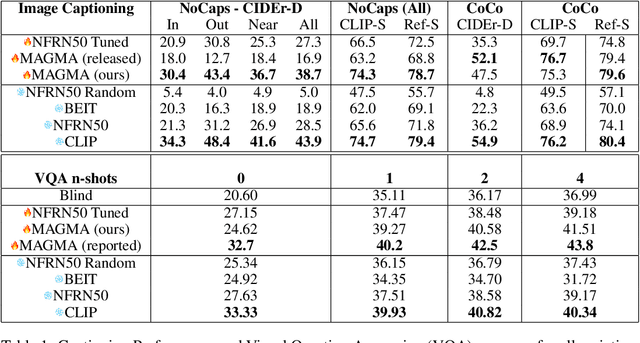

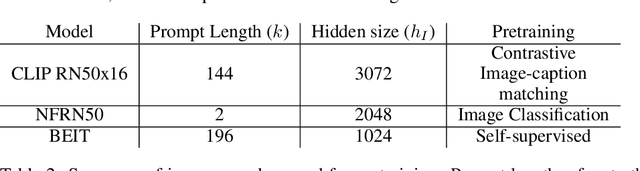
Abstract:The extent to which text-only language models (LMs) learn to represent the physical, non-linguistic world is an open question. Prior work has shown that pretrained LMs can be taught to ``understand'' visual inputs when the models' parameters are updated on image captioning tasks. We test a stronger hypothesis: that the conceptual representations learned by text-only models are functionally equivalent (up to a linear transformation) to those learned by models trained on vision tasks. Specifically, we show that the image representations from vision models can be transferred as continuous prompts to frozen LMs by training only a single linear projection. Using these to prompt the LM achieves competitive performance on captioning and visual question answering tasks compared to models that tune both the image encoder and text decoder (such as the MAGMA model). We compare three image encoders with increasing amounts of linguistic supervision seen during pretraining: BEIT (no linguistic information), NF-ResNET (lexical category information), and CLIP (full natural language descriptions). We find that all three encoders perform equally well at transferring visual property information to the language model (e.g., whether an animal is large or small), but that image encoders pretrained with linguistic supervision more saliently encode category information (e.g., distinguishing hippo vs.\ elephant) and thus perform significantly better on benchmark language-and-vision tasks. Our results indicate that LMs encode conceptual information structurally similarly to vision-based models, even those that are solely trained on images.
Pretraining on Interactions for Learning Grounded Affordance Representations
Jul 05, 2022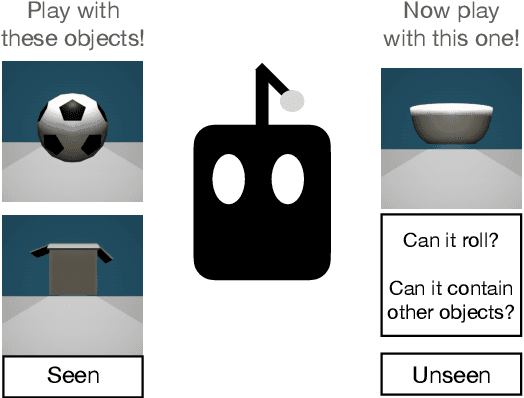
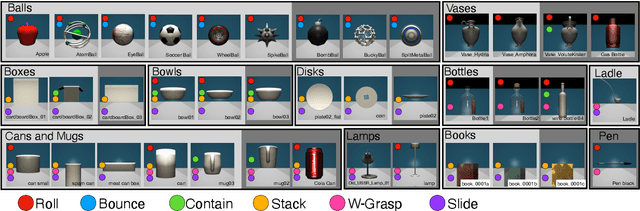
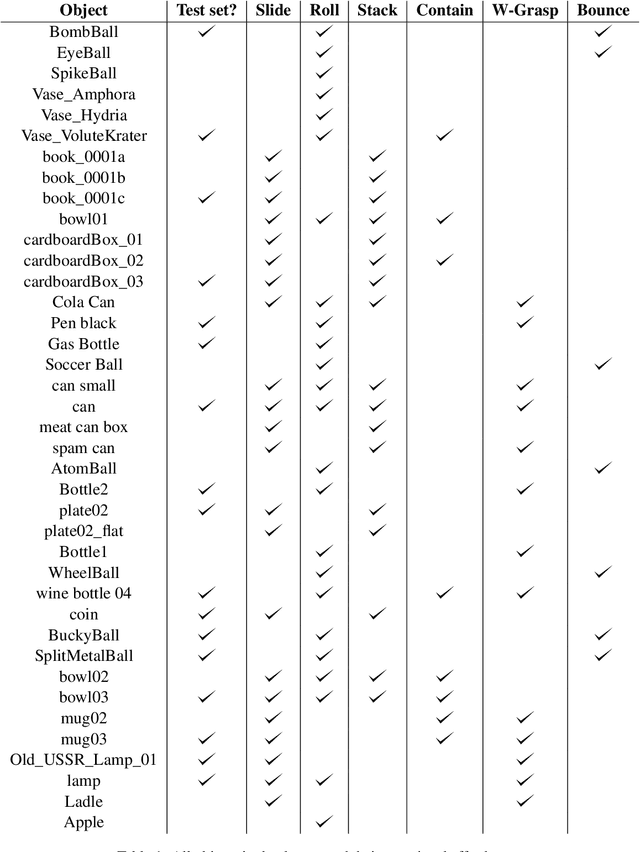

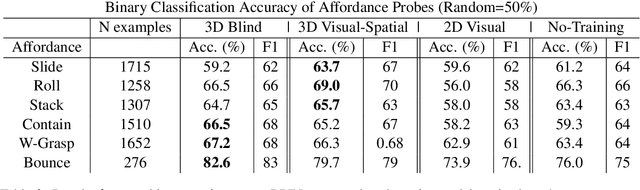
Abstract:Lexical semantics and cognitive science point to affordances (i.e. the actions that objects support) as critical for understanding and representing nouns and verbs. However, study of these semantic features has not yet been integrated with the "foundation" models that currently dominate language representation research. We hypothesize that predictive modeling of object state over time will result in representations that encode object affordance information "for free". We train a neural network to predict objects' trajectories in a simulated interaction and show that our network's latent representations differentiate between both observed and unobserved affordances. We find that models trained using 3D simulations from our SPATIAL dataset outperform conventional 2D computer vision models trained on a similar task, and, on initial inspection, that differences between concepts correspond to expected features (e.g., roll entails rotation). Our results suggest a way in which modern deep learning approaches to grounded language learning can be integrated with traditional formal semantic notions of lexical representations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge