Babak Taati
STAF: Sinusoidal Trainable Activation Functions for Implicit Neural Representation
Feb 02, 2025Abstract:Implicit Neural Representations (INRs) have emerged as a powerful framework for modeling continuous signals. The spectral bias of ReLU-based networks is a well-established limitation, restricting their ability to capture fine-grained details in target signals. While previous works have attempted to mitigate this issue through frequency-based encodings or architectural modifications, these approaches often introduce additional complexity and do not fully address the underlying challenge of learning high-frequency components efficiently. We introduce Sinusoidal Trainable Activation Functions (STAF), designed to directly tackle this limitation by enabling networks to adaptively learn and represent complex signals with higher precision and efficiency. STAF inherently modulates its frequency components, allowing for self-adaptive spectral learning. This capability significantly improves convergence speed and expressivity, making STAF highly effective for both signal representations and inverse problems. Through extensive evaluations, we demonstrate that STAF outperforms state-of-the-art (SOTA) methods in accuracy and reconstruction fidelity with superior Peak Signal-to-Noise Ratio (PSNR). These results establish STAF as a robust solution for overcoming spectral bias and the capacity-convergence gap, making it valuable for computer graphics and related fields. Our codebase is publicly accessible on the https://github.com/AlirezaMorsali/STAF.
STARS: Self-supervised Tuning for 3D Action Recognition in Skeleton Sequences
Jul 15, 2024



Abstract:Self-supervised pretraining methods with masked prediction demonstrate remarkable within-dataset performance in skeleton-based action recognition. However, we show that, unlike contrastive learning approaches, they do not produce well-separated clusters. Additionally, these methods struggle with generalization in few-shot settings. To address these issues, we propose Self-supervised Tuning for 3D Action Recognition in Skeleton sequences (STARS). Specifically, STARS first uses a masked prediction stage using an encoder-decoder architecture. It then employs nearest-neighbor contrastive learning to partially tune the weights of the encoder, enhancing the formation of semantic clusters for different actions. By tuning the encoder for a few epochs, and without using hand-crafted data augmentations, STARS achieves state-of-the-art self-supervised results in various benchmarks, including NTU-60, NTU-120, and PKU-MMD. In addition, STARS exhibits significantly better results than masked prediction models in few-shot settings, where the model has not seen the actions throughout pretraining. Project page: https://soroushmehraban.github.io/stars/
SUM: Saliency Unification through Mamba for Visual Attention Modeling
Jun 25, 2024



Abstract:Visual attention modeling, important for interpreting and prioritizing visual stimuli, plays a significant role in applications such as marketing, multimedia, and robotics. Traditional saliency prediction models, especially those based on Convolutional Neural Networks (CNNs) or Transformers, achieve notable success by leveraging large-scale annotated datasets. However, the current state-of-the-art (SOTA) models that use Transformers are computationally expensive. Additionally, separate models are often required for each image type, lacking a unified approach. In this paper, we propose Saliency Unification through Mamba (SUM), a novel approach that integrates the efficient long-range dependency modeling of Mamba with U-Net to provide a unified model for diverse image types. Using a novel Conditional Visual State Space (C-VSS) block, SUM dynamically adapts to various image types, including natural scenes, web pages, and commercial imagery, ensuring universal applicability across different data types. Our comprehensive evaluations across five benchmarks demonstrate that SUM seamlessly adapts to different visual characteristics and consistently outperforms existing models. These results position SUM as a versatile and powerful tool for advancing visual attention modeling, offering a robust solution universally applicable across different types of visual content.
Benchmarking Skeleton-based Motion Encoder Models for Clinical Applications: Estimating Parkinson's Disease Severity in Walking Sequences
May 28, 2024



Abstract:This study investigates the application of general human motion encoders trained on large-scale human motion datasets for analyzing gait patterns in PD patients. Although these models have learned a wealth of human biomechanical knowledge, their effectiveness in analyzing pathological movements, such as parkinsonian gait, has yet to be fully validated. We propose a comparative framework and evaluate six pre-trained state-of-the-art human motion encoder models on their ability to predict the Movement Disorder Society - Unified Parkinson's Disease Rating Scale (MDS-UPDRS-III) gait scores from motion capture data. We compare these against a traditional gait feature-based predictive model in a recently released large public PD dataset, including PD patients on and off medication. The feature-based model currently shows higher weighted average accuracy, precision, recall, and F1-score. Motion encoder models with closely comparable results demonstrate promise for scalability and efficiency in clinical settings. This potential is underscored by the enhanced performance of the encoder model upon fine-tuning on PD training set. Four of the six human motion models examined provided prediction scores that were significantly different between on- and off-medication states. This finding reveals the sensitivity of motion encoder models to nuanced clinical changes. It also underscores the necessity for continued customization of these models to better capture disease-specific features, thereby reducing the reliance on labor-intensive feature engineering. Lastly, we establish a benchmark for the analysis of skeleton-based motion encoder models in clinical settings. To the best of our knowledge, this is the first study to provide a benchmark that enables state-of-the-art models to be tested and compete in a clinical context. Codes and benchmark leaderboard are available at code.
MotionAGFormer: Enhancing 3D Human Pose Estimation with a Transformer-GCNFormer Network
Oct 25, 2023



Abstract:Recent transformer-based approaches have demonstrated excellent performance in 3D human pose estimation. However, they have a holistic view and by encoding global relationships between all the joints, they do not capture the local dependencies precisely. In this paper, we present a novel Attention-GCNFormer (AGFormer) block that divides the number of channels by using two parallel transformer and GCNFormer streams. Our proposed GCNFormer module exploits the local relationship between adjacent joints, outputting a new representation that is complementary to the transformer output. By fusing these two representation in an adaptive way, AGFormer exhibits the ability to better learn the underlying 3D structure. By stacking multiple AGFormer blocks, we propose MotionAGFormer in four different variants, which can be chosen based on the speed-accuracy trade-off. We evaluate our model on two popular benchmark datasets: Human3.6M and MPI-INF-3DHP. MotionAGFormer-B achieves state-of-the-art results, with P1 errors of 38.4mm and 16.2mm, respectively. Remarkably, it uses a quarter of the parameters and is three times more computationally efficient than the previous leading model on Human3.6M dataset. Code and models are available at https://github.com/TaatiTeam/MotionAGFormer.
Pose2Gait: Extracting Gait Features from Monocular Video of Individuals with Dementia
Aug 22, 2023



Abstract:Video-based ambient monitoring of gait for older adults with dementia has the potential to detect negative changes in health and allow clinicians and caregivers to intervene early to prevent falls or hospitalizations. Computer vision-based pose tracking models can process video data automatically and extract joint locations; however, publicly available models are not optimized for gait analysis on older adults or clinical populations. In this work we train a deep neural network to map from a two dimensional pose sequence, extracted from a video of an individual walking down a hallway toward a wall-mounted camera, to a set of three-dimensional spatiotemporal gait features averaged over the walking sequence. The data of individuals with dementia used in this work was captured at two sites using a wall-mounted system to collect the video and depth information used to train and evaluate our model. Our Pose2Gait model is able to extract velocity and step length values from the video that are correlated with the features from the depth camera, with Spearman's correlation coefficients of .83 and .60 respectively, showing that three dimensional spatiotemporal features can be predicted from monocular video. Future work remains to improve the accuracy of other features, such as step time and step width, and test the utility of the predicted values for detecting meaningful changes in gait during longitudinal ambient monitoring.
Large Language Models are Fixated by Red Herrings: Exploring Creative Problem Solving and Einstellung Effect using the Only Connect Wall Dataset
Jun 19, 2023



Abstract:The quest for human imitative AI has been an enduring topic in AI research since its inception. The technical evolution and emerging capabilities of the latest cohort of large language models (LLMs) have reinvigorated the subject beyond academia to the cultural zeitgeist. While recent NLP evaluation benchmark tasks test some aspects of human-imitative behaviour (e.g., BIG-bench's 'human-like behavior' tasks), few, if not none, examine creative problem solving abilities. Creative problem solving in humans is a well-studied topic in cognitive neuroscience with standardized tests that predominantly use the ability to associate (heterogeneous) connections among clue words as a metric for creativity. Exposure to misleading stimuli - distractors dubbed red herrings - impede human performance in such tasks via the fixation effect and Einstellung paradigm. In cognitive neuroscience studies, such fixations are experimentally induced by pre-exposing participants to orthographically similar incorrect words to subsequent word-fragments or clues. The popular British quiz show Only Connect's Connecting Wall segment essentially mimics Mednick's Remote Associates Test (RAT) formulation with built-in, deliberate red herrings, which makes it an ideal proxy dataset to explore and study fixation effect and Einstellung paradigm from cognitive neuroscience in LLMs. In addition to presenting the novel Only Connect Wall (OCW) dataset, we also report results from our evaluation of selected pre-trained language models and LLMs (including OpenAI's GPT series) on creative problem solving tasks like grouping clue words by heterogeneous connections, and identifying correct open knowledge domain connections in respective groups. The code and link to the dataset are available at https://github.com/TaatiTeam/OCW.
Concurrent Validity of Automatic Speech and Pause Measures During Passage Reading in ALS
Aug 22, 2022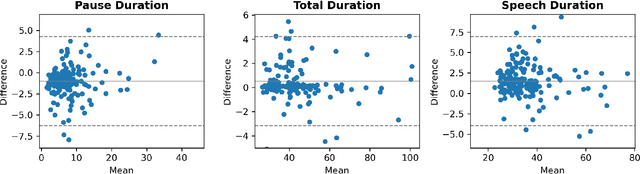
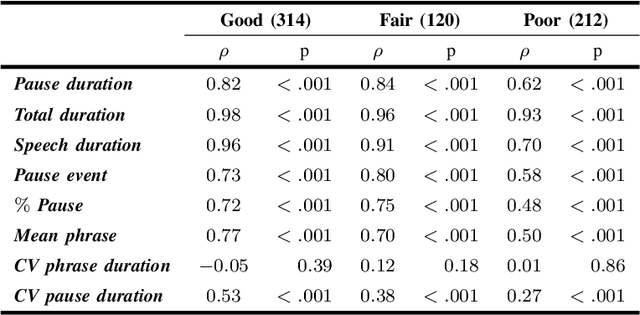
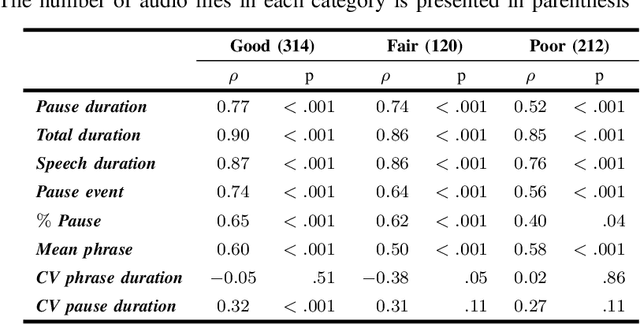
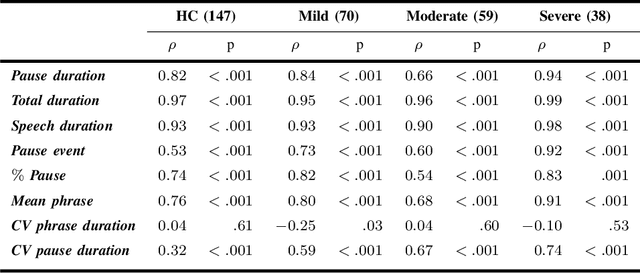
Abstract:The analysis of speech measures in individuals with amyotrophic lateral sclerosis (ALS) can provide essential information for early diagnosis and tracking disease progression. However, current methods for extracting speech and pause features are manual or semi-automatic, which makes them time-consuming and labour-intensive. The advent of speech-text alignment algorithms provides an opportunity for inexpensive, automated, and accurate analysis of speech measures in individuals with ALS. There is a need to validate speech and pause features calculated by these algorithms against current gold standard methods. In this study, we extracted 8 speech/pause features from 646 audio files of individuals with ALS and healthy controls performing passage reading. Two pretrained forced alignment models - one using transformers and another using a Gaussian mixture / hidden Markov architecture - were used for automatic feature extraction. The results were then validated against semi-automatic speech/pause analysis software, with further subgroup analyses based on audio quality and disease severity. Features extracted using transformer-based forced alignment had the highest agreement with gold standards, including in terms of audio quality and disease severity. This study lays the groundwork for future intelligent diagnostic support systems for clinicians, and for novel methods of tracking disease progression remotely from home.
Automated Temporal Segmentation of Orofacial Assessment Videos
Aug 22, 2022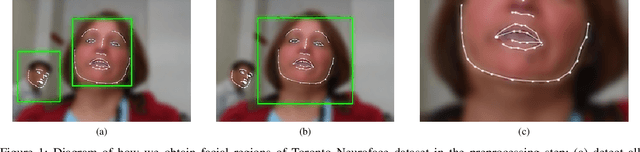
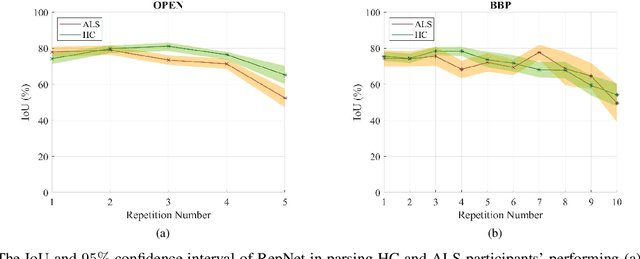
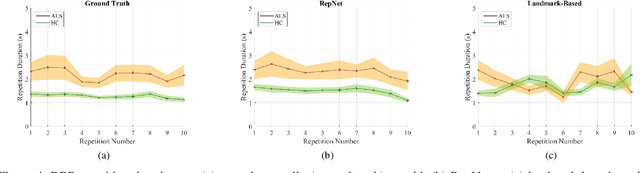
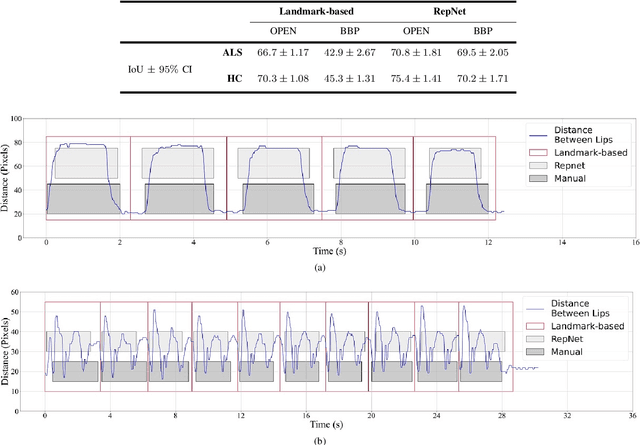
Abstract:Computer vision techniques can help automate or partially automate clinical examination of orofacial impairments to provide accurate and objective assessments. Towards the development of such automated systems, we evaluated two approaches to detect and temporally segment (parse) repetitions in orofacial assessment videos. Recorded videos of participants with amyotrophic lateral sclerosis (ALS) and healthy control (HC) individuals were obtained from the Toronto NeuroFace Dataset. Two approaches for repetition detection and parsing were examined: one based on engineered features from tracked facial landmarks and peak detection in the distance between the vermilion-cutaneous junction of the upper and lower lips (baseline analysis), and another using a pre-trained transformer-based deep learning model called RepNet (Dwibedi et al, 2020), which automatically detects periodicity, and parses periodic and semi-periodic repetitions in video data. In experimental evaluation of two orofacial assessments tasks, - repeating maximum mouth opening (OPEN) and repeating the sentence "Buy Bobby a Puppy" (BBP) - RepNet provided better parsing than the landmark-based approach, quantified by higher mean intersection-over-union (IoU) with respect to ground truth manual parsing. Automated parsing using RepNet also clearly separated HC and ALS participants based on the duration of BBP repetitions, whereas the landmark-based method could not.
Estimating Parkinsonism Severity in Natural Gait Videos of Older Adults with Dementia
May 07, 2021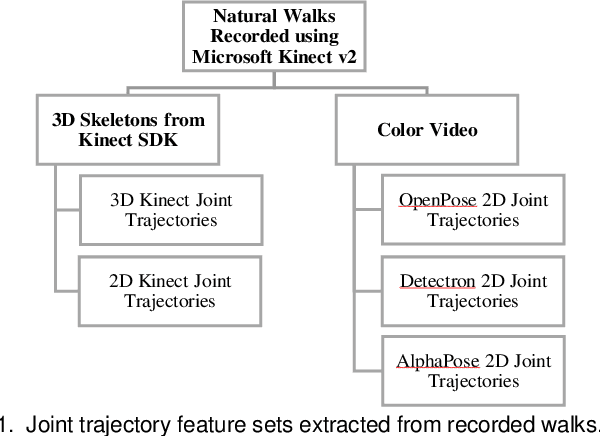
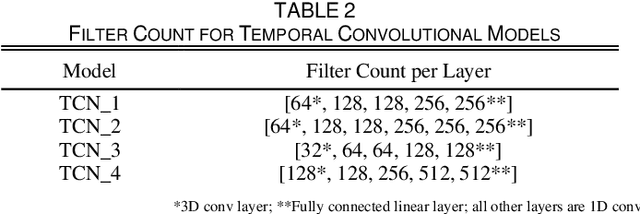


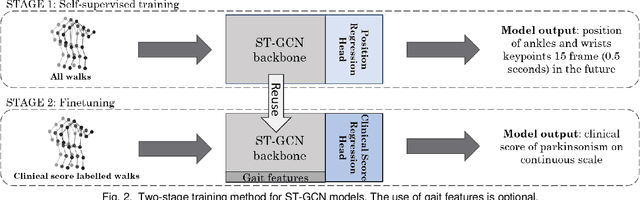
Abstract:Drug-induced parkinsonism affects many older adults with dementia, often causing gait disturbances. New advances in vision-based human pose-estimation have opened possibilities for frequent and unobtrusive analysis of gait in residential settings. This work proposes novel spatial-temporal graph convolutional network (ST-GCN) architectures and training procedures to predict clinical scores of parkinsonism in gait from video of individuals with dementia. We propose a two-stage training approach consisting of a self-supervised pretraining stage that encourages the ST-GCN model to learn about gait patterns before predicting clinical scores in the finetuning stage. The proposed ST-GCN models are evaluated on joint trajectories extracted from video and are compared against traditional (ordinal, linear, random forest) regression models and temporal convolutional network baselines. Three 2D human pose-estimation libraries (OpenPose, Detectron, AlphaPose) and the Microsoft Kinect (2D and 3D) are used to extract joint trajectories of 4787 natural walking bouts from 53 older adults with dementia. A subset of 399 walks from 14 participants is annotated with scores of parkinsonism severity on the gait criteria of the Unified Parkinson's Disease Rating Scale (UPDRS) and the Simpson-Angus Scale (SAS). Our results demonstrate that ST-GCN models operating on 3D joint trajectories extracted from the Kinect consistently outperform all other models and feature sets. Prediction of parkinsonism scores in natural walking bouts of unseen participants remains a challenging task, with the best models achieving macro-averaged F1-scores of 0.53 +/- 0.03 and 0.40 +/- 0.02 for UPDRS-gait and SAS-gait, respectively. Pre-trained model and demo code for this work is available: https://github.com/TaatiTeam/stgcn_parkinsonism_prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge