Yana Yunusova
Concurrent Validity of Automatic Speech and Pause Measures During Passage Reading in ALS
Aug 22, 2022
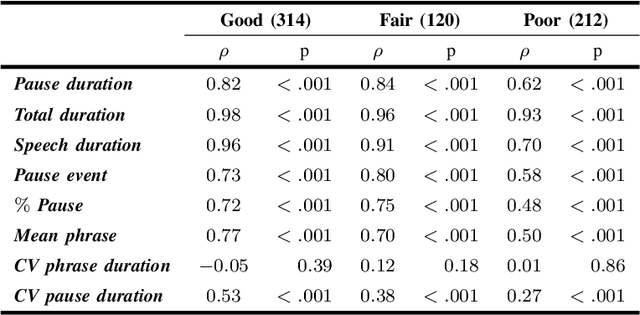
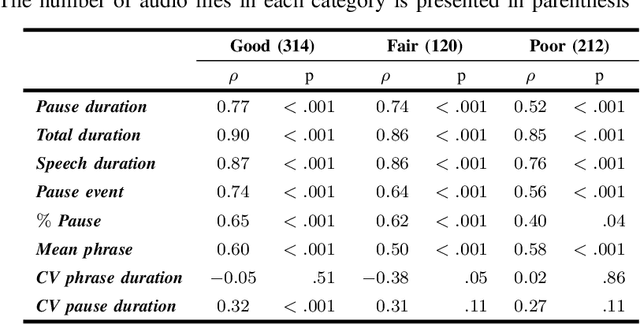
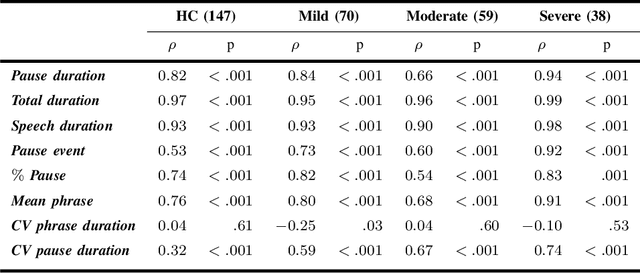
Abstract:The analysis of speech measures in individuals with amyotrophic lateral sclerosis (ALS) can provide essential information for early diagnosis and tracking disease progression. However, current methods for extracting speech and pause features are manual or semi-automatic, which makes them time-consuming and labour-intensive. The advent of speech-text alignment algorithms provides an opportunity for inexpensive, automated, and accurate analysis of speech measures in individuals with ALS. There is a need to validate speech and pause features calculated by these algorithms against current gold standard methods. In this study, we extracted 8 speech/pause features from 646 audio files of individuals with ALS and healthy controls performing passage reading. Two pretrained forced alignment models - one using transformers and another using a Gaussian mixture / hidden Markov architecture - were used for automatic feature extraction. The results were then validated against semi-automatic speech/pause analysis software, with further subgroup analyses based on audio quality and disease severity. Features extracted using transformer-based forced alignment had the highest agreement with gold standards, including in terms of audio quality and disease severity. This study lays the groundwork for future intelligent diagnostic support systems for clinicians, and for novel methods of tracking disease progression remotely from home.
Automated Temporal Segmentation of Orofacial Assessment Videos
Aug 22, 2022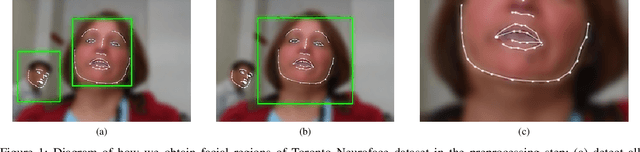
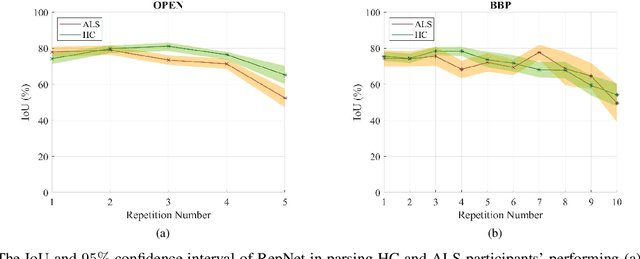
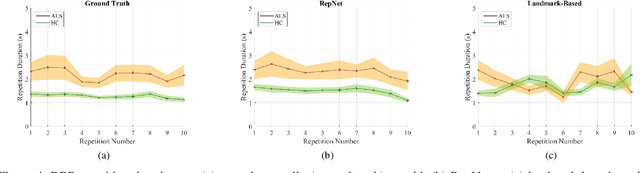
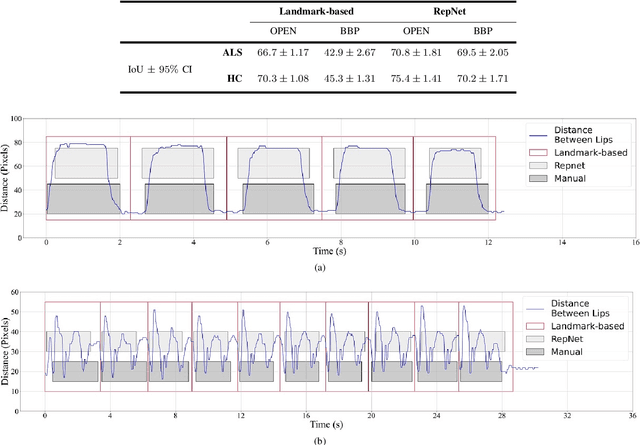
Abstract:Computer vision techniques can help automate or partially automate clinical examination of orofacial impairments to provide accurate and objective assessments. Towards the development of such automated systems, we evaluated two approaches to detect and temporally segment (parse) repetitions in orofacial assessment videos. Recorded videos of participants with amyotrophic lateral sclerosis (ALS) and healthy control (HC) individuals were obtained from the Toronto NeuroFace Dataset. Two approaches for repetition detection and parsing were examined: one based on engineered features from tracked facial landmarks and peak detection in the distance between the vermilion-cutaneous junction of the upper and lower lips (baseline analysis), and another using a pre-trained transformer-based deep learning model called RepNet (Dwibedi et al, 2020), which automatically detects periodicity, and parses periodic and semi-periodic repetitions in video data. In experimental evaluation of two orofacial assessments tasks, - repeating maximum mouth opening (OPEN) and repeating the sentence "Buy Bobby a Puppy" (BBP) - RepNet provided better parsing than the landmark-based approach, quantified by higher mean intersection-over-union (IoU) with respect to ground truth manual parsing. Automated parsing using RepNet also clearly separated HC and ALS participants based on the duration of BBP repetitions, whereas the landmark-based method could not.
Estimation of Orofacial Kinematics in Parkinson's Disease: Comparison of 2D and 3D Markerless Systems for Motion Tracking
Mar 18, 2020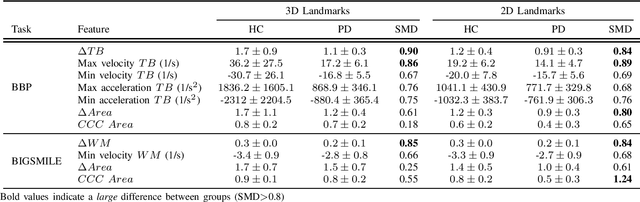
Abstract:Orofacial deficits are common in people with Parkinson's disease (PD) and their evolution might represent an important biomarker of disease progression. We are developing an automated system for assessment of orofacial function in PD that can be used in-home or in-clinic and can provide useful and objective clinical information that informs disease management. Our current approach relies on color and depth cameras for the estimation of 3D facial movements. However, depth cameras are not commonly available, might be expensive, and require specialized software for control and data processing. The objective of this paper was to evaluate if depth cameras are needed to differentiate between healthy controls and PD patients based on features extracted from orofacial kinematics. Results indicate that 2D features, extracted from color cameras only, are as informative as 3D features, extracted from color and depth cameras, differentiating healthy controls from PD patients. These results pave the way for the development of a universal system for automatic and objective assessment of orofacial function in PD.
Toward an Automatic System for Computer-Aided Assessment in Facial Palsy
Oct 25, 2019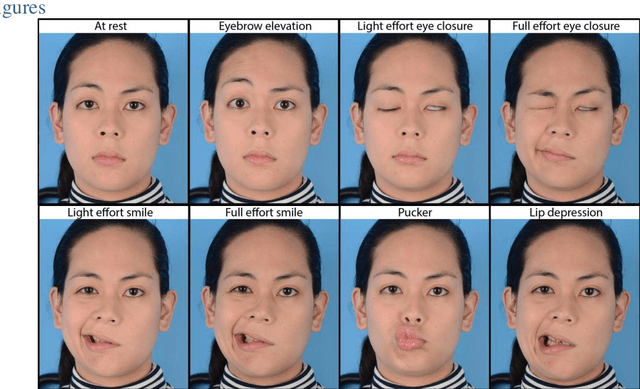
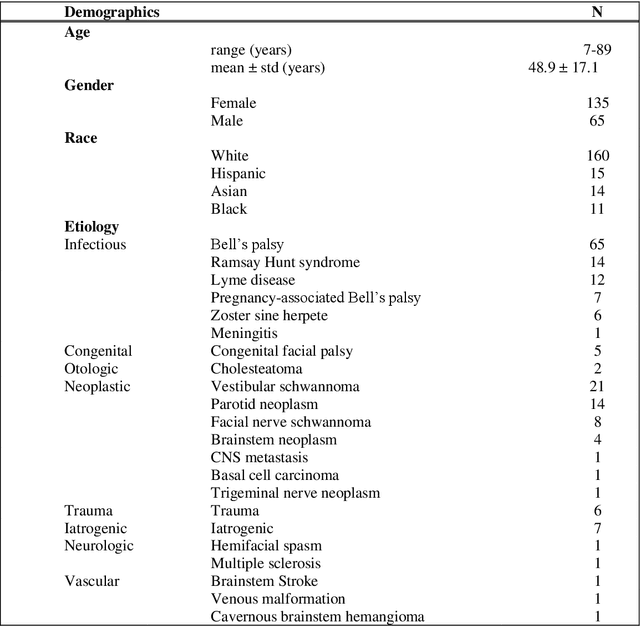
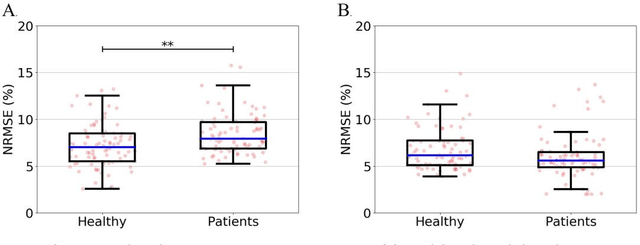
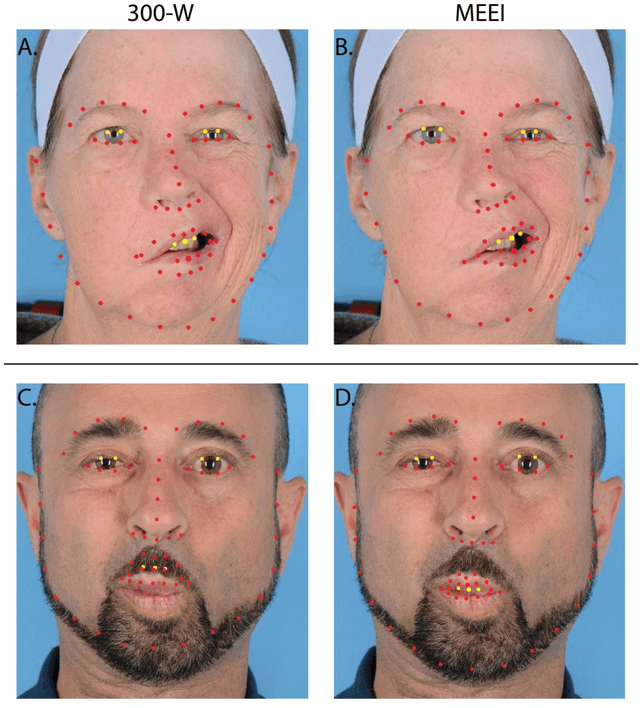
Abstract:Importance: Machine learning (ML) approaches to facial landmark localization carry great clinical potential for quantitative assessment of facial function as they enable high-throughput automated quantification of relevant facial metrics from photographs. However, translation from research settings to clinical applications requires important improvements. Objective: To develop an ML algorithm for accurate facial landmarks localization in photographs of facial palsy patients, and use it as part of an automated computer-aided diagnosis system. Design, Setting, and Participants: Facial landmarks were manually localized in portrait photographs of eight expressions obtained from 200 facial palsy patients and 10 controls. A novel ML model for automated facial landmark localization was trained using this disease-specific database. Model output was compared to manual annotations and the output of a model trained using a larger database consisting only of healthy subjects. Model accuracy was evaluated by the normalized root mean square error (NRMSE) between algorithms' prediction and manual annotations. Results: Publicly available algorithms provide poor results when applied to patients compared to healthy controls (NRMSE, 8.56 +/- 2.16 vs. 7.09 +/- 2.34, p << 0.01). We found significant improvement in facial landmark localization accuracy for the clinical population when using a model trained with a relatively small number patients' photographs (1440) compared to a model trained using several thousand more images of healthy faces (NRMSE, 6.03 +/- 2.43 vs. 8.56 +/- 2.16, p << 0.01). Conclusions: Retraining a landmark detection model with a small number of clinical images significantly improved landmark detection performance in frontal view photographs of the clinical population. These results represent the first steps towards an automatic system for computer-aided assessment in facial palsy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge