Anna Maria Anniciello
BRACS: A Dataset for BReAst Carcinoma Subtyping in H&E Histology Images
Nov 08, 2021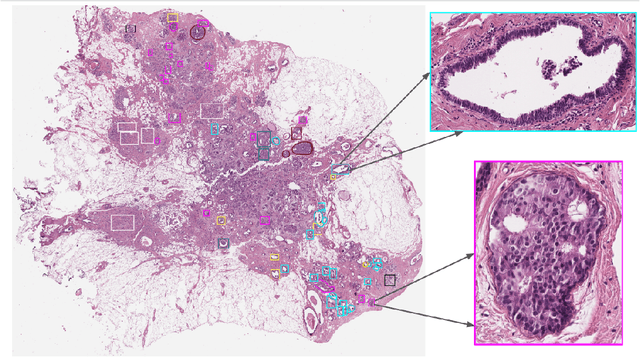
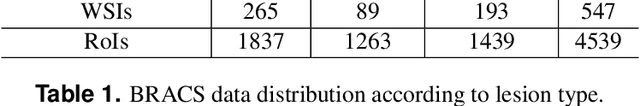

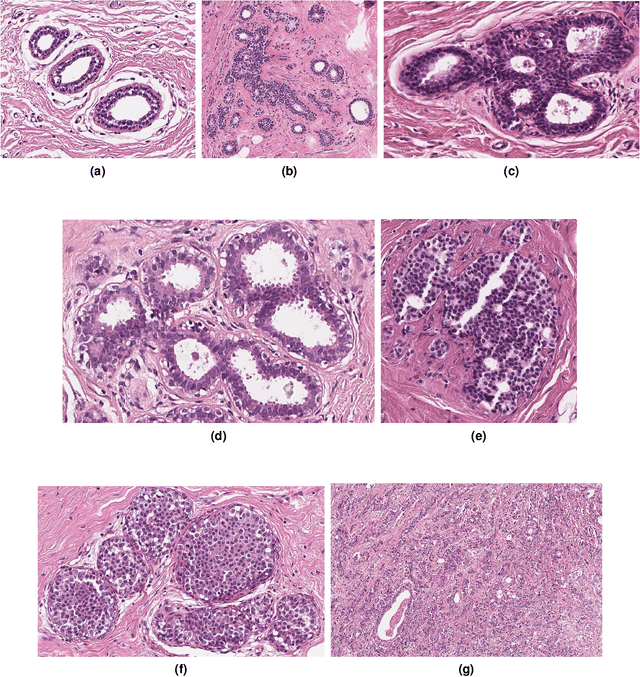
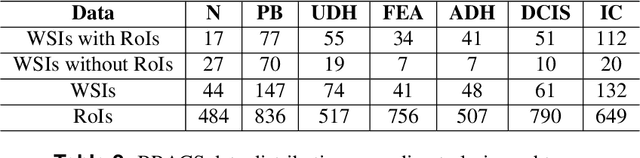
Abstract:Breast cancer is the most commonly diagnosed cancer and registers the highest number of deaths for women with cancer. Recent advancements in diagnostic activities combined with large-scale screening policies have significantly lowered the mortality rates for breast cancer patients. However, the manual inspection of tissue slides by the pathologists is cumbersome, time-consuming, and is subject to significant inter- and intra-observer variability. Recently, the advent of whole-slide scanning systems have empowered the rapid digitization of pathology slides, and enabled to develop digital workflows. These advances further enable to leverage Artificial Intelligence (AI) to assist, automate, and augment pathological diagnosis. But the AI techniques, especially Deep Learning (DL), require a large amount of high-quality annotated data to learn from. Constructing such task-specific datasets poses several challenges, such as, data-acquisition level constrains, time-consuming and expensive annotations, and anonymization of private information. In this paper, we introduce the BReAst Carcinoma Subtyping (BRACS) dataset, a large cohort of annotated Hematoxylin & Eosin (H&E)-stained images to facilitate the characterization of breast lesions. BRACS contains 547 Whole-Slide Images (WSIs), and 4539 Regions of Interest (ROIs) extracted from the WSIs. Each WSI, and respective ROIs, are annotated by the consensus of three board-certified pathologists into different lesion categories. Specifically, BRACS includes three lesion types, i.e., benign, malignant and atypical, which are further subtyped into seven categories. It is, to the best of our knowledge, the largest annotated dataset for breast cancer subtyping both at WSI- and ROI-level. Further, by including the understudied atypical lesions, BRACS offers an unique opportunity for leveraging AI to better understand their characteristics.
Hierarchical Graph Representations in Digital Pathology
Mar 17, 2021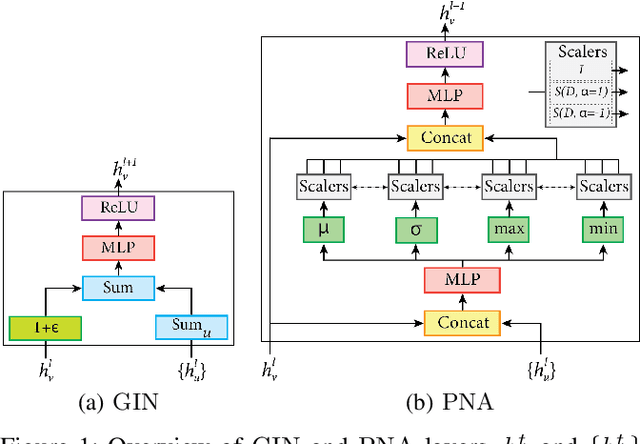
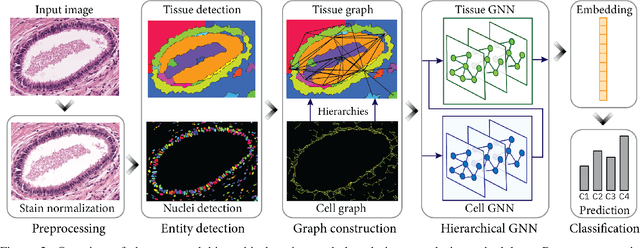
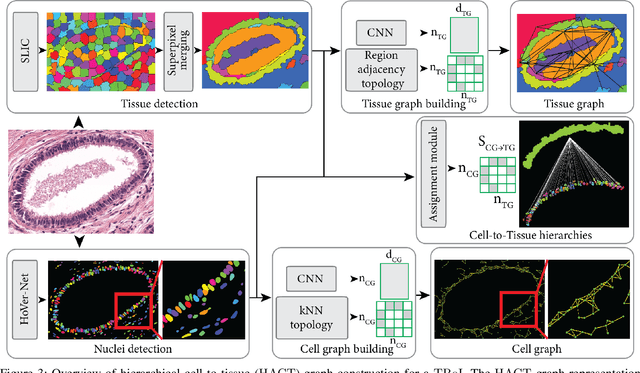
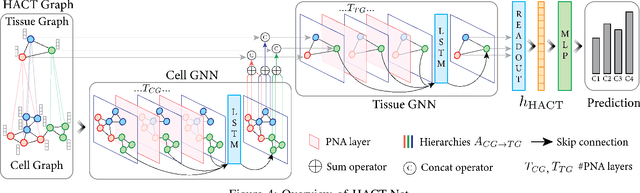
Abstract:Cancer diagnosis, prognosis, and therapy response predictions from tissue specimens highly depend on the phenotype and topological distribution of constituting histological entities. Thus, adequate tissue representations for encoding histological entities is imperative for computer aided cancer patient care. To this end, several approaches have leveraged cell-graphs that encode cell morphology and organization to denote the tissue information. These allow for utilizing machine learning to map tissue representations to tissue functionality to help quantify their relationship. Though cellular information is crucial, it is incomplete alone to comprehensively characterize complex tissue structure. We herein treat the tissue as a hierarchical composition of multiple types of histological entities from fine to coarse level, capturing multivariate tissue information at multiple levels. We propose a novel multi-level hierarchical entity-graph representation of tissue specimens to model hierarchical compositions that encode histological entities as well as their intra- and inter-entity level interactions. Subsequently, a graph neural network is proposed to operate on the hierarchical entity-graph representation to map the tissue structure to tissue functionality. Specifically, for input histology images we utilize well-defined cells and tissue regions to build HierArchical Cell-to-Tissue (HACT) graph representations, and devise HACT-Net, a graph neural network, to classify such HACT representations. As part of this work, we introduce the BReAst Carcinoma Subtyping (BRACS) dataset, a large cohort of H&E stained breast tumor images, to evaluate our proposed methodology against pathologists and state-of-the-art approaches. Through comparative assessment and ablation studies, our method is demonstrated to yield superior classification results compared to alternative methods as well as pathologists.
Quantifying Explainers of Graph Neural Networks in Computational Pathology
Nov 25, 2020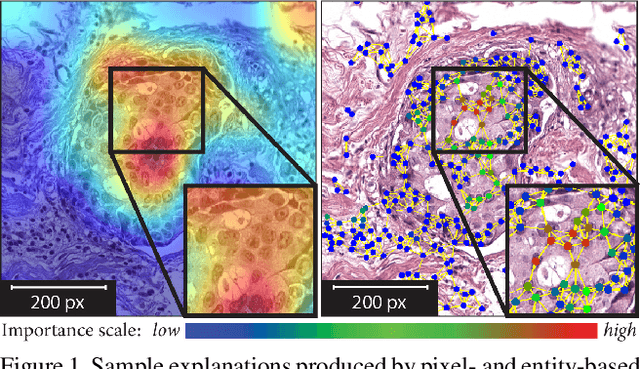
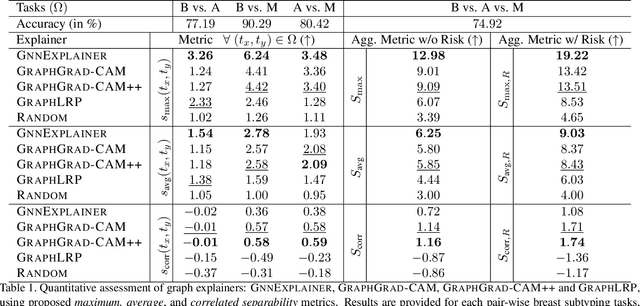
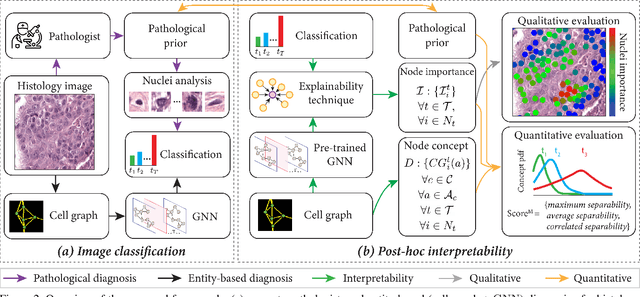
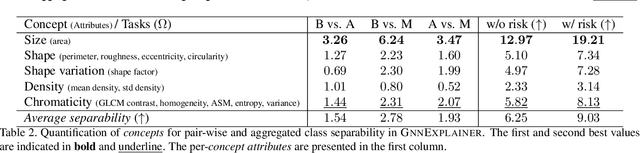
Abstract:Explainability of deep learning methods is imperative to facilitate their clinical adoption in digital pathology. However, popular deep learning methods and explainability techniques (explainers) based on pixel-wise processing disregard biological entities' notion, thus complicating comprehension by pathologists. In this work, we address this by adopting biological entity-based graph processing and graph explainers enabling explanations accessible to pathologists. In this context, a major challenge becomes to discern meaningful explainers, particularly in a standardized and quantifiable fashion. To this end, we propose herein a set of novel quantitative metrics based on statistics of class separability using pathologically measurable concepts to characterize graph explainers. We employ the proposed metrics to evaluate three types of graph explainers, namely the layer-wise relevance propagation, gradient-based saliency, and graph pruning approaches, to explain Cell-Graph representations for Breast Cancer Subtyping. The proposed metrics are also applicable in other domains by using domain-specific intuitive concepts. We validate the qualitative and quantitative findings on the BRACS dataset, a large cohort of breast cancer RoIs, by expert pathologists.
HACT-Net: A Hierarchical Cell-to-Tissue Graph Neural Network for Histopathological Image Classification
Jul 01, 2020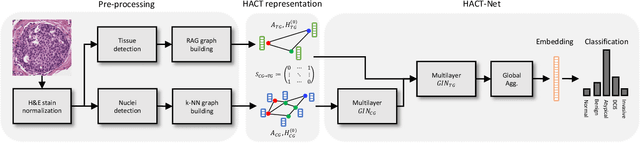
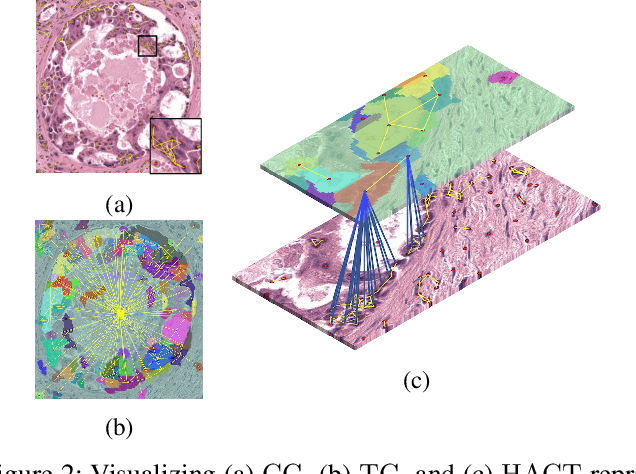
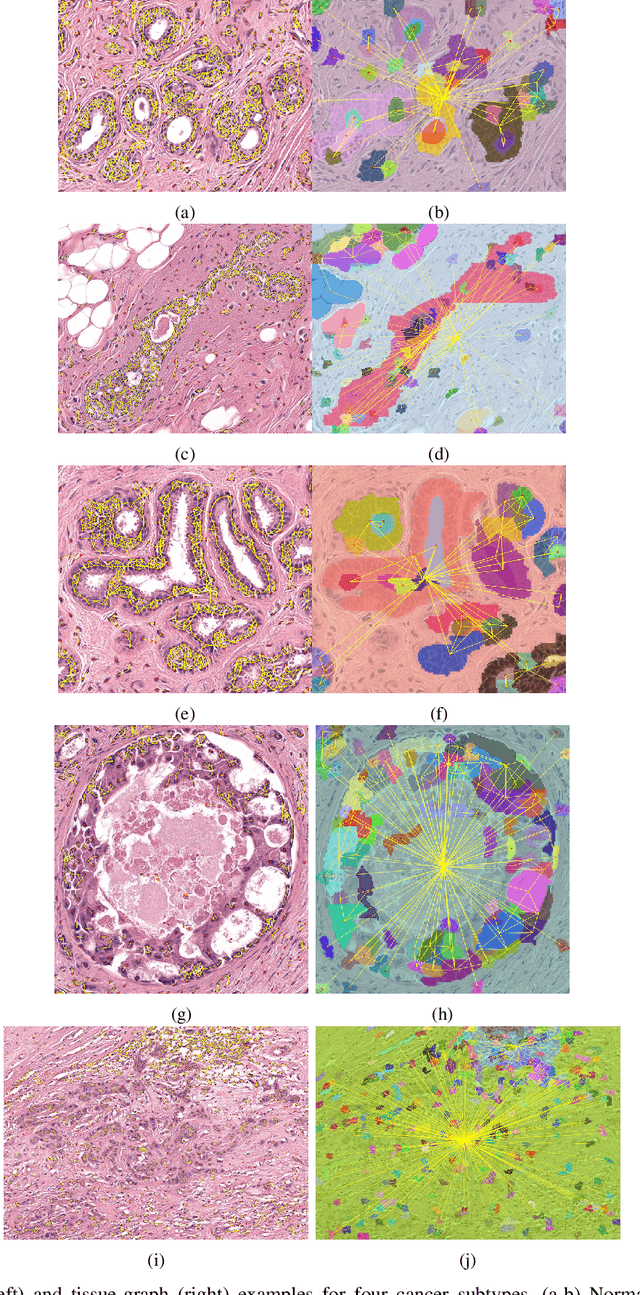
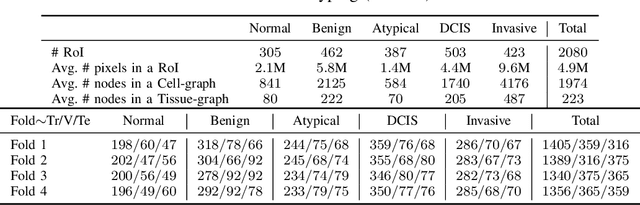
Abstract:Cancer diagnosis, prognosis, and therapeutic response prediction are heavily influenced by the relationship between the histopathological structures and the function of the tissue. Recent approaches acknowledging the structure-function relationship, have linked the structural and spatial patterns of cell organization in tissue via cell-graphs to tumor grades. Though cell organization is imperative, it is insufficient to entirely represent the histopathological structure. We propose a novel hierarchical cell-to-tissue-graph (HACT) representation to improve the structural depiction of the tissue. It consists of a low-level cell-graph, capturing cell morphology and interactions, a high-level tissue-graph, capturing morphology and spatial distribution of tissue parts, and cells-to-tissue hierarchies, encoding the relative spatial distribution of the cells with respect to the tissue distribution. Further, a hierarchical graph neural network (HACT-Net) is proposed to efficiently map the HACT representations to histopathological breast cancer subtypes. We assess the methodology on a large set of annotated tissue regions of interest from H\&E stained breast carcinoma whole-slides. Upon evaluation, the proposed method outperformed recent convolutional neural network and graph neural network approaches for breast cancer multi-class subtyping. The proposed entity-based topological analysis is more inline with the pathological diagnostic procedure of the tissue. It provides more command over the tissue modelling, therefore encourages the further inclusion of pathological priors into task-specific tissue representation.
Towards Explainable Graph Representations in Digital Pathology
Jul 01, 2020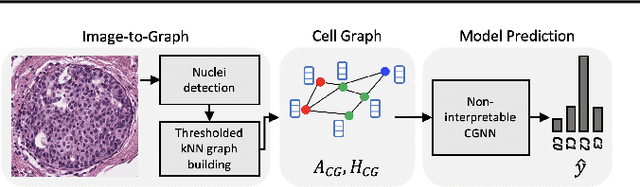
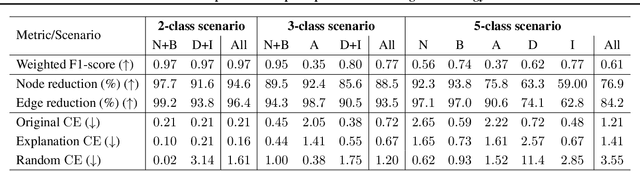
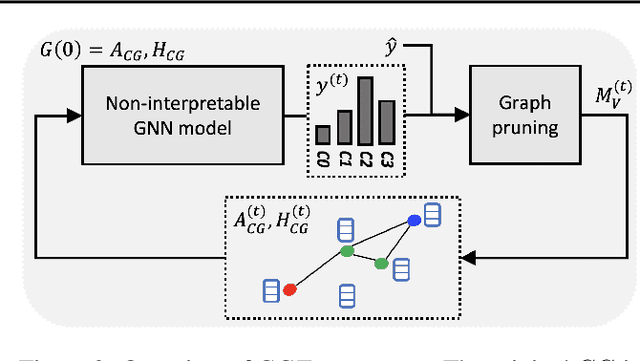
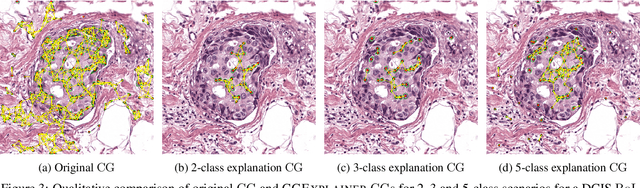
Abstract:Explainability of machine learning (ML) techniques in digital pathology (DP) is of great significance to facilitate their wide adoption in clinics. Recently, graph techniques encoding relevant biological entities have been employed to represent and assess DP images. Such paradigm shift from pixel-wise to entity-wise analysis provides more control over concept representation. In this paper, we introduce a post-hoc explainer to derive compact per-instance explanations emphasizing diagnostically important entities in the graph. Although we focus our analyses to cells and cellular interactions in breast cancer subtyping, the proposed explainer is generic enough to be extended to other topological representations in DP. Qualitative and quantitative analyses demonstrate the efficacy of the explainer in generating comprehensive and compact explanations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge