Andreas Maier
Pattern Recognition Lab, FAU Erlangen-Nürnberg, Germany
X-ray Scatter Estimation Using Deep Splines
Jan 22, 2021
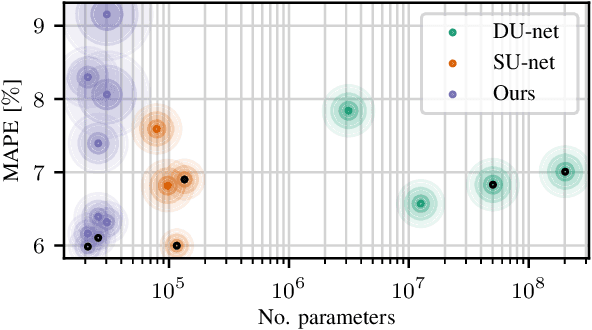
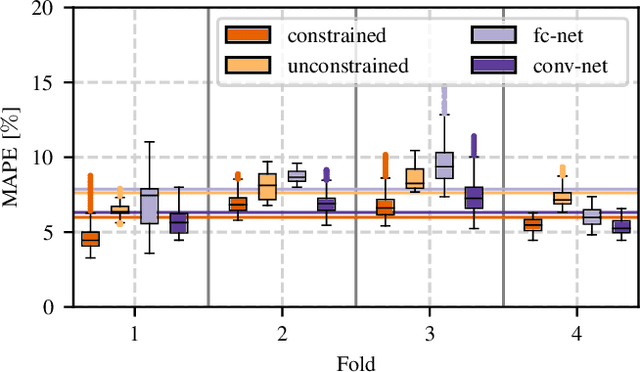
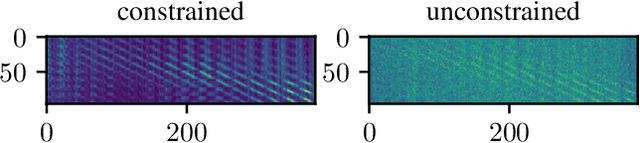
Abstract:Algorithmic X-ray scatter compensation is a desirable technique in flat-panel X-ray imaging and cone-beam computed tomography. State-of-the-art U-net based image translation approaches yielded promising results. As there are no physics constraints applied to the output of the U-Net, it cannot be ruled out that it yields spurious results. Unfortunately, those may be misleading in the context of medical imaging. To overcome this problem, we propose to embed B-splines as a known operator into neural networks. This inherently limits their predictions to well-behaved and smooth functions. In a study using synthetic head and thorax data as well as real thorax phantom data, we found that our approach performed on par with U-net when comparing both algorithms based on quantitative performance metrics. However, our approach not only reduces runtime and parameter complexity, but we also found it much more robust to unseen noise levels. While the U-net responded with visible artifacts, our approach preserved the X-ray signal's frequency characteristics.
Synthetic Glacier SAR Image Generation from Arbitrary Masks Using Pix2Pix Algorithm
Jan 14, 2021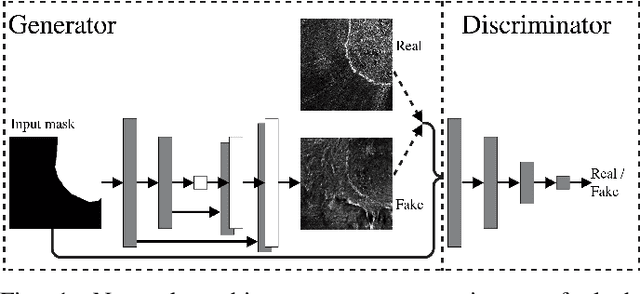
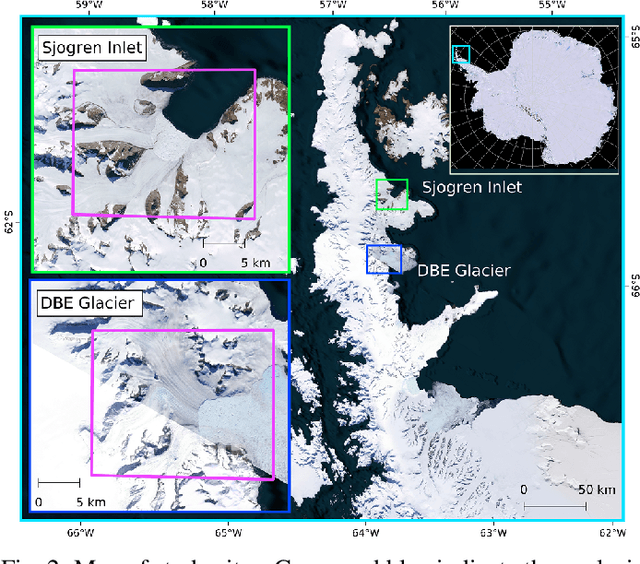
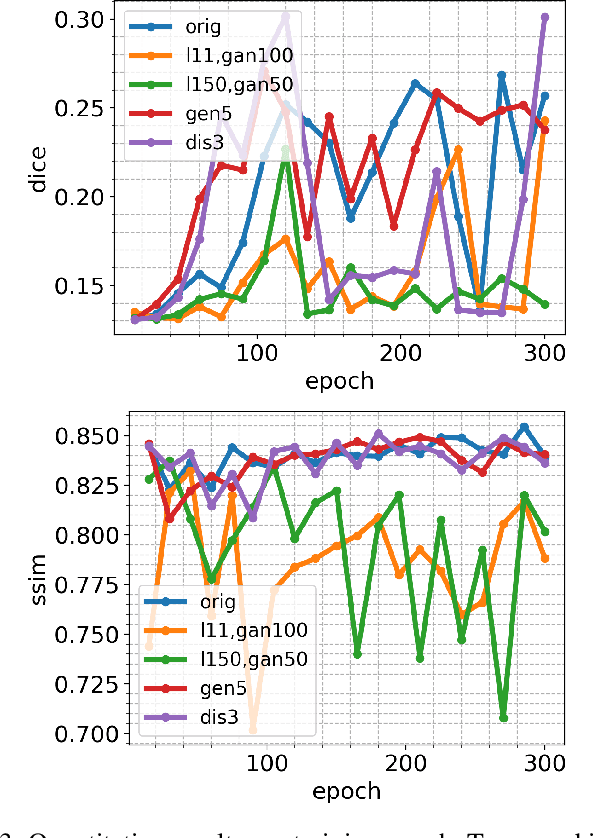

Abstract:Supervised machine learning requires a large amount of labeled data to achieve proper test results. However, generating accurately labeled segmentation maps on remote sensing imagery, including images from synthetic aperture radar (SAR), is tedious and highly subjective. In this work, we propose to alleviate the issue of limited training data by generating synthetic SAR images with the pix2pix algorithm. This algorithm uses conditional Generative Adversarial Networks (cGANs) to generate an artificial image while preserving the structure of the input. In our case, the input is a segmentation mask, from which a corresponding synthetic SAR image is generated. We present different models, perform a comparative study and demonstrate that this approach synthesizes convincing glaciers in SAR images with promising qualitative and quantitative results.
Learning to be EXACT, Cell Detection for Asthma on Partially Annotated Whole Slide Images
Jan 13, 2021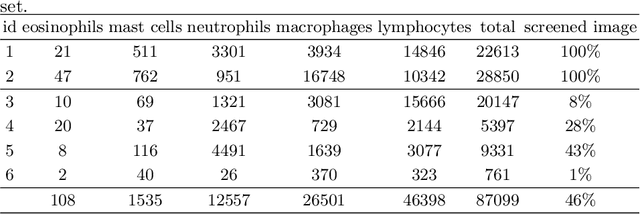
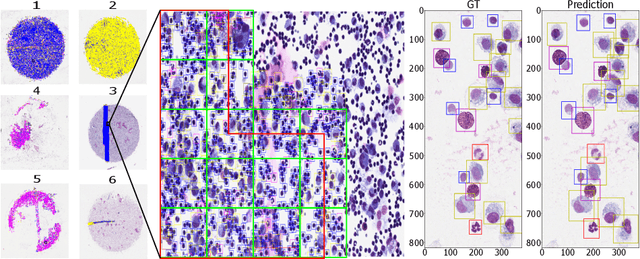
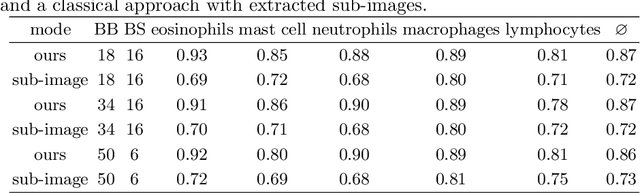
Abstract:Asthma is a chronic inflammatory disorder of the lower respiratory tract and naturally occurs in humans and animals including horses. The annotation of an asthma microscopy whole slide image (WSI) is an extremely labour-intensive task due to the hundreds of thousands of cells per WSI. To overcome the limitation of annotating WSI incompletely, we developed a training pipeline which can train a deep learning-based object detection model with partially annotated WSIs and compensate class imbalances on the fly. With this approach we can freely sample from annotated WSIs areas and are not restricted to fully annotated extracted sub-images of the WSI as with classical approaches. We evaluated our pipeline in a cross-validation setup with a fixed training set using a dataset of six equine WSIs of which four are partially annotated and used for training, and two fully annotated WSI are used for validation and testing. Our WSI-based training approach outperformed classical sub-image-based training methods by up to 15\% $mAP$ and yielded human-like performance when compared to the annotations of ten trained pathologists.
Coronary Plaque Analysis for CT Angiography Clinical Research
Jan 11, 2021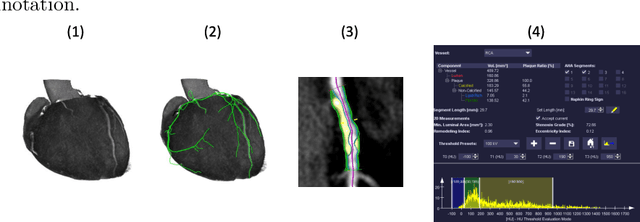
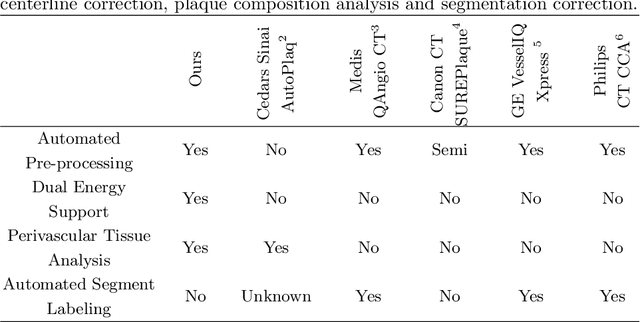
Abstract:The analysis of plaque deposits in the coronary vasculature is an important topic in current clinical research. From a technical side mostly new algorithms for different sub tasks - e.g. centerline extraction or vessel/plaque segmentation - are proposed. However, to enable clinical research with the help of these algorithms, a software solution, which enables manual correction, comprehensive visual feedback and tissue analysis capabilities, is needed. Therefore, we want to present such an integrated software solution. It is able to perform robust automatic centerline extraction and inner and outer vessel wall segmentation, while providing easy to use manual correction tools. Also, it allows for annotation of lesions along the centerlines, which can be further analyzed regarding their tissue composition. Furthermore, it enables research in upcoming technologies and research directions: it does support dual energy CT scans with dedicated plaque analysis and the quantification of the fatty tissue surrounding the vasculature, also in automated set-ups.
Bayesian U-Net for Segmenting Glaciers in SAR Imagery
Jan 08, 2021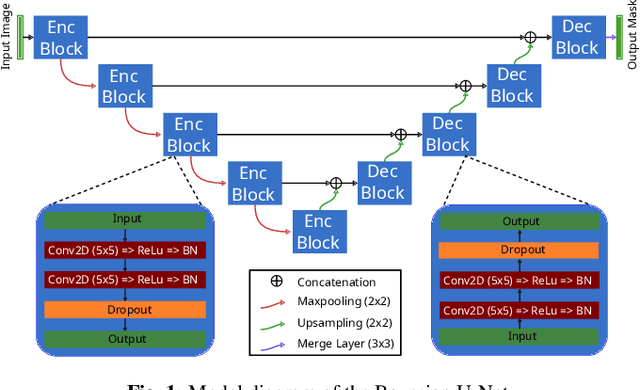
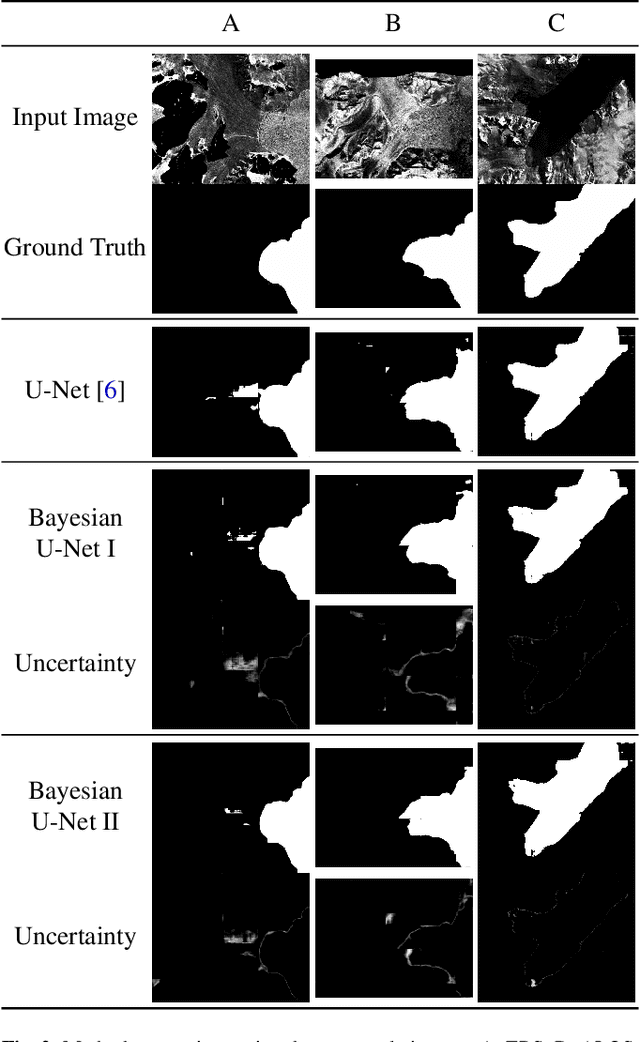
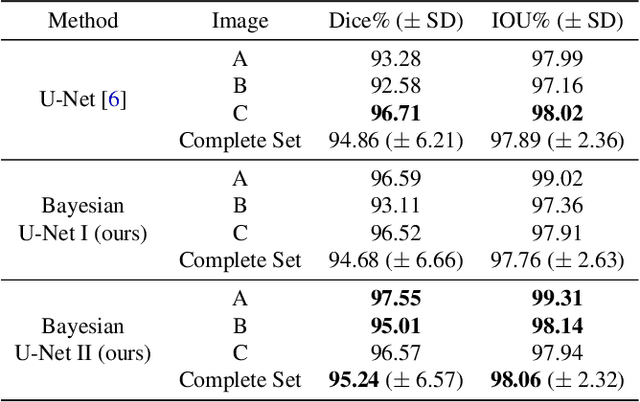

Abstract:Fluctuations of the glacier calving front have an important influence over the ice flow of whole glacier systems. It is therefore important to precisely monitor the position of the calving front. However, the manual delineation of SAR images is a difficult, laborious and subjective task. Convolutional neural networks have previously shown promising results in automating the glacier segmentation in SAR images, making them desirable for further exploration of their possibilities. In this work, we propose to compute uncertainty and use it in an Uncertainty Optimization regime as a novel two-stage process. By using dropout as a random sampling layer in a U-Net architecture, we create a probabilistic Bayesian Neural Network. With several forward passes, we create a sampling distribution, which can estimate the model uncertainty for each pixel in the segmentation mask. The additional uncertainty map information can serve as a guideline for the experts in the manual annotation of the data. Furthermore, feeding the uncertainty map to the network leads to 95.24% Dice similarity, which is an overall improvement in the segmentation performance compared to the state-of-the-art deterministic U-Net-based glacier segmentation pipelines.
Glacier Calving Front Segmentation Using Attention U-Net
Jan 08, 2021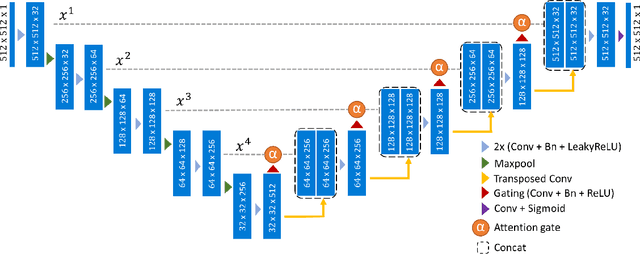
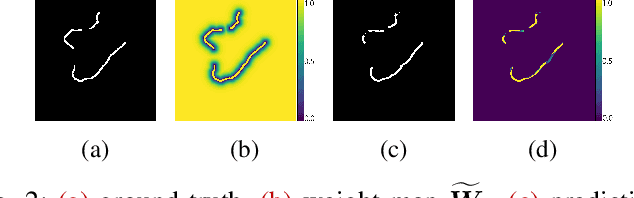
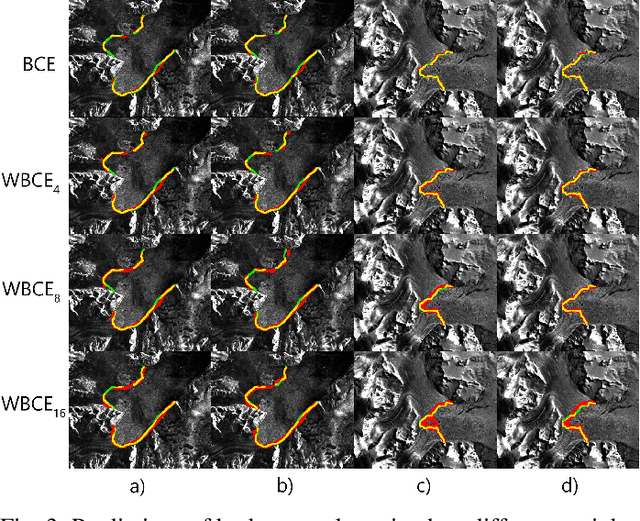
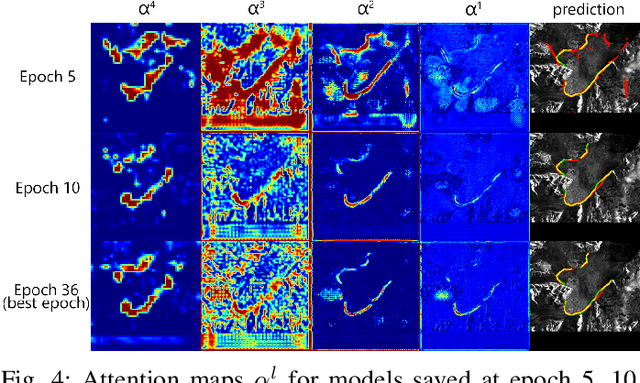
Abstract:An essential climate variable to determine the tidewater glacier status is the location of the calving front position and the separation of seasonal variability from long-term trends. Previous studies have proposed deep learning-based methods to semi-automatically delineate the calving fronts of tidewater glaciers. They used U-Net to segment the ice and non-ice regions and extracted the calving fronts in a post-processing step. In this work, we show a method to segment the glacier calving fronts from SAR images in an end-to-end fashion using Attention U-Net. The main objective is to investigate the attention mechanism in this application. Adding attention modules to the state-of-the-art U-Net network lets us analyze the learning process by extracting its attention maps. We use these maps as a tool to search for proper hyperparameters and loss functions in order to generate higher qualitative results. Our proposed attention U-Net performs comparably to the standard U-Net while providing additional insight into those regions on which the network learned to focus more. In the best case, the attention U-Net achieves a 1.5% better Dice score compared to the canonical U-Net with a glacier front line prediction certainty of up to 237.12 meters.
How Many Annotators Do We Need? -- A Study on the Influence of Inter-Observer Variability on the Reliability of Automatic Mitotic Figure Assessment
Jan 08, 2021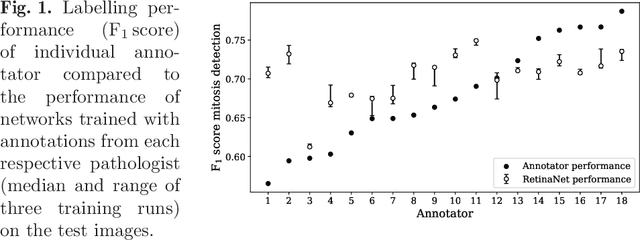
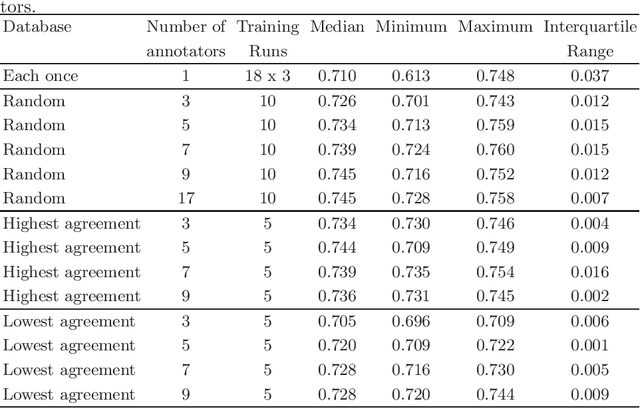
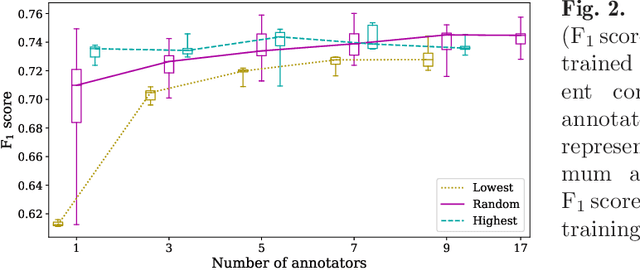
Abstract:Density of mitotic figures in histologic sections is a prognostically relevant characteristic for many tumours. Due to high inter-pathologist variability, deep learning-based algorithms are a promising solution to improve tumour prognostication. Pathologists are the gold standard for database development, however, labelling errors may hamper development of accurate algorithms. In the present work we evaluated the benefit of multi-expert consensus (n = 3, 5, 7, 9, 11) on algorithmic performance. While training with individual databases resulted in highly variable F$_1$ scores, performance was notably increased and more consistent when using the consensus of three annotators. Adding more annotators only resulted in minor improvements. We conclude that databases by few pathologists and high label accuracy may be the best compromise between high algorithmic performance and time investment.
Dataset on Bi- and Multi-Nucleated Tumor Cells in Canine Cutaneous Mast Cell Tumors
Jan 05, 2021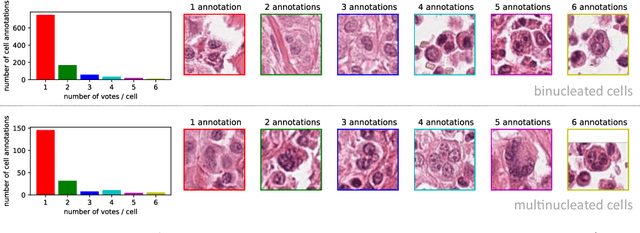
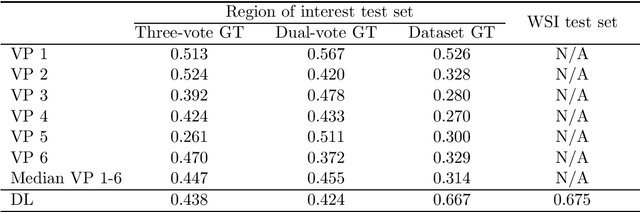
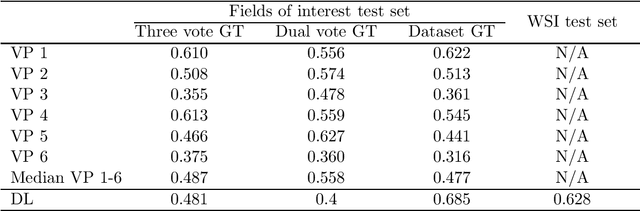
Abstract:Tumor cells with two nuclei (binucleated cells, BiNC) or more nuclei (multinucleated cells, MuNC) indicate an increased amount of cellular genetic material which is thought to facilitate oncogenesis, tumor progression and treatment resistance. In canine cutaneous mast cell tumors (ccMCT), binucleation and multinucleation are parameters used in cytologic and histologic grading schemes (respectively) which correlate with poor patient outcome. For this study, we created the first open source data-set with 19,983 annotations of BiNC and 1,416 annotations of MuNC in 32 histological whole slide images of ccMCT. Labels were created by a pathologist and an algorithmic-aided labeling approach with expert review of each generated candidate. A state-of-the-art deep learning-based model yielded an $F_1$ score of 0.675 for BiNC and 0.623 for MuNC on 11 test whole slide images. In regions of interest ($2.37 mm^2$) extracted from these test images, 6 pathologists had an object detection performance between 0.270 - 0.526 for BiNC and 0.316 - 0.622 for MuNC, while our model archived an $F_1$ score of 0.667 for BiNC and 0.685 for MuNC. This open dataset can facilitate development of automated image analysis for this task and may thereby help to promote standardization of this facet of histologic tumor prognostication.
2-D Respiration Navigation Framework for 3-D Continuous Cardiac Magnetic Resonance Imaging
Dec 26, 2020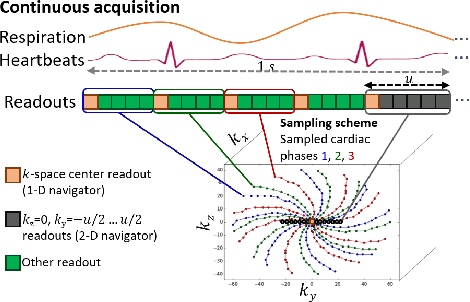
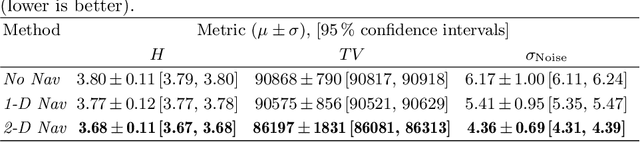

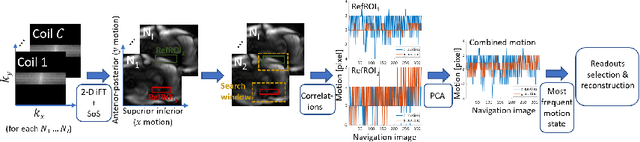
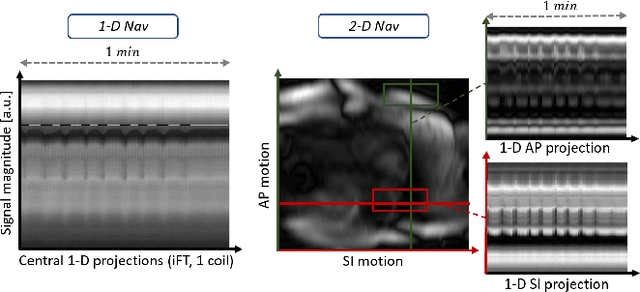
Abstract:Continuous protocols for cardiac magnetic resonance imaging enable sampling of the cardiac anatomy simultaneously resolved into cardiac phases. To avoid respiration artifacts, associated motion during the scan has to be compensated for during reconstruction. In this paper, we propose a sampling adaption to acquire 2-D respiration information during a continuous scan. Further, we develop a pipeline to extract the different respiration states from the acquired signals, which are used to reconstruct data from one respiration phase. Our results show the benefit of the proposed workflow on the image quality compared to no respiration compensation, as well as a previous 1-D respiration navigation approach.
Spatio-temporal Multi-task Learning for Cardiac MRI Left Ventricle Quantification
Dec 24, 2020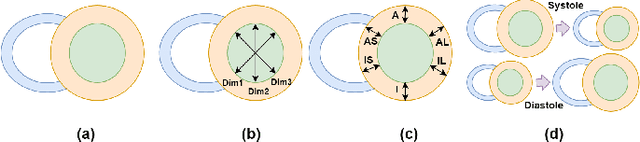
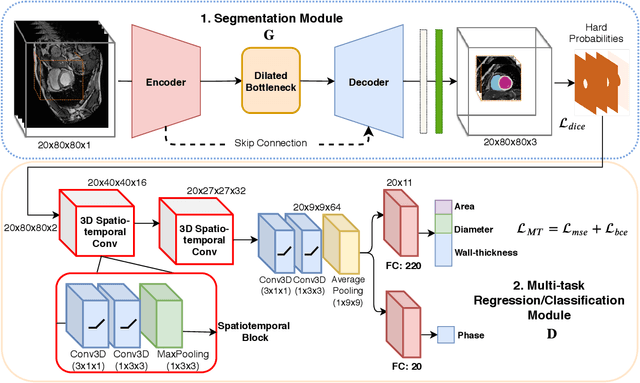
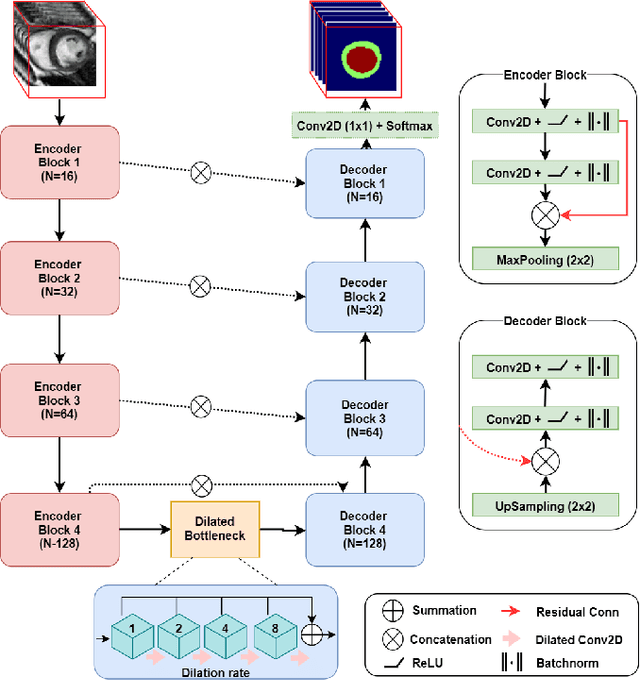
Abstract:Quantitative assessment of cardiac left ventricle (LV) morphology is essential to assess cardiac function and improve the diagnosis of different cardiovascular diseases. In current clinical practice, LV quantification depends on the measurement of myocardial shape indices, which is usually achieved by manual contouring of the endo- and epicardial. However, this process subjected to inter and intra-observer variability, and it is a time-consuming and tedious task. In this paper, we propose a spatio-temporal multi-task learning approach to obtain a complete set of measurements quantifying cardiac LV morphology, regional-wall thickness (RWT), and additionally detecting the cardiac phase cycle (systole and diastole) for a given 3D Cine-magnetic resonance (MR) image sequence. We first segment cardiac LVs using an encoder-decoder network and then introduce a multitask framework to regress 11 LV indices and classify the cardiac phase, as parallel tasks during model optimization. The proposed deep learning model is based on the 3D spatio-temporal convolutions, which extract spatial and temporal features from MR images. We demonstrate the efficacy of the proposed method using cine-MR sequences of 145 subjects and comparing the performance with other state-of-the-art quantification methods. The proposed method obtained high prediction accuracy, with an average mean absolute error (MAE) of 129 $mm^2$, 1.23 $mm$, 1.76 $mm$, Pearson correlation coefficient (PCC) of 96.4%, 87.2%, and 97.5% for LV and myocardium (Myo) cavity regions, 6 RWTs, 3 LV dimensions, and an error rate of 9.0\% for phase classification. The experimental results highlight the robustness of the proposed method, despite varying degrees of cardiac morphology, image appearance, and low contrast in the cardiac MR sequences.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge