Andreas Maier
Pattern Recognition Lab, FAU Erlangen-Nürnberg, Germany
Initial Investigations Towards Non-invasive Monitoring of Chronic Wound Healing Using Deep Learning and Ultrasound Imaging
Jan 25, 2022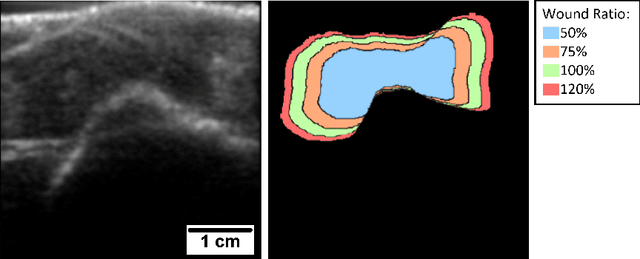

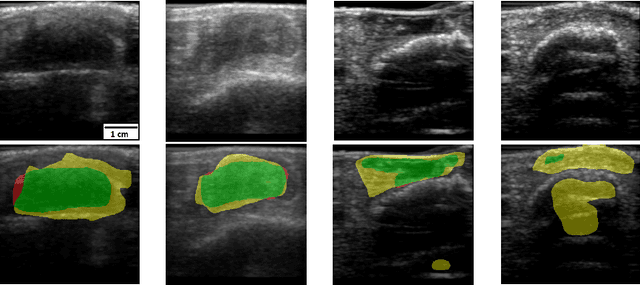
Abstract:Chronic wounds including diabetic and arterial/venous insufficiency injuries have become a major burden for healthcare systems worldwide. Demographic changes suggest that wound care will play an even bigger role in the coming decades. Predicting and monitoring response to therapy in wound care is currently largely based on visual inspection with little information on the underlying tissue. Thus, there is an urgent unmet need for innovative approaches that facilitate personalized diagnostics and treatments at the point-of-care. It has been recently shown that ultrasound imaging can monitor response to therapy in wound care, but this work required onerous manual image annotations. In this study, we present initial results of a deep learning-based automatic segmentation of cross-sectional wound size in ultrasound images and identify requirements and challenges for future research on this application. Evaluation of the segmentation results underscores the potential of the proposed deep learning approach to complement non-invasive imaging with Dice scores of 0.34 (U-Net, FCN) and 0.27 (ResNet-U-Net) but also highlights the need for improving robustness further. We conclude that deep learning-supported analysis of non-invasive ultrasound images is a promising area of research to automatically extract cross-sectional wound size and depth information with potential value in monitoring response to therapy.
Ultra Low-Parameter Denoising: Trainable Bilateral Filter Layers in Computed Tomography
Jan 25, 2022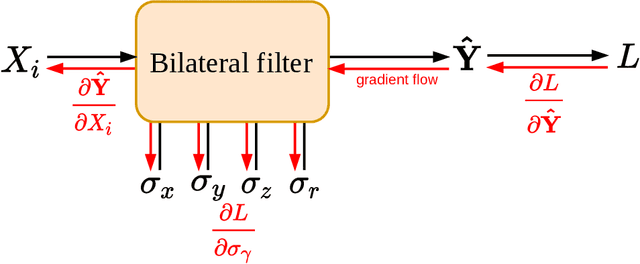
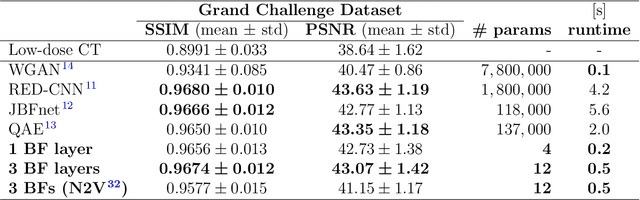
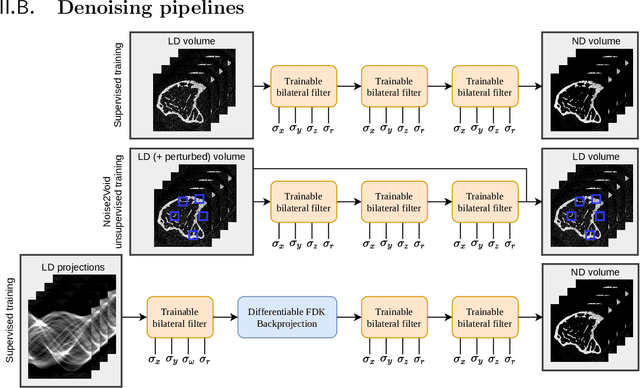
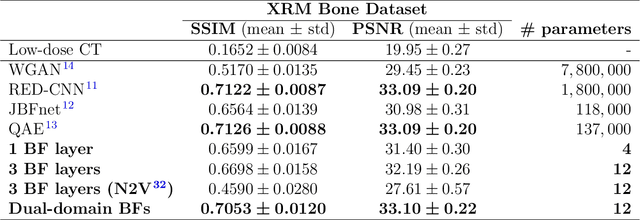
Abstract:Computed tomography is widely used as an imaging tool to visualize three-dimensional structures with expressive bone-soft tissue contrast. However, CT resolution and radiation dose are tightly entangled, highlighting the importance of low-dose CT combined with sophisticated denoising algorithms. Most data-driven denoising techniques are based on deep neural networks and, therefore, contain hundreds of thousands of trainable parameters, making them incomprehensible and prone to prediction failures. Developing understandable and robust denoising algorithms achieving state-of-the-art performance helps to minimize radiation dose while maintaining data integrity. This work presents an open-source CT denoising framework based on the idea of bilateral filtering. We propose a bilateral filter that can be incorporated into a deep learning pipeline and optimized in a purely data-driven way by calculating the gradient flow toward its hyperparameters and its input. Denoising in pure image-to-image pipelines and across different domains such as raw detector data and reconstructed volume, using a differentiable backprojection layer, is demonstrated. Although only using three spatial parameters and one range parameter per filter layer, the proposed denoising pipelines can compete with deep state-of-the-art denoising architectures with several hundred thousand parameters. Competitive denoising performance is achieved on x-ray microscope bone data (0.7053 and 33.10) and the 2016 Low Dose CT Grand Challenge dataset (0.9674 and 43.07) in terms of SSIM and PSNR. Due to the extremely low number of trainable parameters with well-defined effect, prediction reliance and data integrity is guaranteed at any time in the proposed pipelines, in contrast to most other deep learning-based denoising architectures.
Superpixel Pre-Segmentation of HER2 Slides for Efficient Annotation
Jan 19, 2022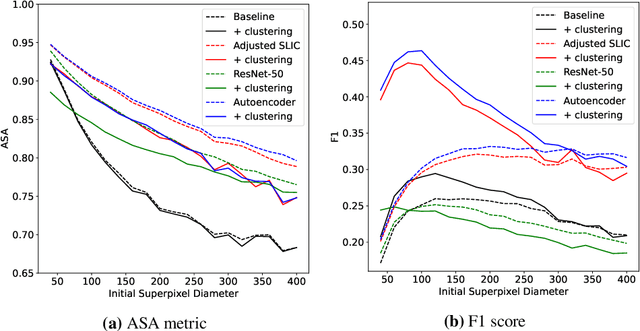
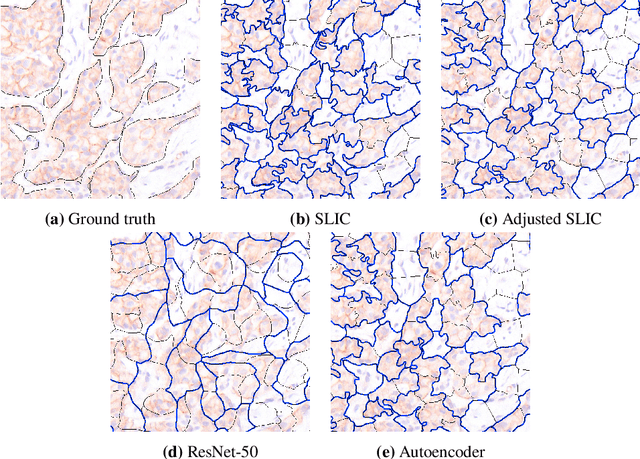
Abstract:Supervised deep learning has shown state-of-the-art performance for medical image segmentation across different applications, including histopathology and cancer research; however, the manual annotation of such data is extremely laborious. In this work, we explore the use of superpixel approaches to compute a pre-segmentation of HER2 stained images for breast cancer diagnosis that facilitates faster manual annotation and correction in a second step. Four methods are compared: Standard Simple Linear Iterative Clustering (SLIC) as a baseline, a domain adapted SLIC, and superpixels based on feature embeddings of a pretrained ResNet-50 and a denoising autoencoder. To tackle oversegmentation, we propose to hierarchically merge superpixels, based on their content in the respective feature space. When evaluating the approaches on fully manually annotated images, we observe that the autoencoder-based superpixels achieve a 23% increase in boundary F1 score compared to the baseline SLIC superpixels. Furthermore, the boundary F1 score increases by 73% when hierarchical clustering is applied on the adapted SLIC and the autoencoder-based superpixels. These evaluations show encouraging first results for a pre-segmentation for efficient manual refinement without the need for an initial set of annotated training data.
Learned Cone-Beam CT Reconstruction Using Neural Ordinary Differential Equations
Jan 19, 2022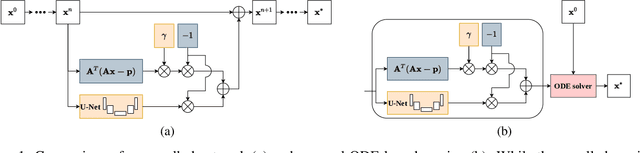
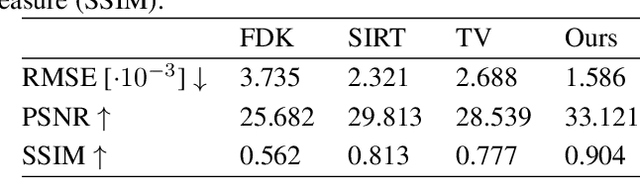
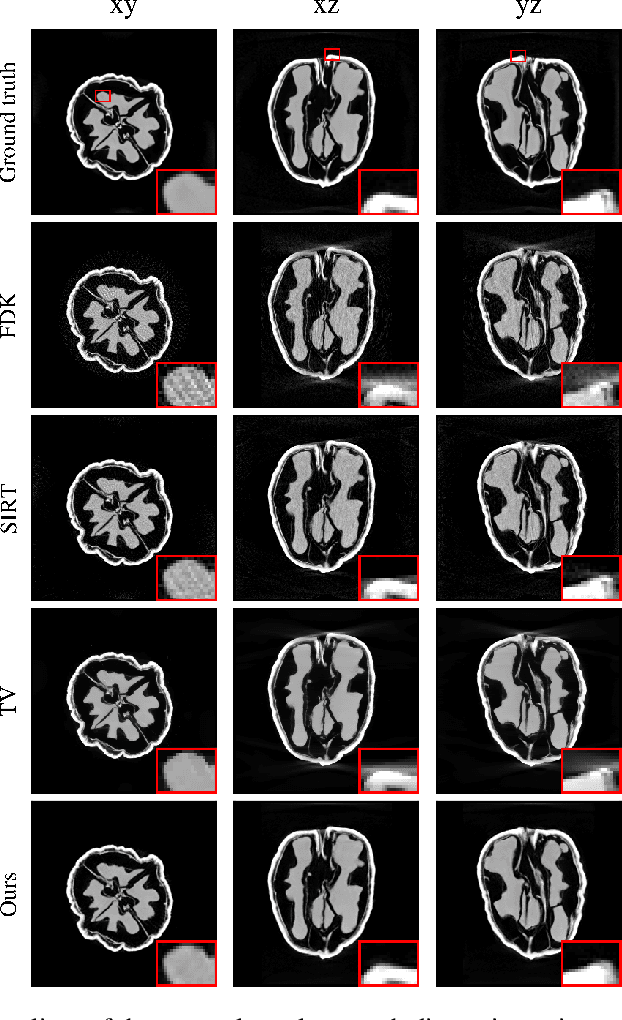
Abstract:Learned iterative reconstruction algorithms for inverse problems offer the flexibility to combine analytical knowledge about the problem with modules learned from data. This way, they achieve high reconstruction performance while ensuring consistency with the measured data. In computed tomography, extending such approaches from 2D fan-beam to 3D cone-beam data is challenging due to the prohibitively high GPU memory that would be needed to train such models. This paper proposes to use neural ordinary differential equations to solve the reconstruction problem in a residual formulation via numerical integration. For training, there is no need to backpropagate through several unrolled network blocks nor through the internals of the solver. Instead, the gradients are obtained very memory-efficiently in the neural ODE setting allowing for training on a single consumer graphics card. The method is able to reduce the root mean squared error by over 30% compared to the best performing classical iterative reconstruction algorithm and produces high quality cone-beam reconstructions even in a sparse view scenario.
Segmentation of the Carotid Lumen and Vessel Wall using Deep Learning and Location Priors
Jan 17, 2022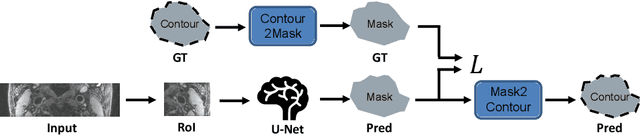
Abstract:In this report we want to present our method and results for the Carotid Artery Vessel Wall Segmentation Challenge. We propose an image-based pipeline utilizing the U-Net architecture and location priors to solve the segmentation problem at hand.
A Keypoint Detection and Description Network Based on the Vessel Structure for Multi-Modal Retinal Image Registration
Jan 06, 2022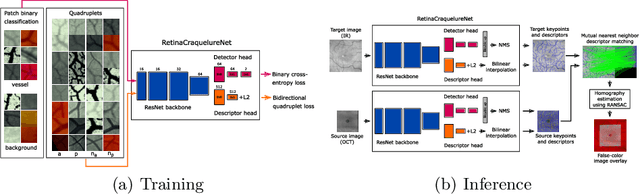
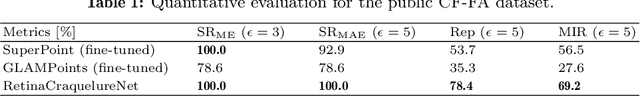

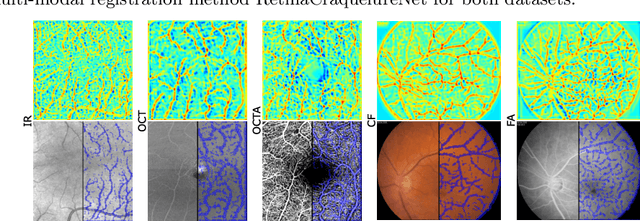

Abstract:Ophthalmological imaging utilizes different imaging systems, such as color fundus, infrared, fluorescein angiography, optical coherence tomography (OCT) or OCT angiography. Multiple images with different modalities or acquisition times are often analyzed for the diagnosis of retinal diseases. Automatically aligning the vessel structures in the images by means of multi-modal registration can support the ophthalmologists in their work. Our method uses a convolutional neural network to extract features of the vessel structure in multi-modal retinal images. We jointly train a keypoint detection and description network on small patches using a classification and a cross-modal descriptor loss function and apply the network to the full image size in the test phase. Our method demonstrates the best registration performance on our and a public multi-modal dataset in comparison to competing methods.
Deep learning for brain metastasis detection and segmentation in longitudinal MRI data
Dec 28, 2021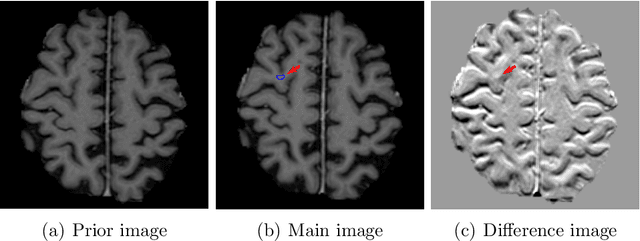
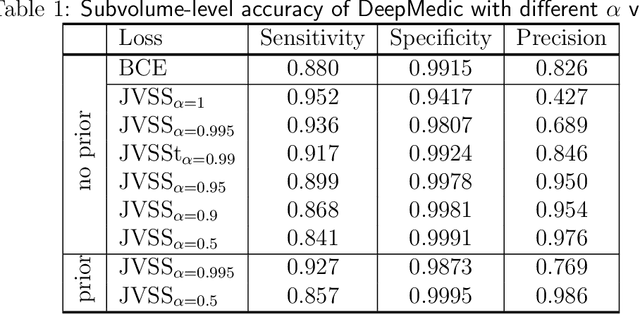
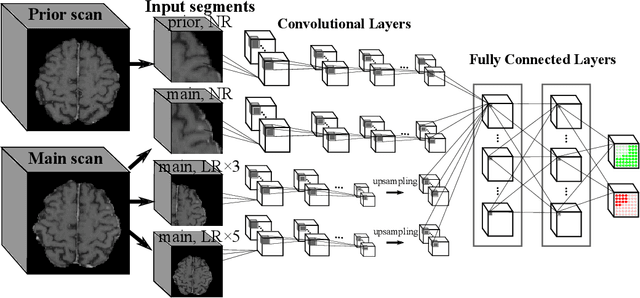
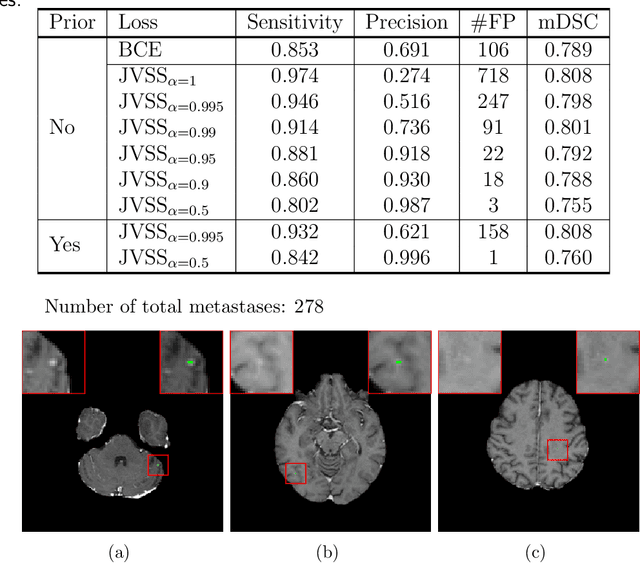
Abstract:Brain metastases occur frequently in patients with metastatic cancer. Early and accurate detection of brain metastases is very essential for treatment planning and prognosis in radiation therapy. To improve brain metastasis detection performance with deep learning, a custom detection loss called volume-level sensitivity-specificity (VSS) is proposed, which rates individual metastasis detection sensitivity and specificity in (sub-)volume levels. As sensitivity and precision are always a trade-off in a metastasis level, either a high sensitivity or a high precision can be achieved by adjusting the weights in the VSS loss without decline in dice score coefficient for segmented metastases. To reduce metastasis-like structures being detected as false positive metastases, a temporal prior volume is proposed as an additional input of the neural network. Our proposed VSS loss improves the sensitivity of brain metastasis detection, increasing the sensitivity from 86.7% to 95.5%. Alternatively, it improves the precision from 68.8% to 97.8%. With the additional temporal prior volume, about 45% of the false positive metastases are reduced in the high sensitivity model and the precision reaches 99.6% for the high specificity model. The mean dice coefficient for all metastases is about 0.81. With the ensemble of the high sensitivity and high specificity models, on average only 1.5 false positive metastases per patient needs further check, while the majority of true positive metastases are confirmed. The ensemble learning is able to distinguish high confidence true positive metastases from metastases candidates that require special expert review or further follow-up, being particularly well-fit to the requirements of expert support in real clinical practice.
Detection of Large Vessel Occlusions using Deep Learning by Deforming Vessel Tree Segmentations
Dec 10, 2021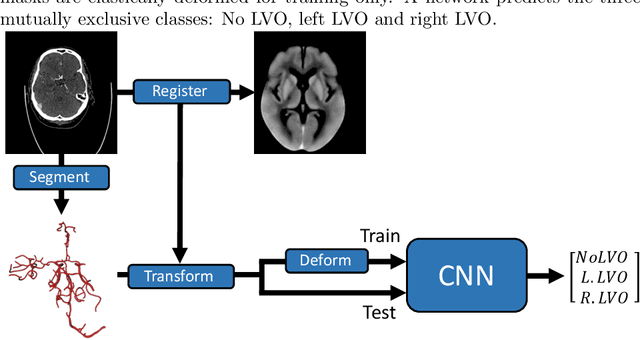
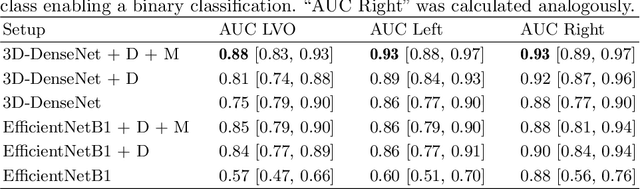
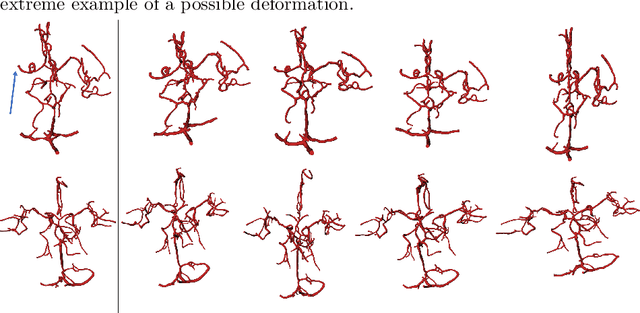
Abstract:Computed Tomography Angiography is a key modality providing insights into the cerebrovascular vessel tree that are crucial for the diagnosis and treatment of ischemic strokes, in particular in cases of large vessel occlusions (LVO). Thus, the clinical workflow greatly benefits from an automated detection of patients suffering from LVOs. This work uses convolutional neural networks for case-level classification trained with elastic deformation of the vessel tree segmentation masks to artificially augment training data. Using only masks as the input to our model uniquely allows us to apply such deformations much more aggressively than one could with conventional image volumes while retaining sample realism. The neural network classifies the presence of an LVO and the affected hemisphere. In a 5-fold cross validated ablation study, we demonstrate that the use of the suggested augmentation enables us to train robust models even from few data sets. Training the EfficientNetB1 architecture on 100 data sets, the proposed augmentation scheme was able to raise the ROC AUC to 0.85 from a baseline value of 0.56 using no augmentation. The best performance was achieved using a 3D-DenseNet yielding an AUC of 0.87. The augmentation had positive impact in classification of the affected hemisphere as well, where the 3D-DenseNet reached an AUC of 0.93 on both sides.
Does Proprietary Software Still Offer Protection of Intellectual Property in the Age of Machine Learning? -- A Case Study using Dual Energy CT Data
Dec 06, 2021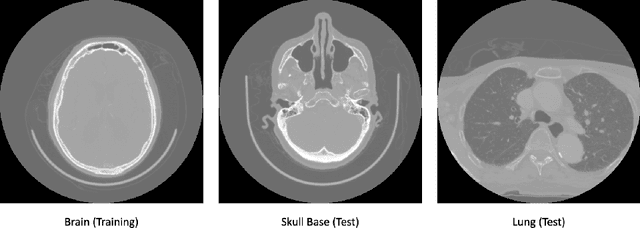
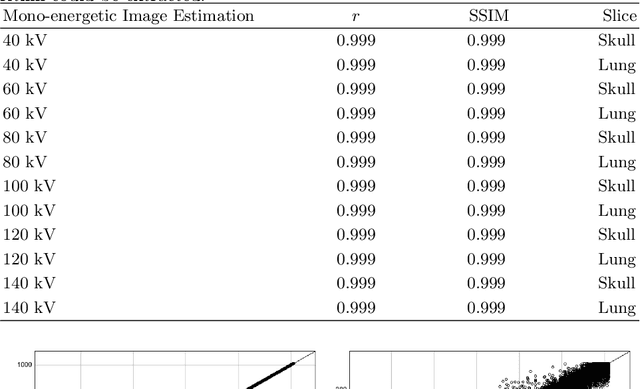
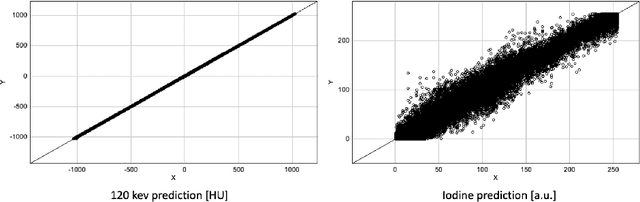
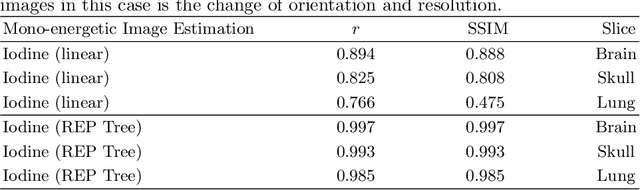
Abstract:In the domain of medical image processing, medical device manufacturers protect their intellectual property in many cases by shipping only compiled software, i.e. binary code which can be executed but is difficult to be understood by a potential attacker. In this paper, we investigate how well this procedure is able to protect image processing algorithms. In particular, we investigate whether the computation of mono-energetic images and iodine maps from dual energy CT data can be reverse-engineered by machine learning methods. Our results indicate that both can be approximated using only one single slice image as training data at a very high accuracy with structural similarity greater than 0.98 in all investigated cases.
Anomaly Detection in IR Images of PV Modules using Supervised Contrastive Learning
Dec 06, 2021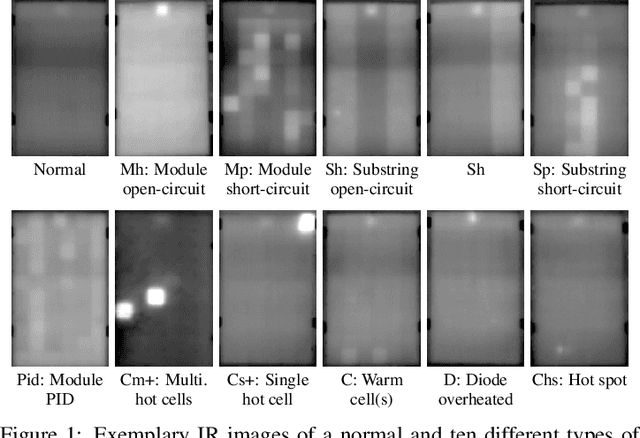
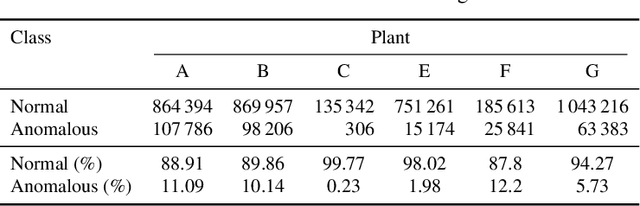
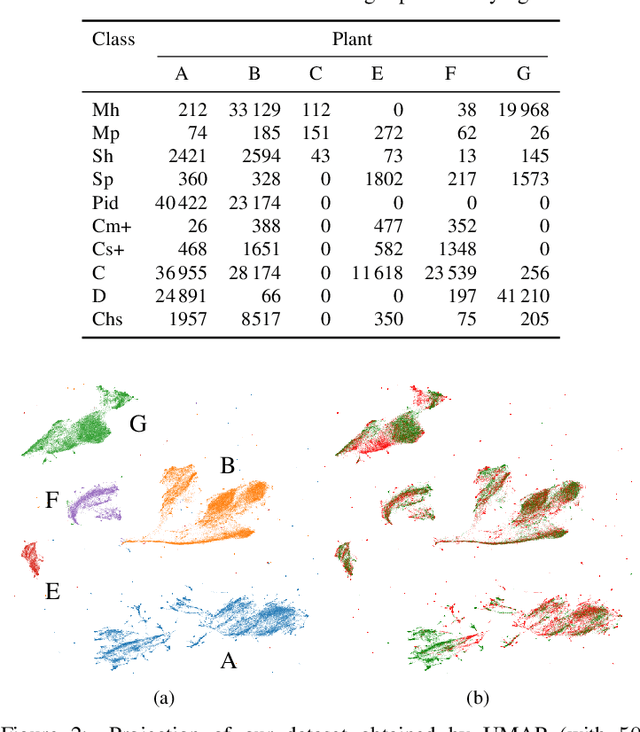

Abstract:Increasing deployment of photovoltaic (PV) plants requires methods for automatic detection of faulty PV modules in modalities, such as infrared (IR) images. Recently, deep learning has become popular for this. However, related works typically sample train and test data from the same distribution ignoring the presence of domain shift between data of different PV plants. Instead, we frame fault detection as more realistic unsupervised domain adaptation problem where we train on labelled data of one source PV plant and make predictions on another target plant. We train a ResNet-34 convolutional neural network with a supervised contrastive loss, on top of which we employ a k-nearest neighbor classifier to detect anomalies. Our method achieves a satisfactory area under the receiver operating characteristic (AUROC) of 73.3 % to 96.6 % on nine combinations of four source and target datasets with 2.92 million IR images of which 8.5 % are anomalous. It even outperforms a binary cross-entropy classifier in some cases. With a fixed decision threshold this results in 79.4 % and 77.1 % correctly classified normal and anomalous images, respectively. Most misclassified anomalies are of low severity, such as hot diodes and small hot spots. Our method is insensitive to hyperparameter settings, converges quickly and reliably detects unknown types of anomalies making it well suited for practice. Possible uses are in automatic PV plant inspection systems or to streamline manual labelling of IR datasets by filtering out normal images. Furthermore, our work serves the community with a more realistic view on PV module fault detection using unsupervised domain adaptation to develop more performant methods with favorable generalization capabilities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge