Amir Pouran Ben Veyseh
Trankit: A Light-Weight Transformer-based Toolkit for Multilingual Natural Language Processing
Jan 14, 2021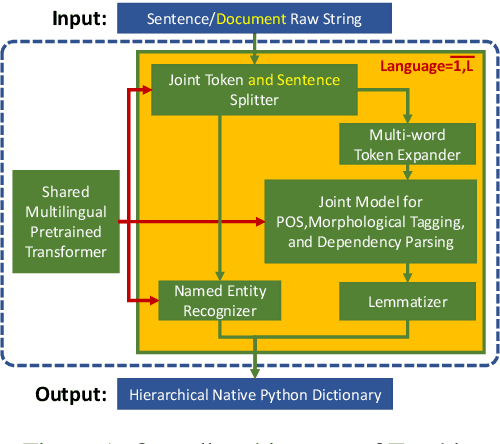
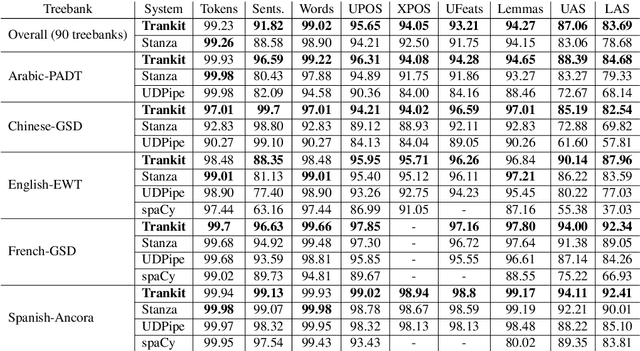
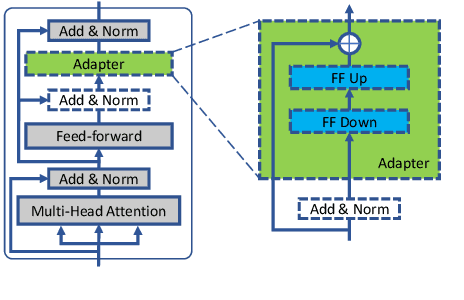
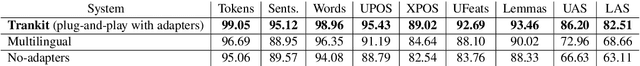
Abstract:We introduce Trankit, a light-weight Transformer-based Toolkit for multilingual Natural Language Processing (NLP). It provides a trainable pipeline for fundamental NLP tasks over 100 languages, and 90 pretrained pipelines for 56 languages. Built on a state-of-the-art pretrained language model, Trankit significantly outperforms prior multilingual NLP pipelines over sentence segmentation, part-of-speech tagging, morphological feature tagging, and dependency parsing while maintaining competitive performance for tokenization, multi-word token expansion, and lemmatization over 90 Universal Dependencies treebanks. Despite the use of a large pretrained transformer, our toolkit is still efficient in memory usage and speed. This is achieved by our novel plug-and-play mechanism with Adapters where a multilingual pretrained transformer is shared across pipelines for different languages. Our toolkit along with pretrained models and code are publicly available at: https://github.com/nlp-uoregon/trankit. A demo website for our toolkit is also available at: http://nlp.uoregon.edu/trankit. Finally, we create a demo video for Trankit at: https://youtu.be/q0KGP3zGjGc.
Acronym Identification and Disambiguation Shared Tasks for Scientific Document Understanding
Jan 06, 2021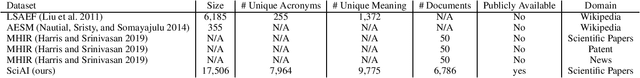

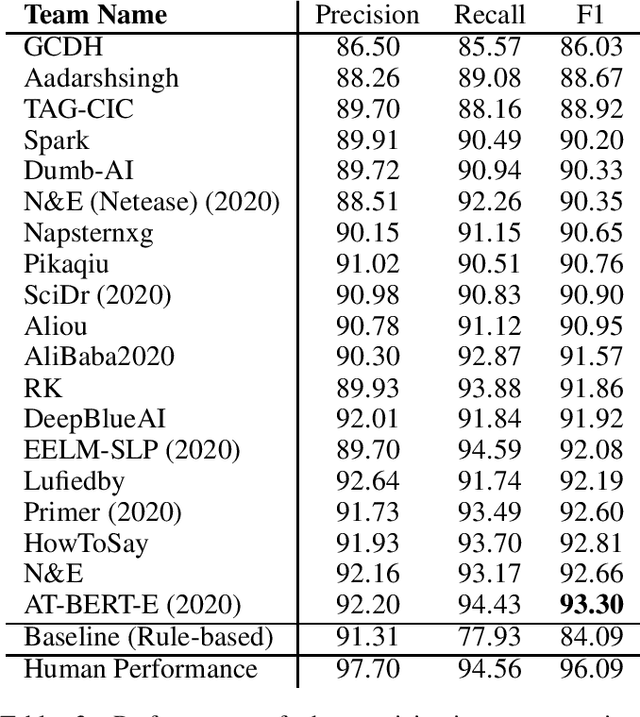
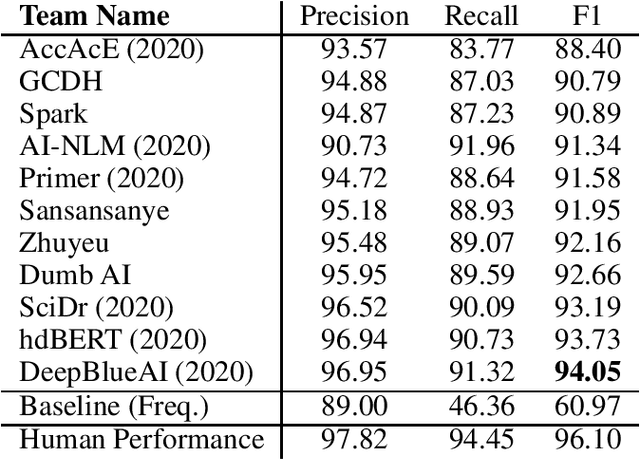
Abstract:Acronyms are the short forms of longer phrases and they are frequently used in writing, especially scholarly writing, to save space and facilitate the communication of information. As such, every text understanding tool should be capable of recognizing acronyms in text (i.e., acronym identification) and also finding their correct meaning (i.e., acronym disambiguation). As most of the prior works on these tasks are restricted to the biomedical domain and use unsupervised methods or models trained on limited datasets, they fail to perform well for scientific document understanding. To push forward research in this direction, we have organized two shared task for acronym identification and acronym disambiguation in scientific documents, named AI@SDU and AD@SDU, respectively. The two shared tasks have attracted 52 and 43 participants, respectively. While the submitted systems make substantial improvements compared to the existing baselines, there are still far from the human-level performance. This paper reviews the two shared tasks and the prominent participating systems for each of them.
What Does This Acronym Mean? Introducing a New Dataset for Acronym Identification and Disambiguation
Oct 28, 2020
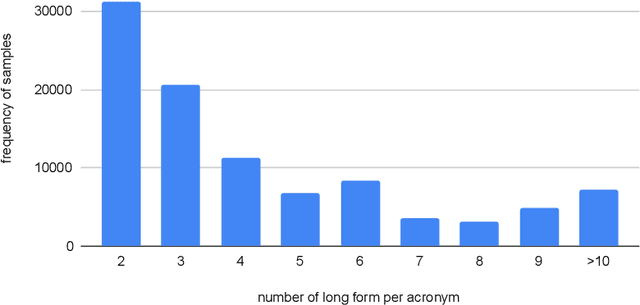
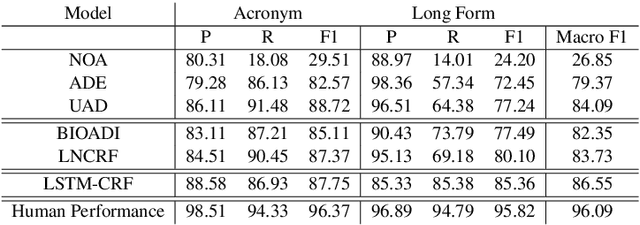
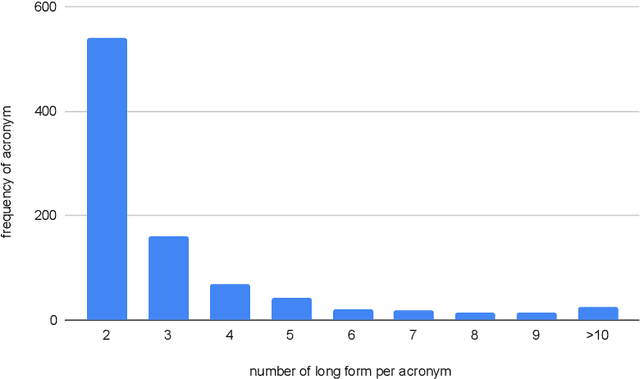
Abstract:Acronyms are the short forms of phrases that facilitate conveying lengthy sentences in documents and serve as one of the mainstays of writing. Due to their importance, identifying acronyms and corresponding phrases (i.e., acronym identification (AI)) and finding the correct meaning of each acronym (i.e., acronym disambiguation (AD)) are crucial for text understanding. Despite the recent progress on this task, there are some limitations in the existing datasets which hinder further improvement. More specifically, limited size of manually annotated AI datasets or noises in the automatically created acronym identification datasets obstruct designing advanced high-performing acronym identification models. Moreover, the existing datasets are mostly limited to the medical domain and ignore other domains. In order to address these two limitations, we first create a manually annotated large AI dataset for scientific domain. This dataset contains 17,506 sentences which is substantially larger than previous scientific AI datasets. Next, we prepare an AD dataset for scientific domain with 62,441 samples which is significantly larger than the previous scientific AD dataset. Our experiments show that the existing state-of-the-art models fall far behind human-level performance on both datasets proposed by this work. In addition, we propose a new deep learning model that utilizes the syntactical structure of the sentence to expand an ambiguous acronym in a sentence. The proposed model outperforms the state-of-the-art models on the new AD dataset, providing a strong baseline for future research on this dataset.
Graph Transformer Networks with Syntactic and Semantic Structures for Event Argument Extraction
Oct 26, 2020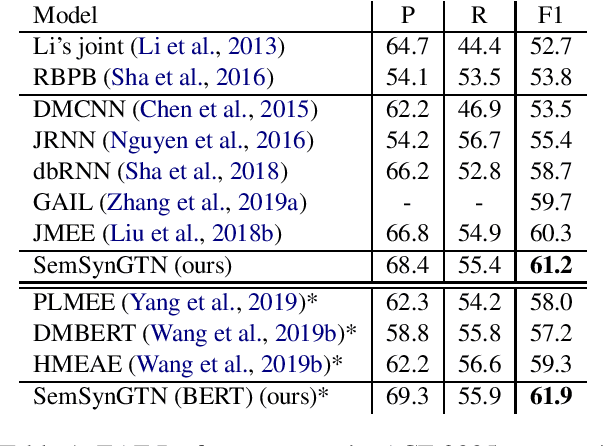
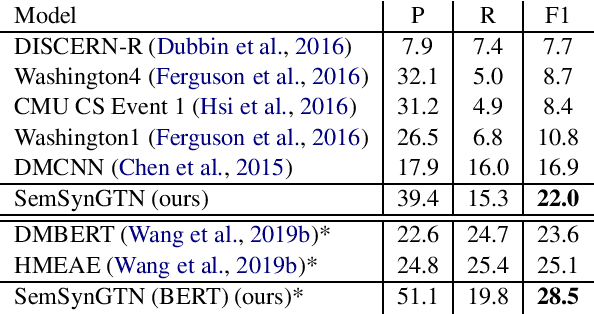
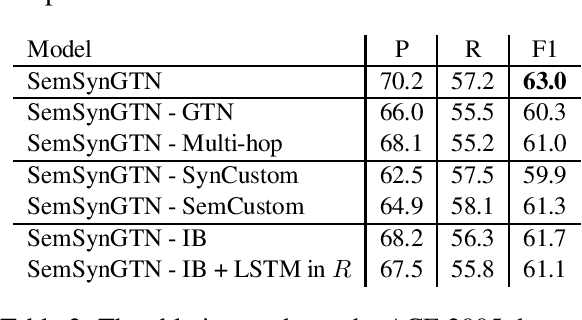
Abstract:The goal of Event Argument Extraction (EAE) is to find the role of each entity mention for a given event trigger word. It has been shown in the previous works that the syntactic structures of the sentences are helpful for the deep learning models for EAE. However, a major problem in such prior works is that they fail to exploit the semantic structures of the sentences to induce effective representations for EAE. Consequently, in this work, we propose a novel model for EAE that exploits both syntactic and semantic structures of the sentences with the Graph Transformer Networks (GTNs) to learn more effective sentence structures for EAE. In addition, we introduce a novel inductive bias based on information bottleneck to improve generalization of the EAE models. Extensive experiments are performed to demonstrate the benefits of the proposed model, leading to state-of-the-art performance for EAE on standard datasets.
Improving Aspect-based Sentiment Analysis with Gated Graph Convolutional Networks and Syntax-based Regulation
Oct 26, 2020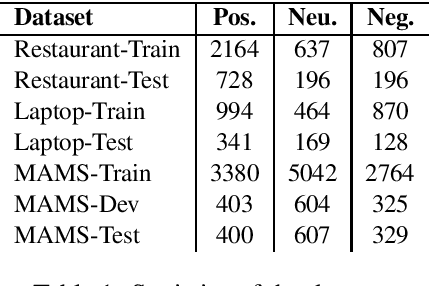
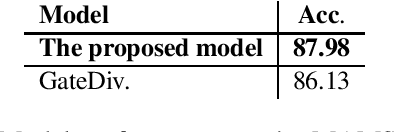
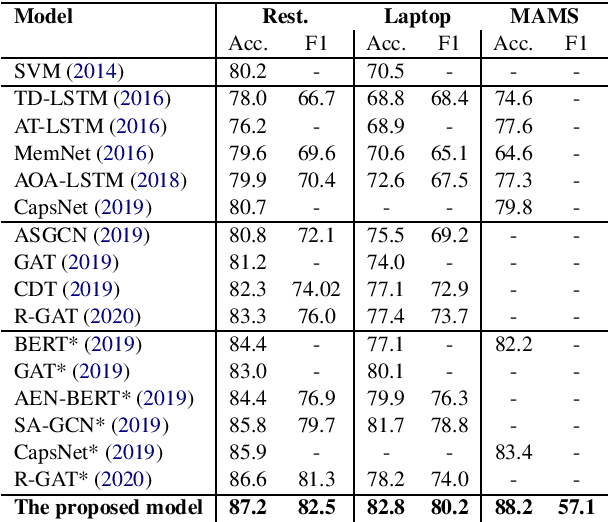
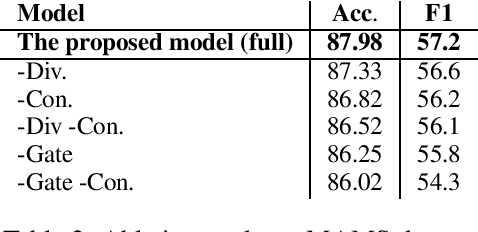

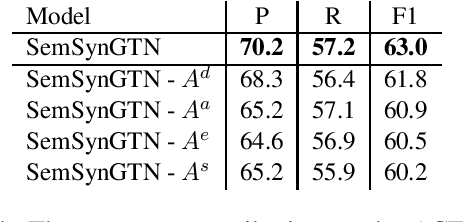
Abstract:Aspect-based Sentiment Analysis (ABSA) seeks to predict the sentiment polarity of a sentence toward a specific aspect. Recently, it has been shown that dependency trees can be integrated into deep learning models to produce the state-of-the-art performance for ABSA. However, these models tend to compute the hidden/representation vectors without considering the aspect terms and fail to benefit from the overall contextual importance scores of the words that can be obtained from the dependency tree for ABSA. In this work, we propose a novel graph-based deep learning model to overcome these two issues of the prior work on ABSA. In our model, gate vectors are generated from the representation vectors of the aspect terms to customize the hidden vectors of the graph-based models toward the aspect terms. In addition, we propose a mechanism to obtain the importance scores for each word in the sentences based on the dependency trees that are then injected into the model to improve the representation vectors for ABSA. The proposed model achieves the state-of-the-art performance on three benchmark datasets.
Introducing Syntactic Structures into Target Opinion Word Extraction with Deep Learning
Oct 26, 2020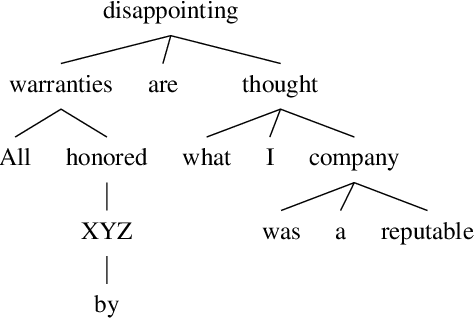
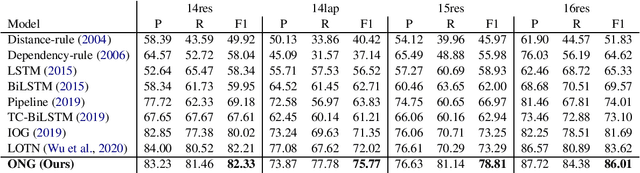
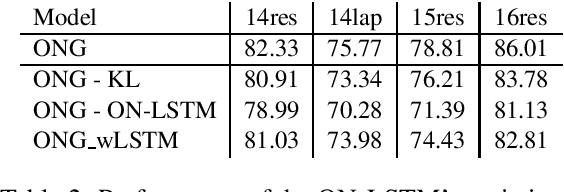
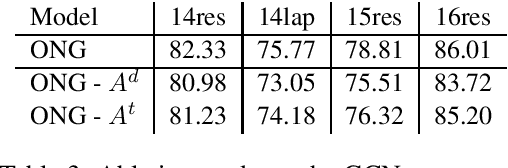
Abstract:Targeted opinion word extraction (TOWE) is a sub-task of aspect based sentiment analysis (ABSA) which aims to find the opinion words for a given aspect-term in a sentence. Despite their success for TOWE, the current deep learning models fail to exploit the syntactic information of the sentences that have been proved to be useful for TOWE in the prior research. In this work, we propose to incorporate the syntactic structures of the sentences into the deep learning models for TOWE, leveraging the syntax-based opinion possibility scores and the syntactic connections between the words. We also introduce a novel regularization technique to improve the performance of the deep learning models based on the representation distinctions between the words in TOWE. The proposed model is extensively analyzed and achieves the state-of-the-art performance on four benchmark datasets.
A Joint Model for Definition Extraction with Syntactic Connection and Semantic Consistency
Nov 17, 2019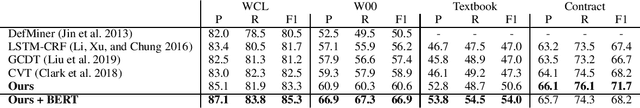
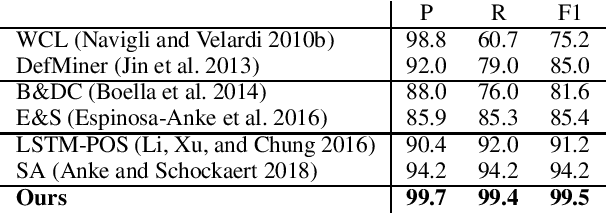
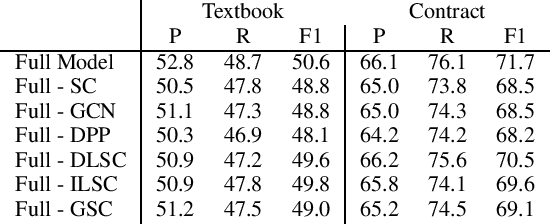
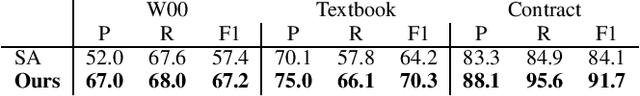
Abstract:Definition Extraction (DE) is one of the well-known topics in Information Extraction that aims to identify terms and their corresponding definitions in unstructured texts. This task can be formalized either as a sentence classification task (i.e., containing term-definition pairs or not) or a sequential labeling task (i.e., identifying the boundaries of the terms and definitions). The previous works for DE have only focused on one of the two approaches, failing to model the inter-dependencies between the two tasks. In this work, we propose a novel model for DE that simultaneously performs the two tasks in a single framework to benefit from their inter-dependencies. Our model features deep learning architectures to exploit the global structures of the input sentences as well as the semantic consistencies between the terms and the definitions, thereby improving the quality of the representation vectors for DE. Besides the joint inference between sentence classification and sequential labeling, the proposed model is fundamentally different from the prior work for DE in that the prior work has only employed the local structures of the input sentences (i.e., word-to-word relations), and not yet considered the semantic consistencies between terms and definitions. In order to implement these novel ideas, our model presents a multi-task learning framework that employs graph convolutional neural networks and predicts the dependency paths between the terms and the definitions. We also seek to enforce the consistency between the representations of the terms and definitions both globally (i.e., increasing semantic consistency between the representations of the entire sentences and the terms/definitions) and locally (i.e., promoting the similarity between the representations of the terms and the definitions).
Improving Slot Filling by Utilizing Contextual Information
Nov 05, 2019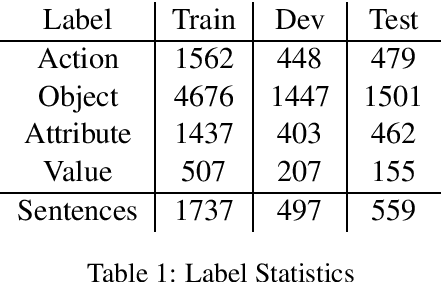
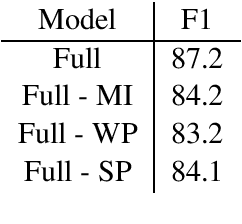
Abstract:Slot Filling is the task of extracting the semantic concept from a given natural language utterance. Recently it has been shown that using contextual information, either in work representations (e.g., BERT embedding) or in the computation graph of the model, could improve the performance of the model. However, recent work uses the contextual information in a restricted manner, e.g., by concatenating the word representation and its context feature vector, limiting the model from learning any direct association between the context and the label of word. We introduce a new deep model utilizing the contextual information for each work in the given sentence in a multi-task setting. Our model enforce consistency between the feature vectors of the context and the word while increasing the expressiveness of the context about the label of the word. Our empirical analysis on a slot filling dataset proves the superiority of the model over the baselines.
Improving Cross-Domain Performance for Relation Extraction via Dependency Prediction and Information Flow Control
Jul 07, 2019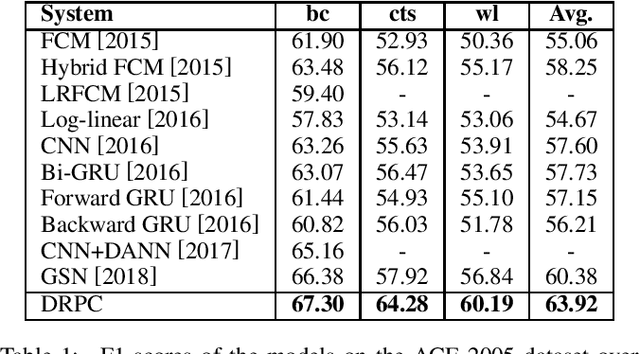
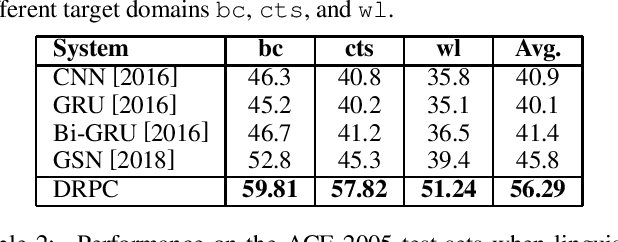
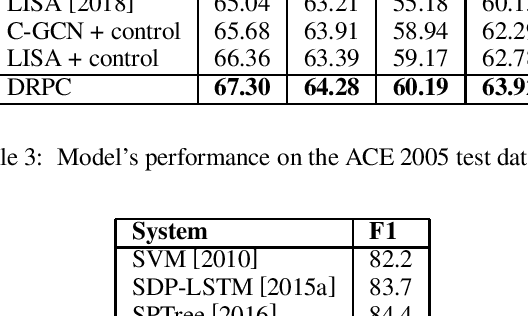
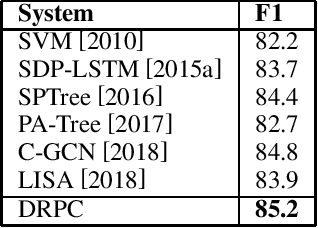
Abstract:Relation Extraction (RE) is one of the fundamental tasks in Information Extraction and Natural Language Processing. Dependency trees have been shown to be a very useful source of information for this task. The current deep learning models for relation extraction has mainly exploited this dependency information by guiding their computation along the structures of the dependency trees. One potential problem with this approach is it might prevent the models from capturing important context information beyond syntactic structures and cause the poor cross-domain generalization. This paper introduces a novel method to use dependency trees in RE for deep learning models that jointly predicts dependency and semantics relations. We also propose a new mechanism to control the information flow in the model based on the input entity mentions. Our extensive experiments on benchmark datasets show that the proposed model outperforms the existing methods for RE significantly.
Graph based Neural Networks for Event Factuality Prediction using Syntactic and Semantic Structures
Jul 07, 2019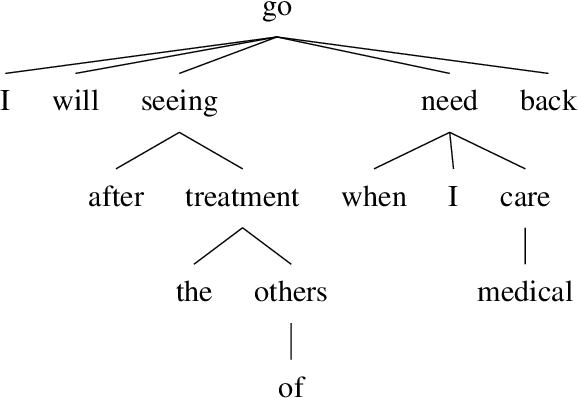
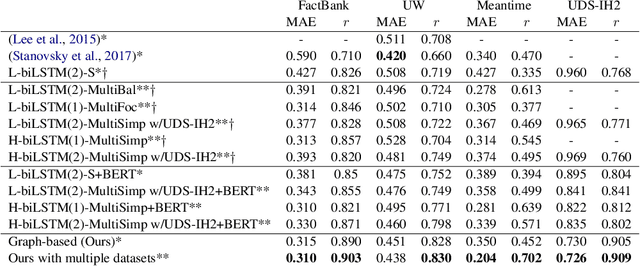
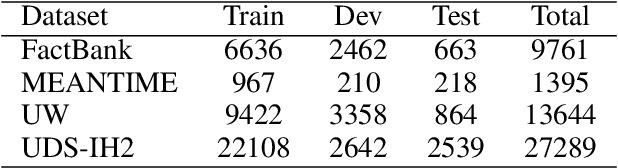
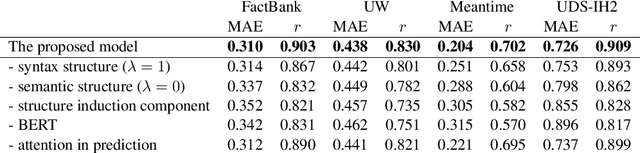
Abstract:Event factuality prediction (EFP) is the task of assessing the degree to which an event mentioned in a sentence has happened. For this task, both syntactic and semantic information are crucial to identify the important context words. The previous work for EFP has only combined these information in a simple way that cannot fully exploit their coordination. In this work, we introduce a novel graph-based neural network for EFP that can integrate the semantic and syntactic information more effectively. Our experiments demonstrate the advantage of the proposed model for EFP.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge