Adam P. Harrison
Accurate Weakly-Supervised Deep Lesion Segmentation using Large-Scale Clinical Annotations: Slice-Propagated 3D Mask Generation from 2D RECIST
Jul 02, 2018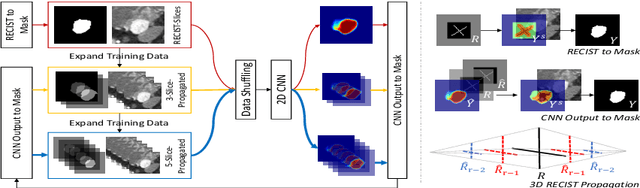
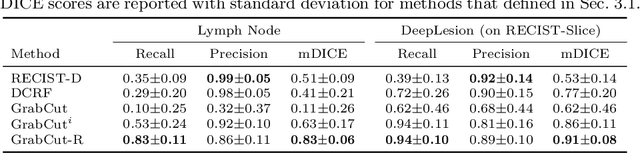
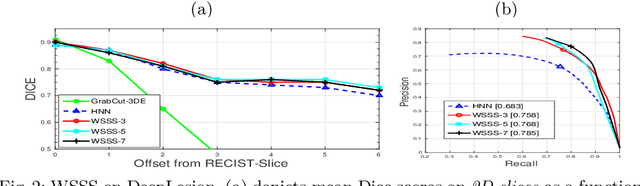
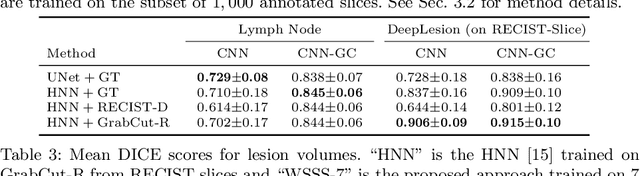
Abstract:Volumetric lesion segmentation from computed tomography (CT) images is a powerful means to precisely assess multiple time-point lesion/tumor changes. However, because manual 3D segmentation is prohibitively time consuming, current practices rely on an imprecise surrogate called response evaluation criteria in solid tumors (RECIST). Despite their coarseness, RECIST markers are commonly found in current hospital picture and archiving systems (PACS), meaning they can provide a potentially powerful, yet extraordinarily challenging, source of weak supervision for full 3D segmentation. Toward this end, we introduce a convolutional neural network (CNN) based weakly supervised slice-propagated segmentation (WSSS) method to 1) generate the initial lesion segmentation on the axial RECIST-slice; 2) learn the data distribution on RECIST-slices; 3) extrapolate to segment the whole lesion slice by slice to finally obtain a volumetric segmentation. To validate the proposed method, we first test its performance on a fully annotated lymph node dataset, where WSSS performs comparably to its fully supervised counterparts. We then test on a comprehensive lesion dataset with 32,735 RECIST marks, where we report a mean Dice score of 92% on RECIST-marked slices and 76% on the entire 3D volumes.
Semi-Automatic RECIST Labeling on CT Scans with Cascaded Convolutional Neural Networks
Jun 25, 2018
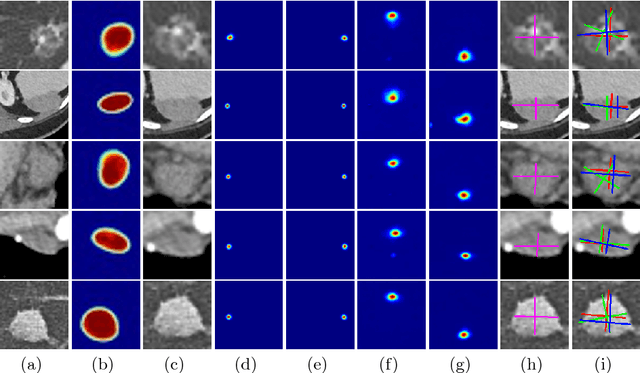
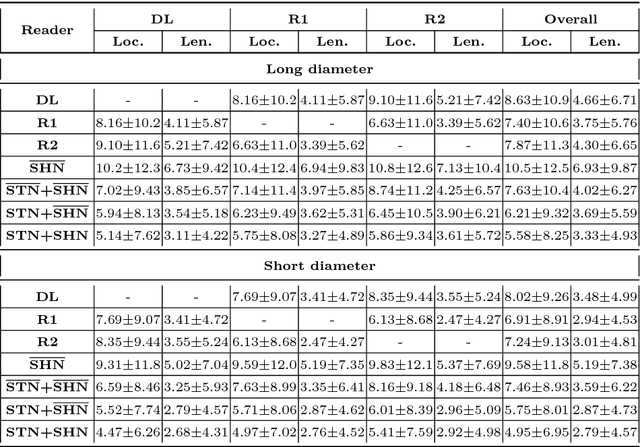
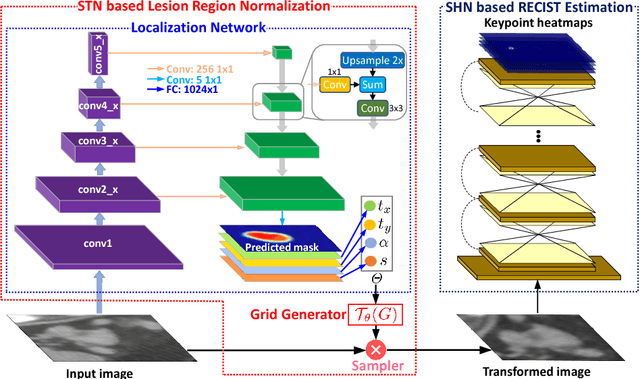

Abstract:Response evaluation criteria in solid tumors (RECIST) is the standard measurement for tumor extent to evaluate treatment responses in cancer patients. As such, RECIST annotations must be accurate. However, RECIST annotations manually labeled by radiologists require professional knowledge and are time-consuming, subjective, and prone to inconsistency among different observers. To alleviate these problems, we propose a cascaded convolutional neural network based method to semi-automatically label RECIST annotations and drastically reduce annotation time. The proposed method consists of two stages: lesion region normalization and RECIST estimation. We employ the spatial transformer network (STN) for lesion region normalization, where a localization network is designed to predict the lesion region and the transformation parameters with a multi-task learning strategy. For RECIST estimation, we adapt the stacked hourglass network (SHN), introducing a relationship constraint loss to improve the estimation precision. STN and SHN can both be learned in an end-to-end fashion. We train our system on the DeepLesion dataset, obtaining a consensus model trained on RECIST annotations performed by multiple radiologists over a multi-year period. Importantly, when judged against the inter-reader variability of two additional radiologist raters, our system performs more stably and with less variability, suggesting that RECIST annotations can be reliably obtained with reduced labor and time.
CT-Realistic Lung Nodule Simulation from 3D Conditional Generative Adversarial Networks for Robust Lung Segmentation
Jun 11, 2018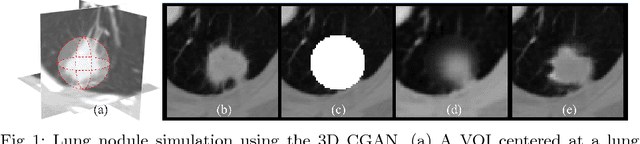
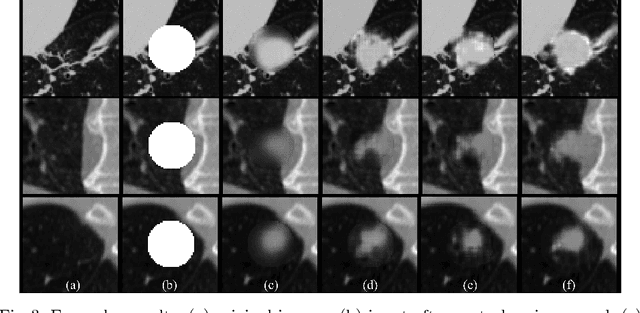
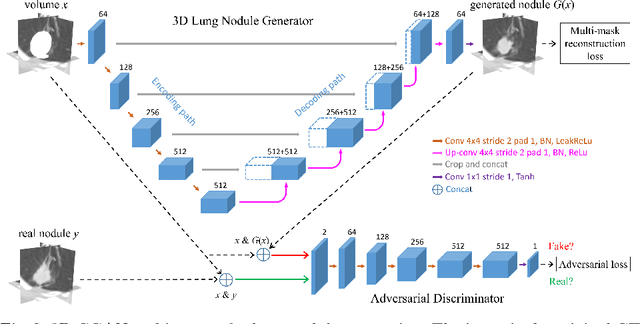

Abstract:Data availability plays a critical role for the performance of deep learning systems. This challenge is especially acute within the medical image domain, particularly when pathologies are involved, due to two factors: 1) limited number of cases, and 2) large variations in location, scale, and appearance. In this work, we investigate whether augmenting a dataset with artificially generated lung nodules can improve the robustness of the progressive holistically nested network (P-HNN) model for pathological lung segmentation of CT scans. To achieve this goal, we develop a 3D generative adversarial network (GAN) that effectively learns lung nodule property distributions in 3D space. In order to embed the nodules within their background context, we condition the GAN based on a volume of interest whose central part containing the nodule has been erased. To further improve realism and blending with the background, we propose a novel multi-mask reconstruction loss. We train our method on over 1000 nodules from the LIDC dataset. Qualitative results demonstrate the effectiveness of our method compared to the state-of-art. We then use our GAN to generate simulated training images where nodules lie on the lung border, which are cases where the published P-HNN model struggles. Qualitative and quantitative results demonstrate that armed with these simulated images, the P-HNN model learns to better segment lung regions under these challenging situations. As a result, our system provides a promising means to help overcome the data paucity that commonly afflicts medical imaging.
White matter hyperintensity segmentation from T1 and FLAIR images using fully convolutional neural networks enhanced with residual connections
Mar 19, 2018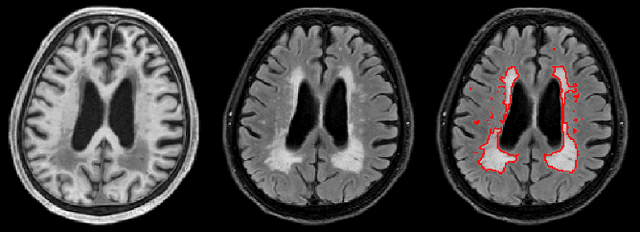
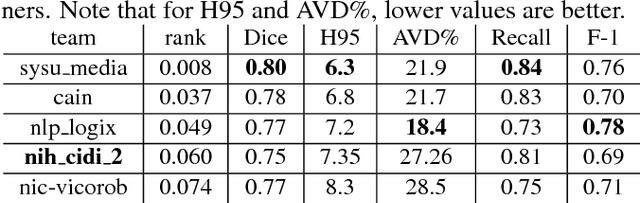
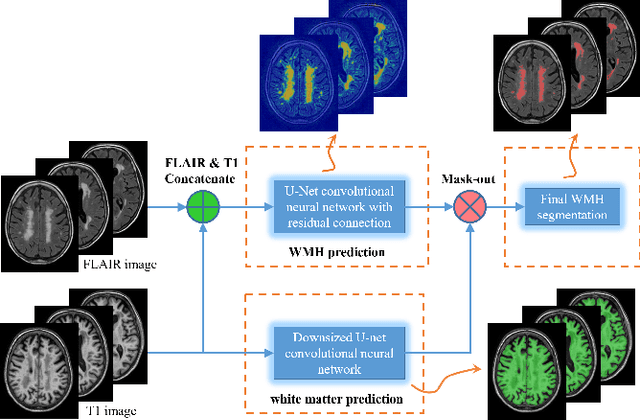
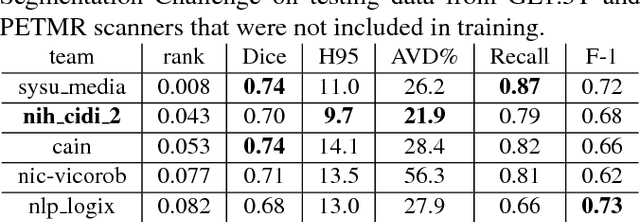
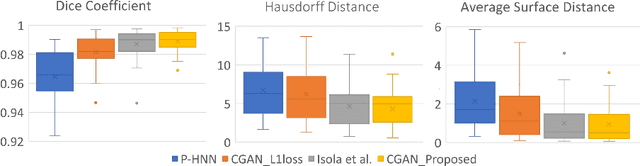
Abstract:Segmentation and quantification of white matter hyperintensities (WMHs) are of great importance in studying and understanding various neurological and geriatric disorders. Although automatic methods have been proposed for WMH segmentation on magnetic resonance imaging (MRI), manual corrections are often necessary to achieve clinically practical results. Major challenges for WMH segmentation stem from their inhomogeneous MRI intensities, random location and size distributions, and MRI noise. The presence of other brain anatomies or diseases with enhanced intensities adds further difficulties. To cope with these challenges, we present a specifically designed fully convolutional neural network (FCN) with residual connections to segment WMHs by using combined T1 and fluid-attenuated inversion recovery (FLAIR) images. Our customized FCN is designed to be straightforward and generalizable, providing efficient end-to-end training due to its enhanced information propagation. We tested our method on the open WMH Segmentation Challenge MICCAI2017 dataset, and, despite our method's relative simplicity, results show that it performs amongst the leading techniques across five metrics. More importantly, our method achieves the best score for hausdorff distance and average volume difference in testing datasets from two MRI scanners that were not included in training, demonstrating better generalization ability of our proposed method over its competitors.
Random Hinge Forest for Differentiable Learning
Mar 01, 2018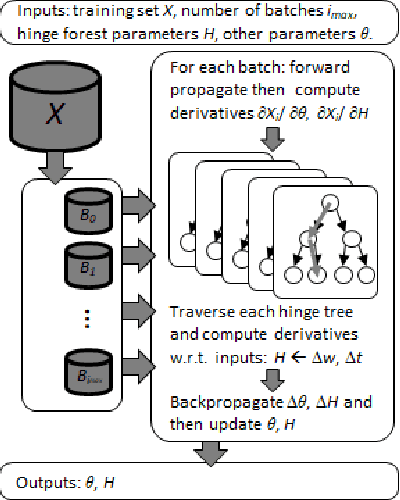
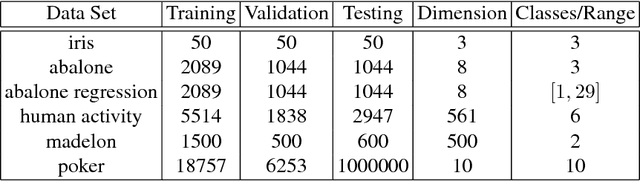
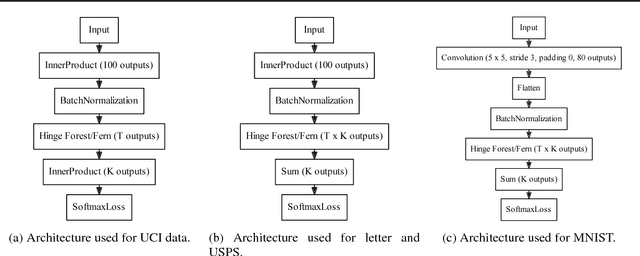
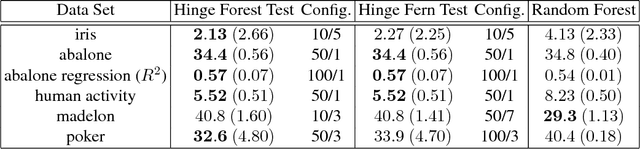
Abstract:We propose random hinge forests, a simple, efficient, and novel variant of decision forests. Importantly, random hinge forests can be readily incorporated as a general component within arbitrary computation graphs that are optimized end-to-end with stochastic gradient descent or variants thereof. We derive random hinge forest and ferns, focusing on their sparse and efficient nature, their min-max margin property, strategies to initialize them for arbitrary network architectures, and the class of optimizers most suitable for optimizing random hinge forest. The performance and versatility of random hinge forests are demonstrated by experiments incorporating a variety of of small and large UCI machine learning data sets and also ones involving the MNIST, Letter, and USPS image datasets. We compare random hinge forests with random forests and the more recent backpropagating deep neural decision forests.
Accurate Weakly Supervised Deep Lesion Segmentation on CT Scans: Self-Paced 3D Mask Generation from RECIST
Jan 25, 2018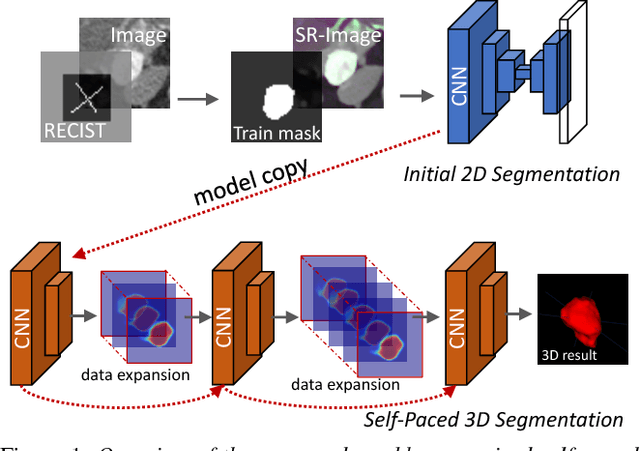
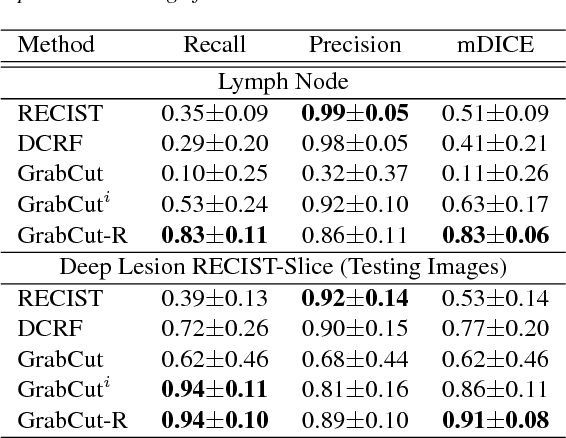
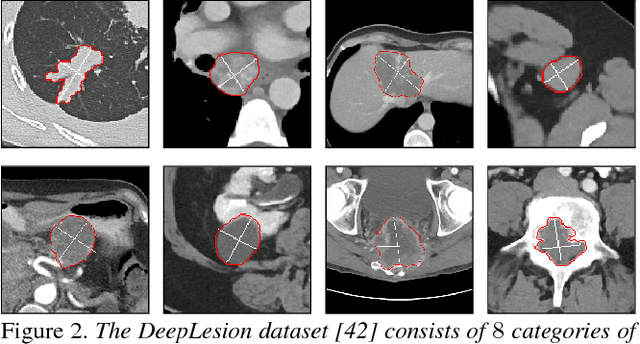
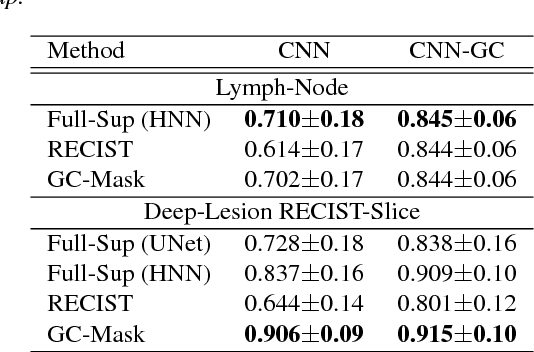
Abstract:Volumetric lesion segmentation via medical imaging is a powerful means to precisely assess multiple time-point lesion/tumor changes. Because manual 3D segmentation is prohibitively time consuming and requires radiological experience, current practices rely on an imprecise surrogate called response evaluation criteria in solid tumors (RECIST). Despite their coarseness, RECIST marks are commonly found in current hospital picture and archiving systems (PACS), meaning they can provide a potentially powerful, yet extraordinarily challenging, source of weak supervision for full 3D segmentation. Toward this end, we introduce a convolutional neural network based weakly supervised self-paced segmentation (WSSS) method to 1) generate the initial lesion segmentation on the axial RECIST-slice; 2) learn the data distribution on RECIST-slices; 3) adapt to segment the whole volume slice by slice to finally obtain a volumetric segmentation. In addition, we explore how super-resolution images (2~5 times beyond the physical CT imaging), generated from a proposed stacked generative adversarial network, can aid the WSSS performance. We employ the DeepLesion dataset, a comprehensive CT-image lesion dataset of 32,735 PACS-bookmarked findings, which include lesions, tumors, and lymph nodes of varying sizes, categories, body regions and surrounding contexts. These are drawn from 10,594 studies of 4,459 patients. We also validate on a lymph-node dataset, where 3D ground truth masks are available for all images. For the DeepLesion dataset, we report mean Dice coefficients of 93% on RECIST-slices and 76% in 3D lesion volumes. We further validate using a subjective user study, where an experienced radiologist accepted our WSSS-generated lesion segmentation results with a high probability of 92.4%.
Pathological Pulmonary Lobe Segmentation from CT Images using Progressive Holistically Nested Neural Networks and Random Walker
Aug 15, 2017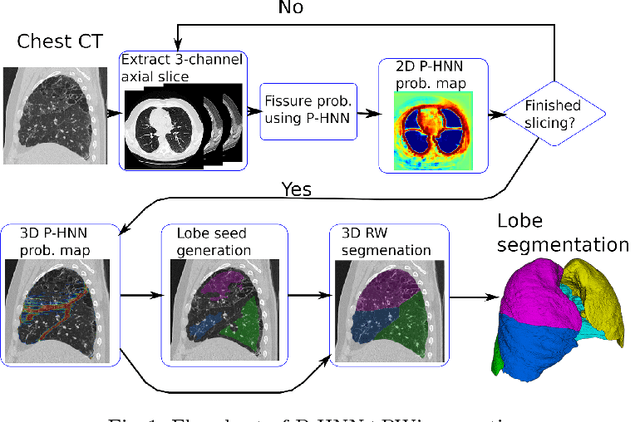
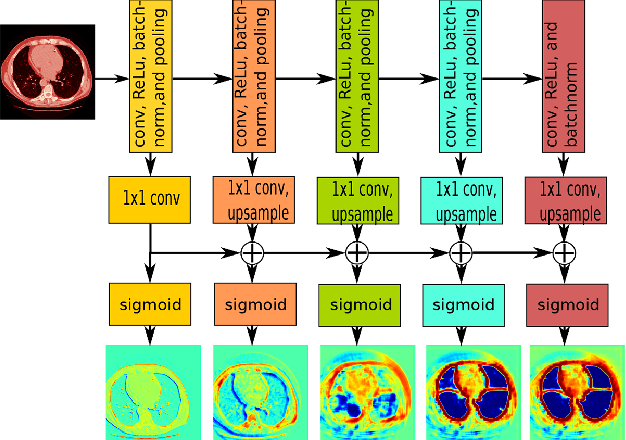
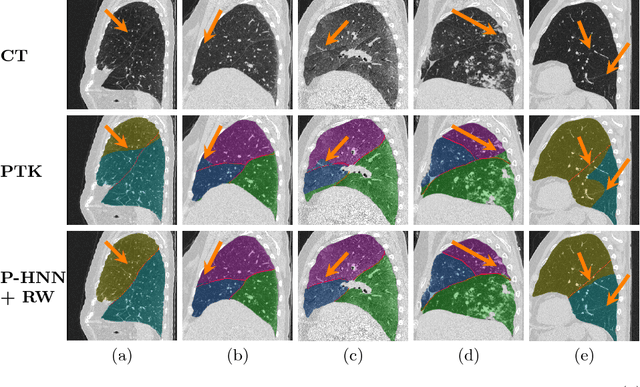
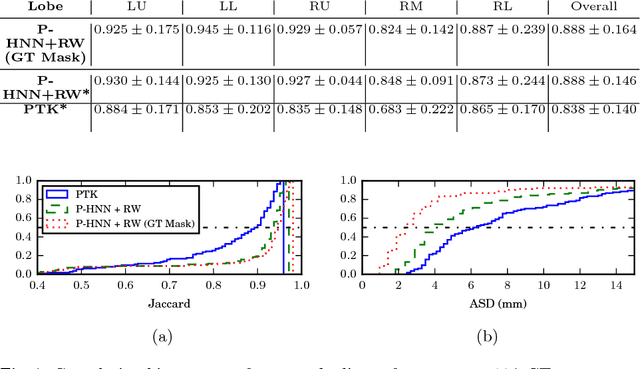
Abstract:Automatic pathological pulmonary lobe segmentation(PPLS) enables regional analyses of lung disease, a clinically important capability. Due to often incomplete lobe boundaries, PPLS is difficult even for experts, and most prior art requires inference from contextual information. To address this, we propose a novel PPLS method that couples deep learning with the random walker (RW) algorithm. We first employ the recent progressive holistically-nested network (P-HNN) model to identify potential lobar boundaries, then generate final segmentations using a RW that is seeded and weighted by the P-HNN output. We are the first to apply deep learning to PPLS. The advantages are independence from prior airway/vessel segmentations, increased robustness in diseased lungs, and methodological simplicity that does not sacrifice accuracy. Our method posts a high mean Jaccard score of 0.888$\pm$0.164 on a held-out set of 154 CT scans from lung-disease patients, while also significantly (p < 0.001) outperforming a state-of-the-art method.
Progressive and Multi-Path Holistically Nested Neural Networks for Pathological Lung Segmentation from CT Images
Jun 12, 2017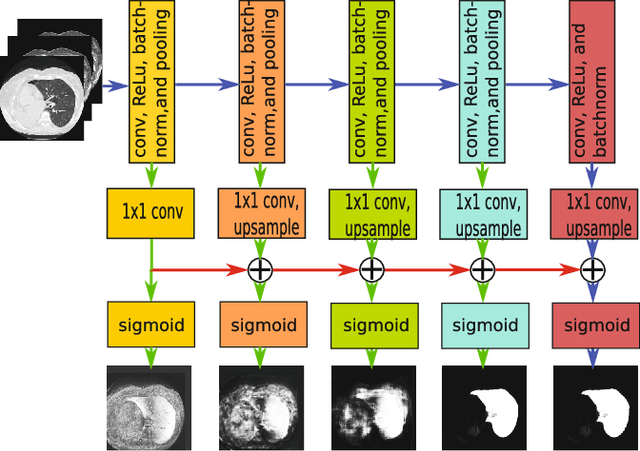

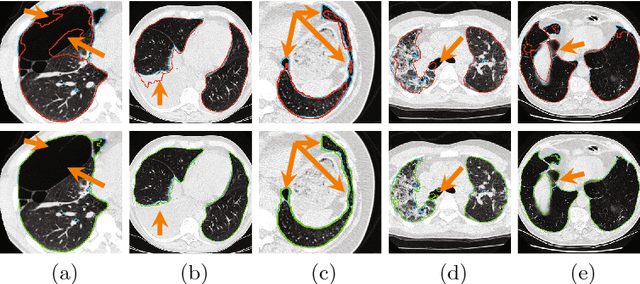
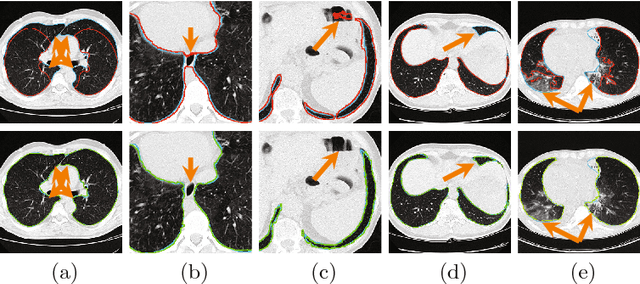
Abstract:Pathological lung segmentation (PLS) is an important, yet challenging, medical image application due to the wide variability of pathological lung appearance and shape. Because PLS is often a pre-requisite for other imaging analytics, methodological simplicity and generality are key factors in usability. Along those lines, we present a bottom-up deep-learning based approach that is expressive enough to handle variations in appearance, while remaining unaffected by any variations in shape. We incorporate the deeply supervised learning framework, but enhance it with a simple, yet effective, progressive multi-path scheme, which more reliably merges outputs from different network stages. The result is a deep model able to produce finer detailed masks, which we call progressive holistically-nested networks (P-HNNs). Using extensive cross-validation, our method is tested on multi-institutional datasets comprising 929 CT scans (848 publicly available), of pathological lungs, reporting mean dice scores of 0.985 and demonstrating significant qualitative and quantitative improvements over state-of-the art approaches.
* 8 Pages, 4 figures, MICCAI 2007
Spatial Aggregation of Holistically-Nested Convolutional Neural Networks for Automated Pancreas Localization and Segmentation
Jan 31, 2017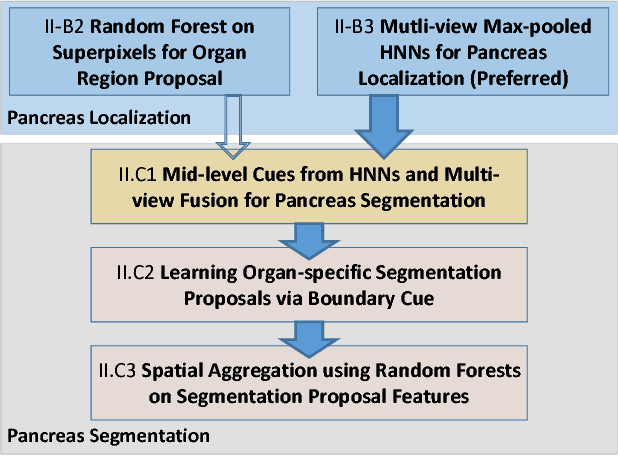
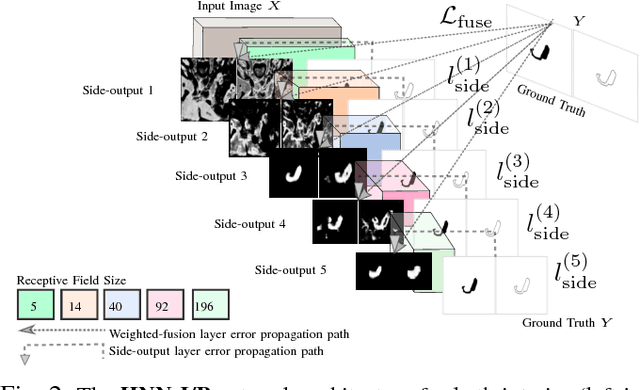
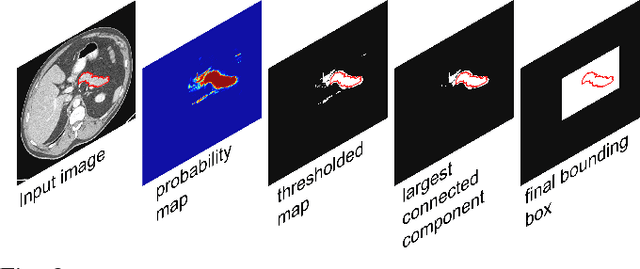
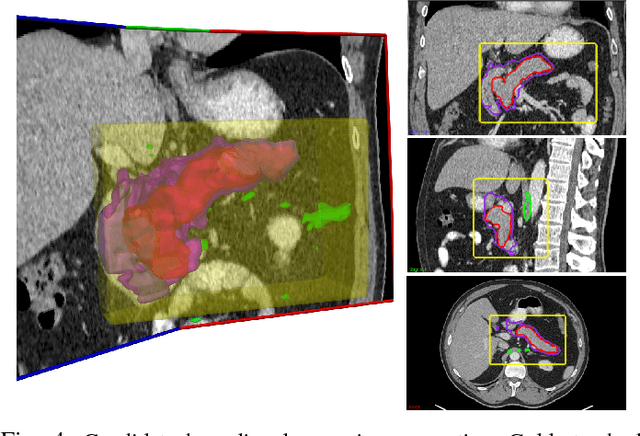
Abstract:Accurate and automatic organ segmentation from 3D radiological scans is an important yet challenging problem for medical image analysis. Specifically, the pancreas demonstrates very high inter-patient anatomical variability in both its shape and volume. In this paper, we present an automated system using 3D computed tomography (CT) volumes via a two-stage cascaded approach: pancreas localization and segmentation. For the first step, we localize the pancreas from the entire 3D CT scan, providing a reliable bounding box for the more refined segmentation step. We introduce a fully deep-learning approach, based on an efficient application of holistically-nested convolutional networks (HNNs) on the three orthogonal axial, sagittal, and coronal views. The resulting HNN per-pixel probability maps are then fused using pooling to reliably produce a 3D bounding box of the pancreas that maximizes the recall. We show that our introduced localizer compares favorably to both a conventional non-deep-learning method and a recent hybrid approach based on spatial aggregation of superpixels using random forest classification. The second, segmentation, phase operates within the computed bounding box and integrates semantic mid-level cues of deeply-learned organ interior and boundary maps, obtained by two additional and separate realizations of HNNs. By integrating these two mid-level cues, our method is capable of generating boundary-preserving pixel-wise class label maps that result in the final pancreas segmentation. Quantitative evaluation is performed on a publicly available dataset of 82 patient CT scans using 4-fold cross-validation (CV). We achieve a Dice similarity coefficient (DSC) of 81.27+/-6.27% in validation, which significantly outperforms previous state-of-the art methods that report DSCs of 71.80+/-10.70% and 78.01+/-8.20%, respectively, using the same dataset.
Holistic Interstitial Lung Disease Detection using Deep Convolutional Neural Networks: Multi-label Learning and Unordered Pooling
Jan 19, 2017
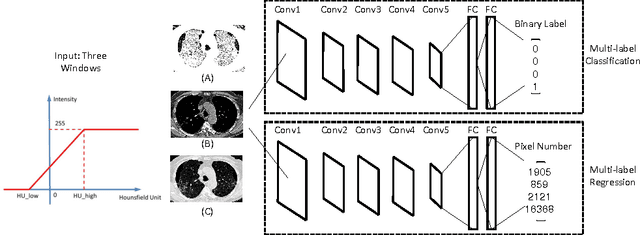
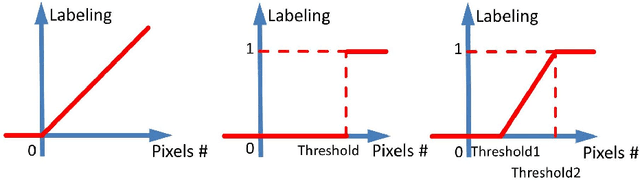
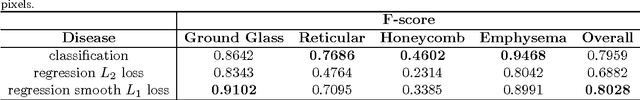
Abstract:Accurately predicting and detecting interstitial lung disease (ILD) patterns given any computed tomography (CT) slice without any pre-processing prerequisites, such as manually delineated regions of interest (ROIs), is a clinically desirable, yet challenging goal. The majority of existing work relies on manually-provided ILD ROIs to extract sampled 2D image patches from CT slices and, from there, performs patch-based ILD categorization. Acquiring manual ROIs is labor intensive and serves as a bottleneck towards fully-automated CT imaging ILD screening over large-scale populations. Furthermore, despite the considerable high frequency of more than one ILD pattern on a single CT slice, previous works are only designed to detect one ILD pattern per slice or patch. To tackle these two critical challenges, we present multi-label deep convolutional neural networks (CNNs) for detecting ILDs from holistic CT slices (instead of ROIs or sub-images). Conventional single-labeled CNN models can be augmented to cope with the possible presence of multiple ILD pattern labels, via 1) continuous-valued deep regression based robust norm loss functions or 2) a categorical objective as the sum of element-wise binary logistic losses. Our methods are evaluated and validated using a publicly available database of 658 patient CT scans under five-fold cross-validation, achieving promising performance on detecting four major ILD patterns: Ground Glass, Reticular, Honeycomb, and Emphysema. We also investigate the effectiveness of a CNN activation-based deep-feature encoding scheme using Fisher vector encoding, which treats ILD detection as spatially-unordered deep texture classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge