Abhijit Guha Roy
Concurrent Spatial and Channel Squeeze & Excitation in Fully Convolutional Networks
Jun 08, 2018
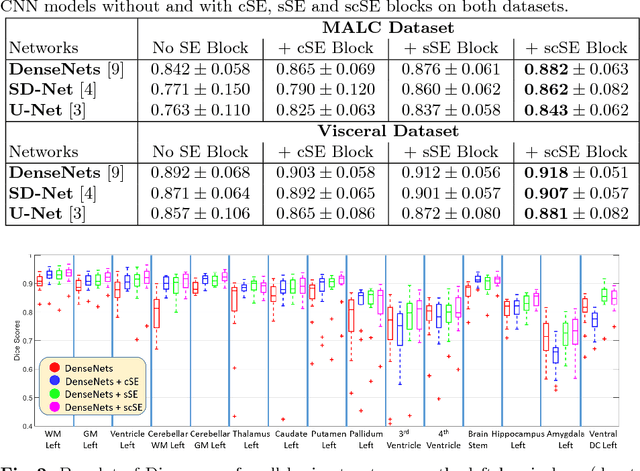
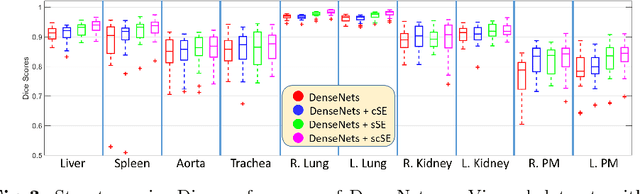
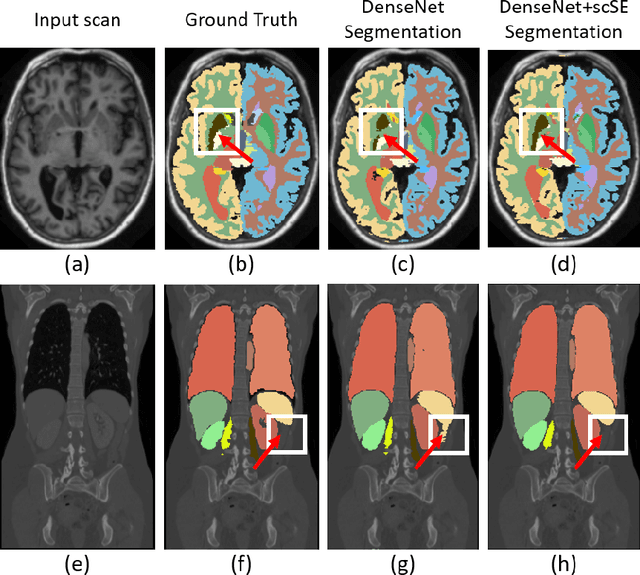
Abstract:Fully convolutional neural networks (F-CNNs) have set the state-of-the-art in image segmentation for a plethora of applications. Architectural innovations within F-CNNs have mainly focused on improving spatial encoding or network connectivity to aid gradient flow. In this paper, we explore an alternate direction of recalibrating the feature maps adaptively, to boost meaningful features, while suppressing weak ones. We draw inspiration from the recently proposed squeeze & excitation (SE) module for channel recalibration of feature maps for image classification. Towards this end, we introduce three variants of SE modules for image segmentation, (i) squeezing spatially and exciting channel-wise (cSE), (ii) squeezing channel-wise and exciting spatially (sSE) and (iii) concurrent spatial and channel squeeze & excitation (scSE). We effectively incorporate these SE modules within three different state-of-the-art F-CNNs (DenseNet, SD-Net, U-Net) and observe consistent improvement of performance across all architectures, while minimally effecting model complexity. Evaluations are performed on two challenging applications: whole brain segmentation on MRI scans (Multi-Atlas Labelling Challenge Dataset) and organ segmentation on whole body contrast enhanced CT scans (Visceral Dataset).
Inherent Brain Segmentation Quality Control from Fully ConvNet Monte Carlo Sampling
Jun 08, 2018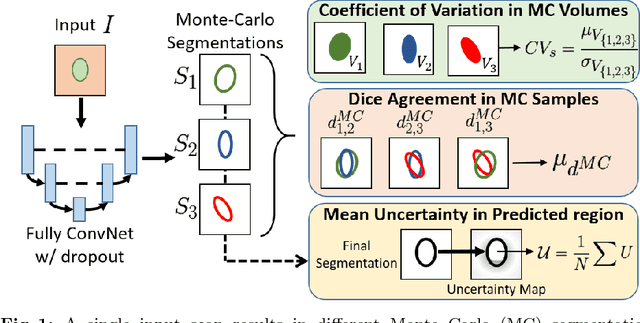
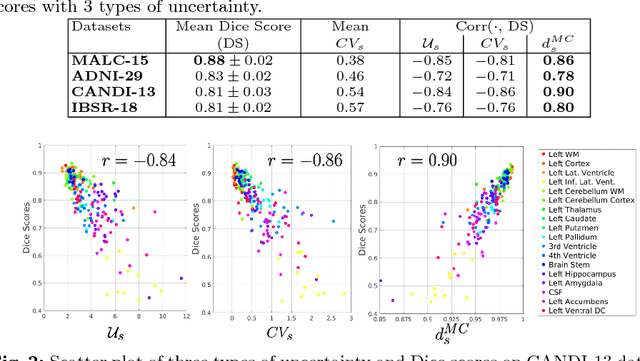
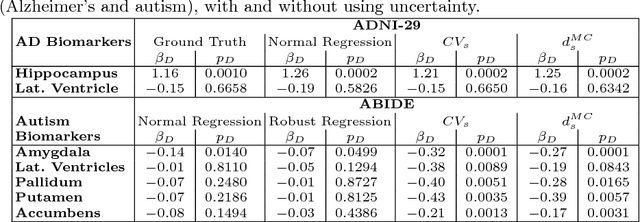
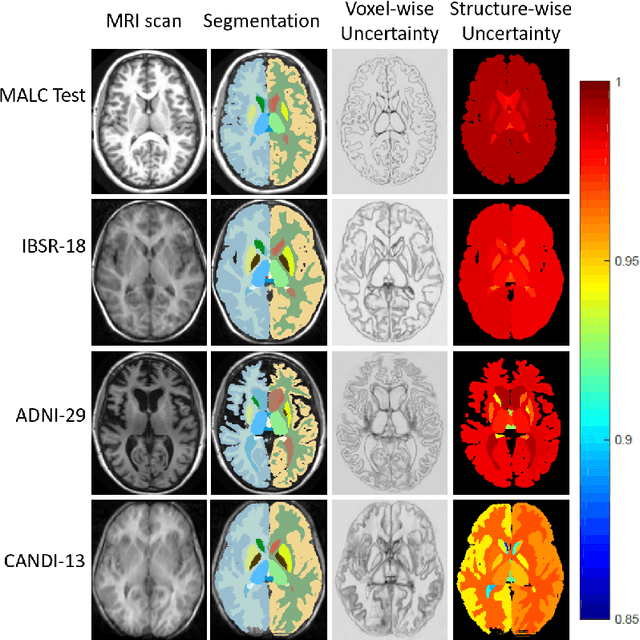
Abstract:We introduce inherent measures for effective quality control of brain segmentation based on a Bayesian fully convolutional neural network, using model uncertainty. Monte Carlo samples from the posterior distribution are efficiently generated using dropout at test time. Based on these samples, we introduce next to a voxel-wise uncertainty map also three metrics for structure-wise uncertainty. We then incorporate these structure-wise uncertainty in group analyses as a measure of confidence in the observation. Our results show that the metrics are highly correlated to segmentation accuracy and therefore present an inherent measure of segmentation quality. Furthermore, group analysis with uncertainty results in effect sizes closer to that of manual annotations. The introduced uncertainty metrics can not only be very useful in translation to clinical practice but also provide automated quality control and group analyses in processing large data repositories.
QuickNAT: Segmenting MRI Neuroanatomy in 20 seconds
Jan 12, 2018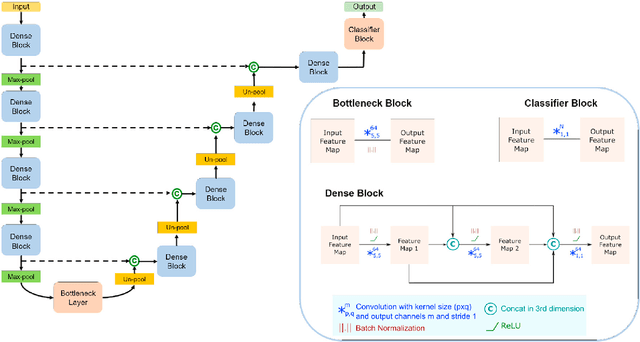

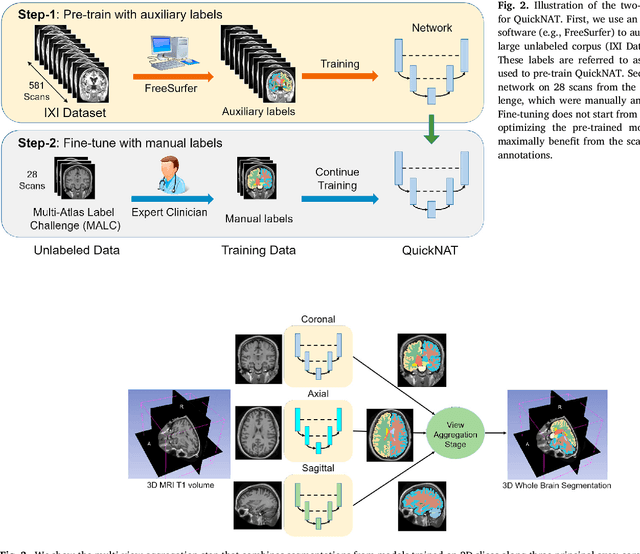
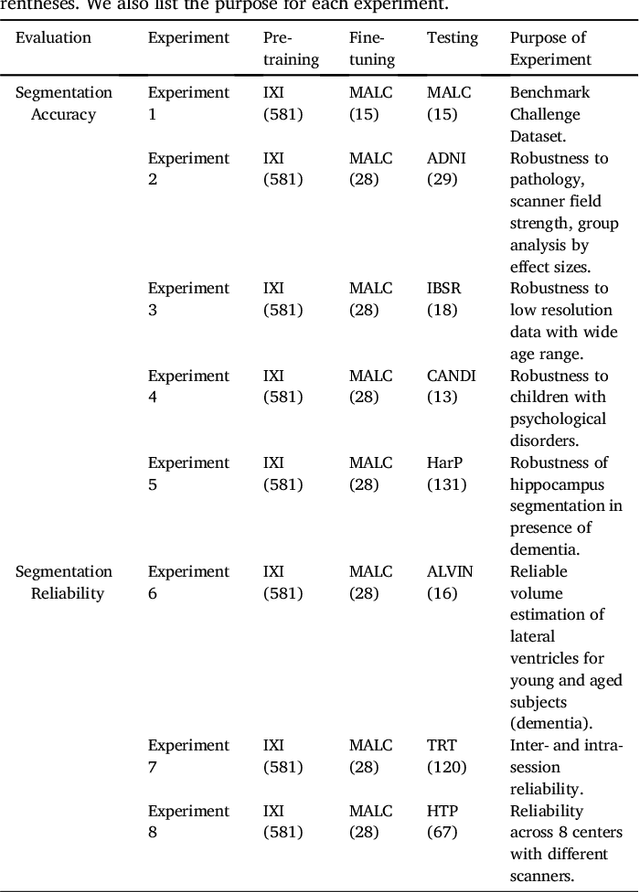
Abstract:Whole brain segmentation from structural magnetic resonance imaging is a prerequisite for most morphological analyses, but requires hours of processing time and therefore delays the availability of image markers after scan acquisition. We introduce QuickNAT, a fully convolution neural network that segments a brain scan in 20 seconds. To enable training of the complex network with limited annotated data, we propose to pre-train on auxiliary labels created from existing segmentation software and to subsequently fine-tune on manual labels. In an extensive set of evaluations on eight datasets that cover a wide age range, pathology, and different scanners, we demonstrate that QuickNAT achieves superior performance to state-of-the-art methods, while being about 700 times faster. This drastic speed up greatly facilitates the processing of large data repositories and supports the translation of imaging biomarkers by making them almost instantaneously available.
ReLayNet: Retinal Layer and Fluid Segmentation of Macular Optical Coherence Tomography using Fully Convolutional Network
Jul 07, 2017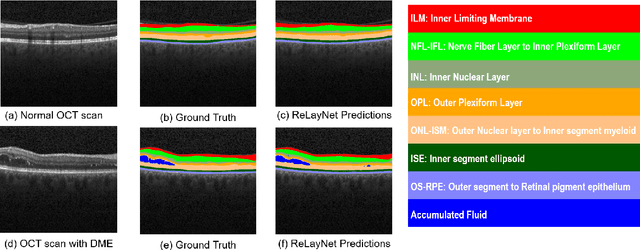
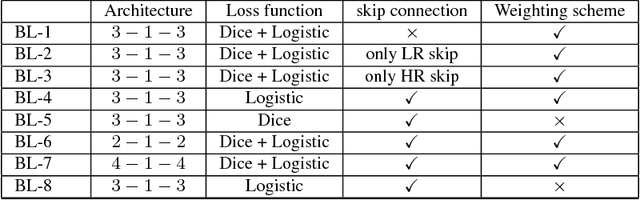

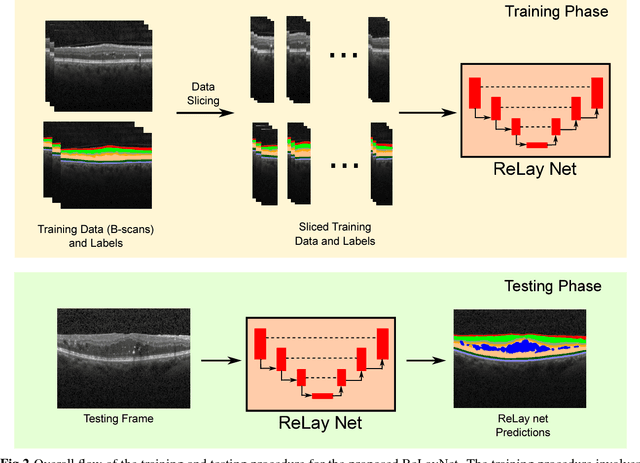
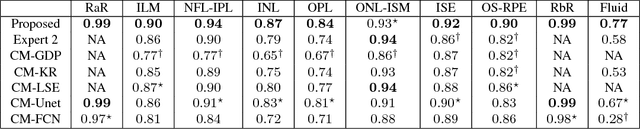
Abstract:Optical coherence tomography (OCT) is used for non-invasive diagnosis of diabetic macular edema assessing the retinal layers. In this paper, we propose a new fully convolutional deep architecture, termed ReLayNet, for end-to-end segmentation of retinal layers and fluid masses in eye OCT scans. ReLayNet uses a contracting path of convolutional blocks (encoders) to learn a hierarchy of contextual features, followed by an expansive path of convolutional blocks (decoders) for semantic segmentation. ReLayNet is trained to optimize a joint loss function comprising of weighted logistic regression and Dice overlap loss. The framework is validated on a publicly available benchmark dataset with comparisons against five state-of-the-art segmentation methods including two deep learning based approaches to substantiate its effectiveness.
Error Corrective Boosting for Learning Fully Convolutional Networks with Limited Data
Jul 02, 2017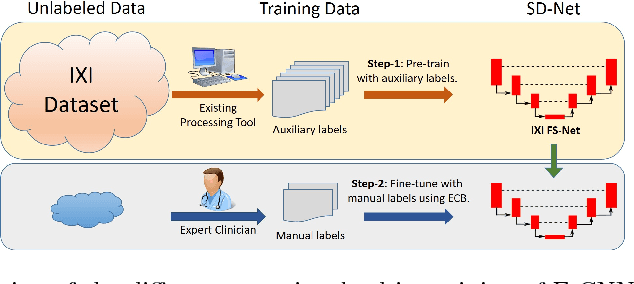
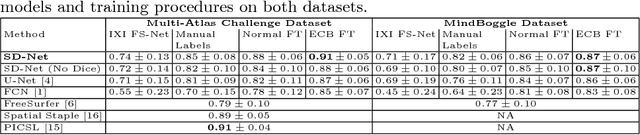
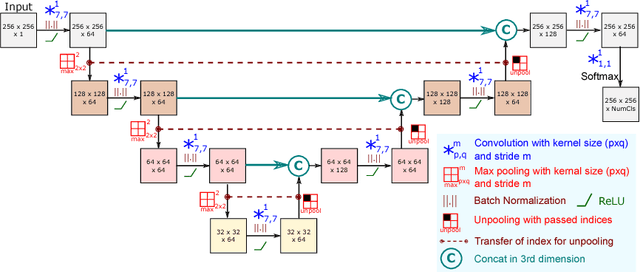
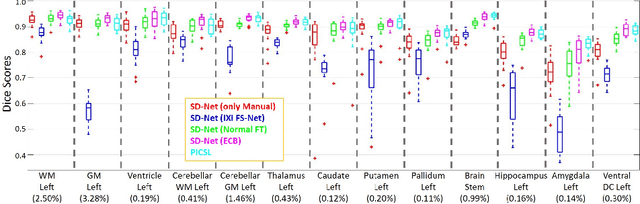
Abstract:Training deep fully convolutional neural networks (F-CNNs) for semantic image segmentation requires access to abundant labeled data. While large datasets of unlabeled image data are available in medical applications, access to manually labeled data is very limited. We propose to automatically create auxiliary labels on initially unlabeled data with existing tools and to use them for pre-training. For the subsequent fine-tuning of the network with manually labeled data, we introduce error corrective boosting (ECB), which emphasizes parameter updates on classes with lower accuracy. Furthermore, we introduce SkipDeconv-Net (SD-Net), a new F-CNN architecture for brain segmentation that combines skip connections with the unpooling strategy for upsampling. The SD-Net addresses challenges of severe class imbalance and errors along boundaries. With application to whole-brain MRI T1 scan segmentation, we generate auxiliary labels on a large dataset with FreeSurfer and fine-tune on two datasets with manual annotations. Our results show that the inclusion of auxiliary labels and ECB yields significant improvements. SD-Net segments a 3D scan in 7 secs in comparison to 30 hours for the closest multi-atlas segmentation method, while reaching similar performance. It also outperforms the latest state-of-the-art F-CNN models.
Deep Residual Hashing
Dec 16, 2016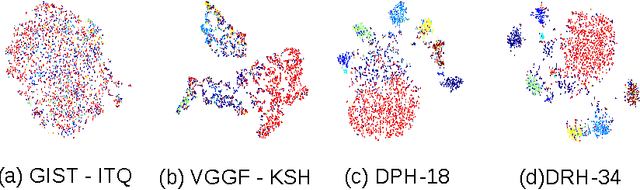
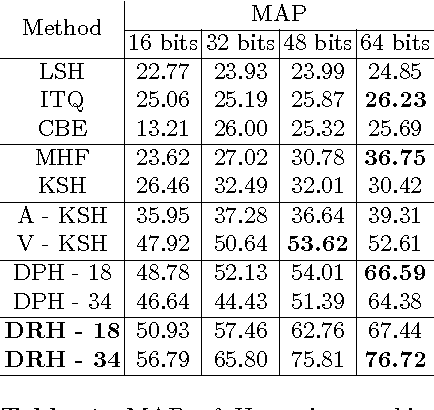
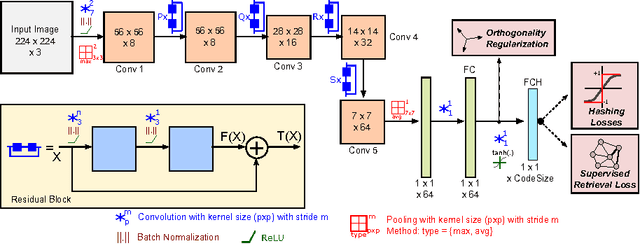
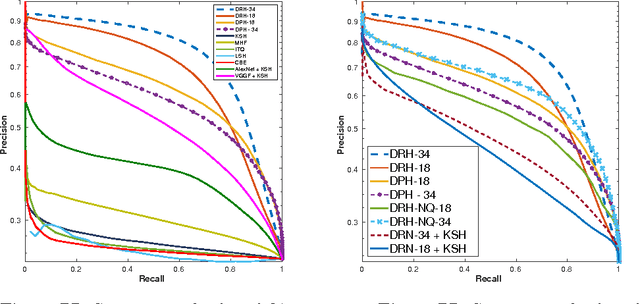
Abstract:Hashing aims at generating highly compact similarity preserving code words which are well suited for large-scale image retrieval tasks. Most existing hashing methods first encode the images as a vector of hand-crafted features followed by a separate binarization step to generate hash codes. This two-stage process may produce sub-optimal encoding. In this paper, for the first time, we propose a deep architecture for supervised hashing through residual learning, termed Deep Residual Hashing (DRH), for an end-to-end simultaneous representation learning and hash coding. The DRH model constitutes four key elements: (1) a sub-network with multiple stacked residual blocks; (2) hashing layer for binarization; (3) supervised retrieval loss function based on neighbourhood component analysis for similarity preserving embedding; and (4) hashing related losses and regularisation to control the quantization error and improve the quality of hash coding. We present results of extensive experiments on a large public chest x-ray image database with co-morbidities and discuss the outcome showing substantial improvements over the latest state-of-the art methods.
Deep Neural Ensemble for Retinal Vessel Segmentation in Fundus Images towards Achieving Label-free Angiography
Sep 19, 2016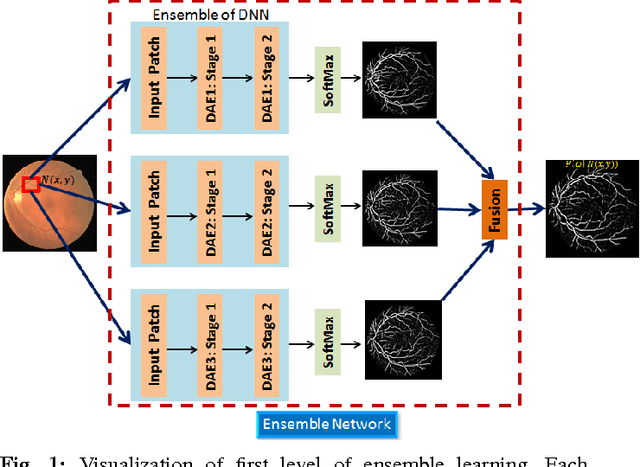
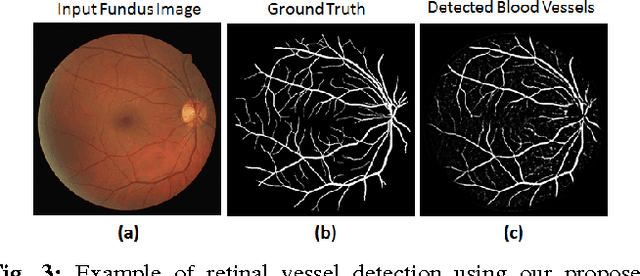
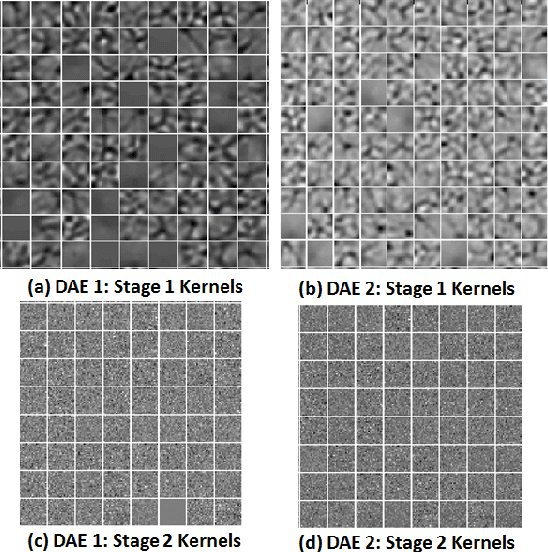
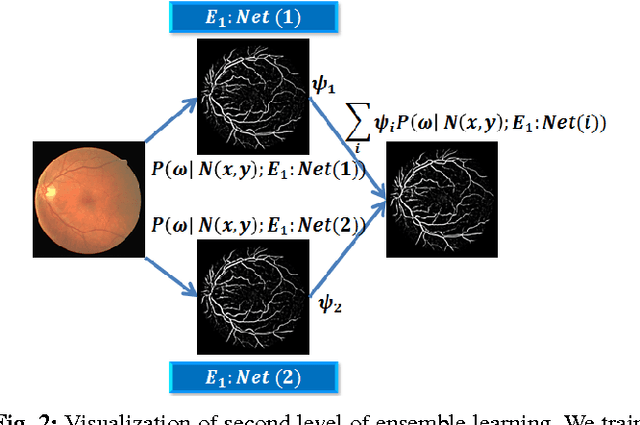


Abstract:Automated segmentation of retinal blood vessels in label-free fundus images entails a pivotal role in computed aided diagnosis of ophthalmic pathologies, viz., diabetic retinopathy, hypertensive disorders and cardiovascular diseases. The challenge remains active in medical image analysis research due to varied distribution of blood vessels, which manifest variations in their dimensions of physical appearance against a noisy background. In this paper we formulate the segmentation challenge as a classification task. Specifically, we employ unsupervised hierarchical feature learning using ensemble of two level of sparsely trained denoised stacked autoencoder. First level training with bootstrap samples ensures decoupling and second level ensemble formed by different network architectures ensures architectural revision. We show that ensemble training of auto-encoders fosters diversity in learning dictionary of visual kernels for vessel segmentation. SoftMax classifier is used for fine tuning each member auto-encoder and multiple strategies are explored for 2-level fusion of ensemble members. On DRIVE dataset, we achieve maximum average accuracy of 95.33\% with an impressively low standard deviation of 0.003 and Kappa agreement coefficient of 0.708 . Comparison with other major algorithms substantiates the high efficacy of our model.
DASA: Domain Adaptation in Stacked Autoencoders using Systematic Dropout
Mar 19, 2016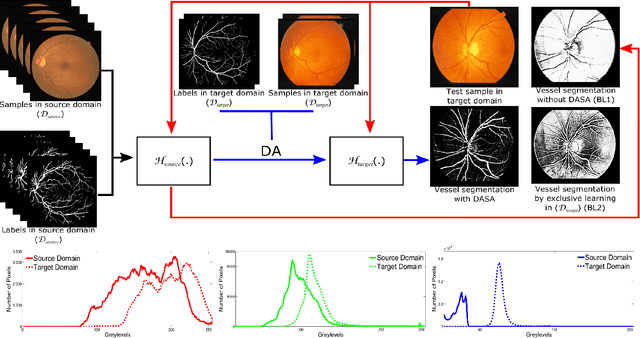
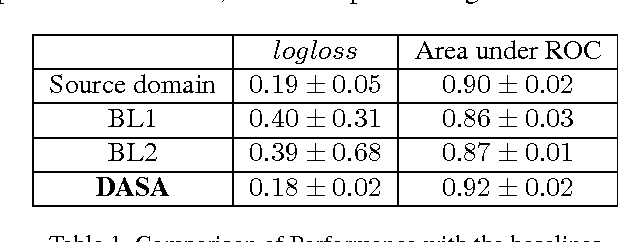
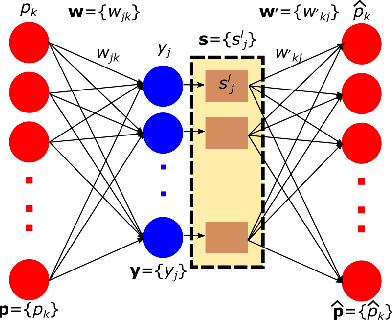
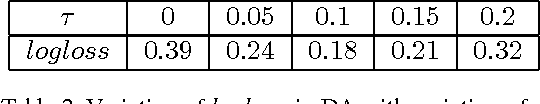
Abstract:Domain adaptation deals with adapting behaviour of machine learning based systems trained using samples in source domain to their deployment in target domain where the statistics of samples in both domains are dissimilar. The task of directly training or adapting a learner in the target domain is challenged by lack of abundant labeled samples. In this paper we propose a technique for domain adaptation in stacked autoencoder (SAE) based deep neural networks (DNN) performed in two stages: (i) unsupervised weight adaptation using systematic dropouts in mini-batch training, (ii) supervised fine-tuning with limited number of labeled samples in target domain. We experimentally evaluate performance in the problem of retinal vessel segmentation where the SAE-DNN is trained using large number of labeled samples in the source domain (DRIVE dataset) and adapted using less number of labeled samples in target domain (STARE dataset). The performance of SAE-DNN measured using $logloss$ in source domain is $0.19$, without and with adaptation are $0.40$ and $0.18$, and $0.39$ when trained exclusively with limited samples in target domain. The area under ROC curve is observed respectively as $0.90$, $0.86$, $0.92$ and $0.87$. The high efficiency of vessel segmentation with DASA strongly substantiates our claim.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge