Abhijeet Patil
Predicting Genetic Mutations from Single-Cell Bone Marrow Images in Acute Myeloid Leukemia Using Noise-Robust Deep Learning Models
Jun 15, 2025Abstract:In this study, we propose a robust methodology for identification of myeloid blasts followed by prediction of genetic mutation in single-cell images of blasts, tackling challenges associated with label accuracy and data noise. We trained an initial binary classifier to distinguish between leukemic (blasts) and non-leukemic cells images, achieving 90 percent accuracy. To evaluate the models generalization, we applied this model to a separate large unlabeled dataset and validated the predictions with two haemato-pathologists, finding an approximate error rate of 20 percent in the leukemic and non-leukemic labels. Assuming this level of label noise, we further trained a four-class model on images predicted as blasts to classify specific mutations. The mutation labels were known for only a bag of cell images extracted from a single slide. Despite the tumor label noise, our mutation classification model achieved 85 percent accuracy across four mutation classes, demonstrating resilience to label inconsistencies. This study highlights the capability of machine learning models to work with noisy labels effectively while providing accurate, clinically relevant mutation predictions, which is promising for diagnostic applications in areas such as haemato-pathology.
A Cytology Dataset for Early Detection of Oral Squamous Cell Carcinoma
Jun 11, 2025Abstract:Oral squamous cell carcinoma OSCC is a major global health burden, particularly in several regions across Asia, Africa, and South America, where it accounts for a significant proportion of cancer cases. Early detection dramatically improves outcomes, with stage I cancers achieving up to 90 percent survival. However, traditional diagnosis based on histopathology has limited accessibility in low-resource settings because it is invasive, resource-intensive, and reliant on expert pathologists. On the other hand, oral cytology of brush biopsy offers a minimally invasive and lower cost alternative, provided that the remaining challenges, inter observer variability and unavailability of expert pathologists can be addressed using artificial intelligence. Development and validation of robust AI solutions requires access to large, labeled, and multi-source datasets to train high capacity models that generalize across domain shifts. We introduce the first large and multicenter oral cytology dataset, comprising annotated slides stained with Papanicolaou(PAP) and May-Grunwald-Giemsa(MGG) protocols, collected from ten tertiary medical centers in India. The dataset is labeled and annotated by expert pathologists for cellular anomaly classification and detection, is designed to advance AI driven diagnostic methods. By filling the gap in publicly available oral cytology datasets, this resource aims to enhance automated detection, reduce diagnostic errors, and improve early OSCC diagnosis in resource-constrained settings, ultimately contributing to reduced mortality and better patient outcomes worldwide.
PathoGen-X: A Cross-Modal Genomic Feature Trans-Align Network for Enhanced Survival Prediction from Histopathology Images
Nov 01, 2024



Abstract:Accurate survival prediction is essential for personalized cancer treatment. However, genomic data - often a more powerful predictor than pathology data - is costly and inaccessible. We present the cross-modal genomic feature translation and alignment network for enhanced survival prediction from histopathology images (PathoGen-X). It is a deep learning framework that leverages both genomic and imaging data during training, relying solely on imaging data at testing. PathoGen-X employs transformer-based networks to align and translate image features into the genomic feature space, enhancing weaker imaging signals with stronger genomic signals. Unlike other methods, PathoGen-X translates and aligns features without projecting them to a shared latent space and requires fewer paired samples. Evaluated on TCGA-BRCA, TCGA-LUAD, and TCGA-GBM datasets, PathoGen-X demonstrates strong survival prediction performance, emphasizing the potential of enriched imaging models for accessible cancer prognosis.
Evaluation Metric for Quality Control and Generative Models in Histopathology Images
Nov 01, 2024



Abstract:Our study introduces ResNet-L2 (RL2), a novel metric for evaluating generative models and image quality in histopathology, addressing limitations of traditional metrics, such as Frechet inception distance (FID), when the data is scarce. RL2 leverages ResNet features with a normalizing flow to calculate RMSE distance in the latent space, providing reliable assessments across diverse histopathology datasets. We evaluated the performance of RL2 on degradation types, such as blur, Gaussian noise, salt-and-pepper noise, and rectangular patches, as well as diffusion processes. RL2's monotonic response to increasing degradation makes it well-suited for models that assess image quality, proving a valuable advancement for evaluating image generation techniques in histopathology. It can also be used to discard low-quality patches while sampling from a whole slide image. It is also significantly lighter and faster compared to traditional metrics and requires fewer images to give stable metric value.
Semantic Segmentation Based Quality Control of Histopathology Whole Slide Images
Oct 04, 2024



Abstract:We developed a software pipeline for quality control (QC) of histopathology whole slide images (WSIs) that segments various regions, such as blurs of different levels, tissue regions, tissue folds, and pen marks. Given the necessity and increasing availability of GPUs for processing WSIs, the proposed pipeline comprises multiple lightweight deep learning models to strike a balance between accuracy and speed. The pipeline was evaluated in all TCGAs, which is the largest publicly available WSI dataset containing more than 11,000 histopathological images from 28 organs. It was compared to a previous work, which was not based on deep learning, and it showed consistent improvement in segmentation results across organs. To minimize annotation effort for tissue and blur segmentation, annotated images were automatically prepared by mosaicking patches (sub-images) from various WSIs whose labels were identified using a patch classification tool HistoROI. Due to the generality of our trained QC pipeline and its extensive testing the potential impact of this work is broad. It can be used for automated pre-processing any WSI cohort to enhance the accuracy and reliability of large-scale histopathology image analysis for both research and clinical use. We have made the trained models, training scripts, training data, and inference results publicly available at https://github.com/abhijeetptl5/wsisegqc, which should enable the research community to use the pipeline right out of the box or further customize it to new datasets and applications in the future.
Efficient Quality Control of Whole Slide Pathology Images with Human-in-the-loop Training
Sep 29, 2024



Abstract:Histopathology whole slide images (WSIs) are being widely used to develop deep learning-based diagnostic solutions, especially for precision oncology. Most of these diagnostic softwares are vulnerable to biases and impurities in the training and test data which can lead to inaccurate diagnoses. For instance, WSIs contain multiple types of tissue regions, at least some of which might not be relevant to the diagnosis. We introduce HistoROI, a robust yet lightweight deep learning-based classifier to segregate WSI into six broad tissue regions -- epithelium, stroma, lymphocytes, adipose, artifacts, and miscellaneous. HistoROI is trained using a novel human-in-the-loop and active learning paradigm that ensures variations in training data for labeling-efficient generalization. HistoROI consistently performs well across multiple organs, despite being trained on only a single dataset, demonstrating strong generalization. Further, we have examined the utility of HistoROI in improving the performance of downstream deep learning-based tasks using the CAMELYON breast cancer lymph node and TCGA lung cancer datasets. For the former dataset, the area under the receiver operating characteristic curve (AUC) for metastasis versus normal tissue of a neural network trained using weakly supervised learning increased from 0.88 to 0.92 by filtering the data using HistoROI. Similarly, the AUC increased from 0.88 to 0.93 for the classification between adenocarcinoma and squamous cell carcinoma on the lung cancer dataset. We also found that the performance of the HistoROI improves upon HistoQC for artifact detection on a test dataset of 93 annotated WSIs. The limitations of the proposed model are analyzed, and potential extensions are also discussed.
* 18 pages
HER2 and FISH Status Prediction in Breast Biopsy H&E-Stained Images Using Deep Learning
Aug 25, 2024



Abstract:The current standard for detecting human epidermal growth factor receptor 2 (HER2) status in breast cancer patients relies on HER2 amplification, identified through fluorescence in situ hybridization (FISH) or immunohistochemistry (IHC). However, hematoxylin and eosin (H\&E) tumor stains are more widely available, and accurately predicting HER2 status using H\&E could reduce costs and expedite treatment selection. Deep Learning algorithms for H&E have shown effectiveness in predicting various cancer features and clinical outcomes, including moderate success in HER2 status prediction. In this work, we employed a customized weak supervision classification technique combined with MoCo-v2 contrastive learning to predict HER2 status. We trained our pipeline on 182 publicly available H&E Whole Slide Images (WSIs) from The Cancer Genome Atlas (TCGA), for which annotations by the pathology team at Yale School of Medicine are publicly available. Our pipeline achieved an Area Under the Curve (AUC) of 0.85 across four different test folds. Additionally, we tested our model on 44 H&E slides from the TCGA-BRCA dataset, which had an HER2 score of 2+ and included corresponding HER2 status and FISH test results. These cases are considered equivocal for IHC, requiring an expensive FISH test on their IHC slides for disambiguation. Our pipeline demonstrated an AUC of 0.81 on these challenging H&E slides. Reducing the need for FISH test can have significant implications in cancer treatment equity for underserved populations.
The ACROBAT 2022 Challenge: Automatic Registration Of Breast Cancer Tissue
May 29, 2023Abstract:The alignment of tissue between histopathological whole-slide-images (WSI) is crucial for research and clinical applications. Advances in computing, deep learning, and availability of large WSI datasets have revolutionised WSI analysis. Therefore, the current state-of-the-art in WSI registration is unclear. To address this, we conducted the ACROBAT challenge, based on the largest WSI registration dataset to date, including 4,212 WSIs from 1,152 breast cancer patients. The challenge objective was to align WSIs of tissue that was stained with routine diagnostic immunohistochemistry to its H&E-stained counterpart. We compare the performance of eight WSI registration algorithms, including an investigation of the impact of different WSI properties and clinical covariates. We find that conceptually distinct WSI registration methods can lead to highly accurate registration performances and identify covariates that impact performances across methods. These results establish the current state-of-the-art in WSI registration and guide researchers in selecting and developing methods.
Fast, Self Supervised, Fully Convolutional Color Normalization of H&E Stained Images
Nov 30, 2020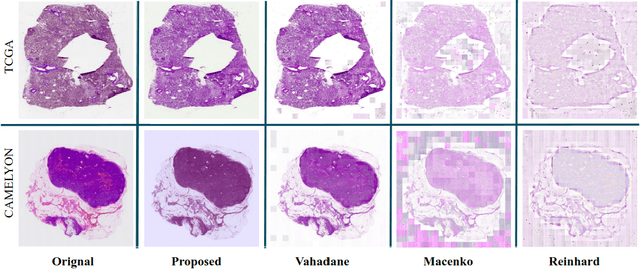
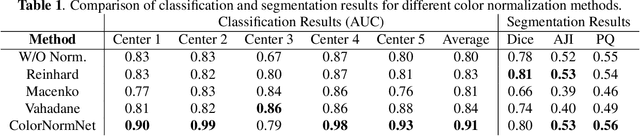
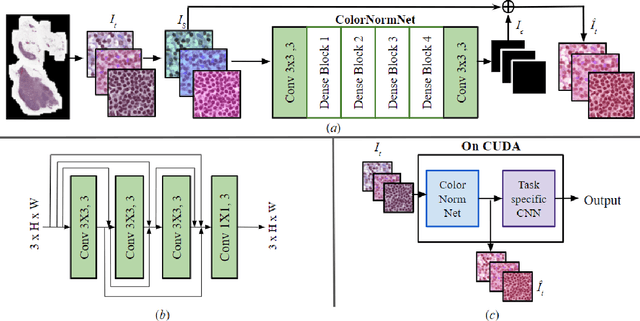
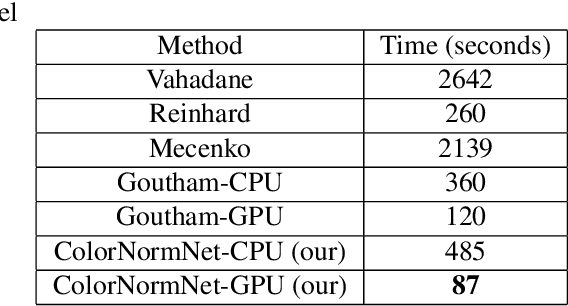
Abstract:Performance of deep learning algorithms decreases drastically if the data distributions of the training and testing sets are different. Due to variations in staining protocols, reagent brands, and habits of technicians, color variation in digital histopathology images is quite common. Color variation causes problems for the deployment of deep learning-based solutions for automatic diagnosis system in histopathology. Previously proposed color normalization methods consider a small patch as a reference for normalization, which creates artifacts on out-of-distribution source images. These methods are also slow as most of the computation is performed on CPUs instead of the GPUs. We propose a color normalization technique, which is fast during its self-supervised training as well as inference. Our method is based on a lightweight fully-convolutional neural network and can be easily attached to a deep learning-based pipeline as a pre-processing block. For classification and segmentation tasks on CAMELYON17 and MoNuSeg datasets respectively, the proposed method is faster and gives a greater increase in accuracy than the state of the art methods.
Visualization for Histopathology Images using Graph Convolutional Neural Networks
Jun 16, 2020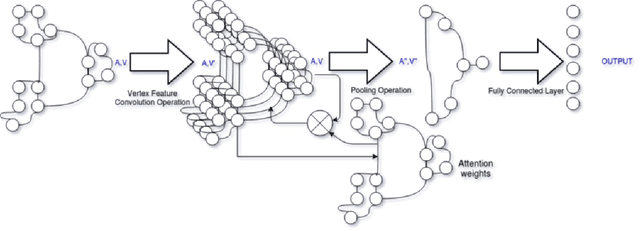
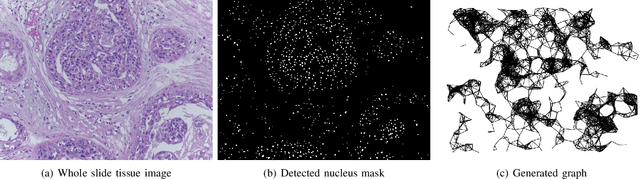
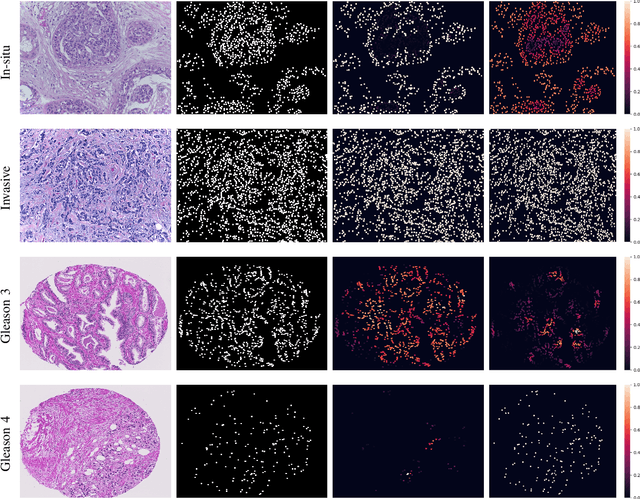
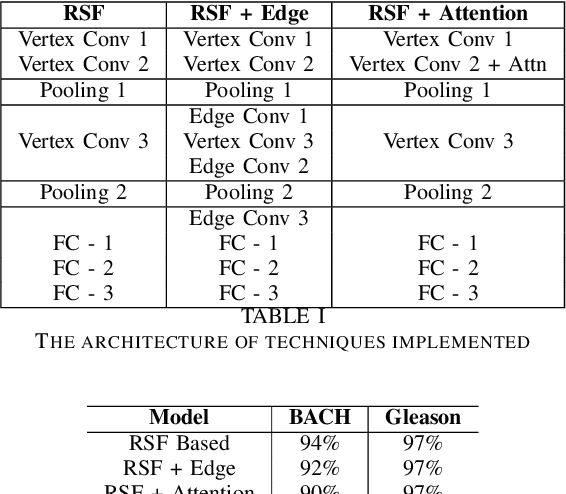
Abstract:With the increase in the use of deep learning for computer-aided diagnosis in medical images, the criticism of the black-box nature of the deep learning models is also on the rise. The medical community needs interpretable models for both due diligence and advancing the understanding of disease and treatment mechanisms. In histology, in particular, while there is rich detail available at the cellular level and that of spatial relationships between cells, it is difficult to modify convolutional neural networks to point out the relevant visual features. We adopt an approach to model histology tissue as a graph of nuclei and develop a graph convolutional network framework based on attention mechanism and node occlusion for disease diagnosis. The proposed method highlights the relative contribution of each cell nucleus in the whole-slide image. Our visualization of such networks trained to distinguish between invasive and in-situ breast cancers, and Gleason 3 and 4 prostate cancers generate interpretable visual maps that correspond well with our understanding of the structures that are important to experts for their diagnosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge