Swapnil Rane
Semantic Segmentation Based Quality Control of Histopathology Whole Slide Images
Oct 04, 2024



Abstract:We developed a software pipeline for quality control (QC) of histopathology whole slide images (WSIs) that segments various regions, such as blurs of different levels, tissue regions, tissue folds, and pen marks. Given the necessity and increasing availability of GPUs for processing WSIs, the proposed pipeline comprises multiple lightweight deep learning models to strike a balance between accuracy and speed. The pipeline was evaluated in all TCGAs, which is the largest publicly available WSI dataset containing more than 11,000 histopathological images from 28 organs. It was compared to a previous work, which was not based on deep learning, and it showed consistent improvement in segmentation results across organs. To minimize annotation effort for tissue and blur segmentation, annotated images were automatically prepared by mosaicking patches (sub-images) from various WSIs whose labels were identified using a patch classification tool HistoROI. Due to the generality of our trained QC pipeline and its extensive testing the potential impact of this work is broad. It can be used for automated pre-processing any WSI cohort to enhance the accuracy and reliability of large-scale histopathology image analysis for both research and clinical use. We have made the trained models, training scripts, training data, and inference results publicly available at https://github.com/abhijeetptl5/wsisegqc, which should enable the research community to use the pipeline right out of the box or further customize it to new datasets and applications in the future.
Efficient Quality Control of Whole Slide Pathology Images with Human-in-the-loop Training
Sep 29, 2024



Abstract:Histopathology whole slide images (WSIs) are being widely used to develop deep learning-based diagnostic solutions, especially for precision oncology. Most of these diagnostic softwares are vulnerable to biases and impurities in the training and test data which can lead to inaccurate diagnoses. For instance, WSIs contain multiple types of tissue regions, at least some of which might not be relevant to the diagnosis. We introduce HistoROI, a robust yet lightweight deep learning-based classifier to segregate WSI into six broad tissue regions -- epithelium, stroma, lymphocytes, adipose, artifacts, and miscellaneous. HistoROI is trained using a novel human-in-the-loop and active learning paradigm that ensures variations in training data for labeling-efficient generalization. HistoROI consistently performs well across multiple organs, despite being trained on only a single dataset, demonstrating strong generalization. Further, we have examined the utility of HistoROI in improving the performance of downstream deep learning-based tasks using the CAMELYON breast cancer lymph node and TCGA lung cancer datasets. For the former dataset, the area under the receiver operating characteristic curve (AUC) for metastasis versus normal tissue of a neural network trained using weakly supervised learning increased from 0.88 to 0.92 by filtering the data using HistoROI. Similarly, the AUC increased from 0.88 to 0.93 for the classification between adenocarcinoma and squamous cell carcinoma on the lung cancer dataset. We also found that the performance of the HistoROI improves upon HistoQC for artifact detection on a test dataset of 93 annotated WSIs. The limitations of the proposed model are analyzed, and potential extensions are also discussed.
* 18 pages
Combining Datasets with Different Label Sets for Improved Nucleus Segmentation and Classification
Oct 05, 2023



Abstract:Segmentation and classification of cell nuclei in histopathology images using deep neural networks (DNNs) can save pathologists' time for diagnosing various diseases, including cancers, by automating cell counting and morphometric assessments. It is now well-known that the accuracy of DNNs increases with the sizes of annotated datasets available for training. Although multiple datasets of histopathology images with nuclear annotations and class labels have been made publicly available, the set of class labels differ across these datasets. We propose a method to train DNNs for instance segmentation and classification on multiple datasets where the set of classes across the datasets are related but not the same. Specifically, our method is designed to utilize a coarse-to-fine class hierarchy, where the set of classes labeled and annotated in a dataset can be at any level of the hierarchy, as long as the classes are mutually exclusive. Within a dataset, the set of classes need not even be at the same level of the class hierarchy tree. Our results demonstrate that segmentation and classification metrics for the class set used by the test split of a dataset can improve by pre-training on another dataset that may even have a different set of classes due to the expansion of the training set enabled by our method. Furthermore, generalization to previously unseen datasets also improves by combining multiple other datasets with different sets of classes for training. The improvement is both qualitative and quantitative. The proposed method can be adapted for various loss functions, DNN architectures, and application domains.
EGFR Mutation Prediction of Lung Biopsy Images using Deep Learning
Aug 26, 2022
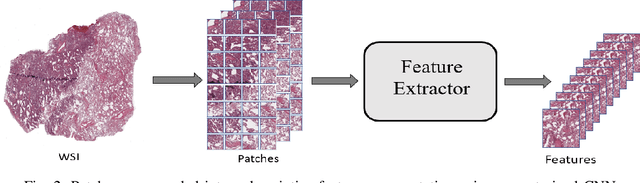
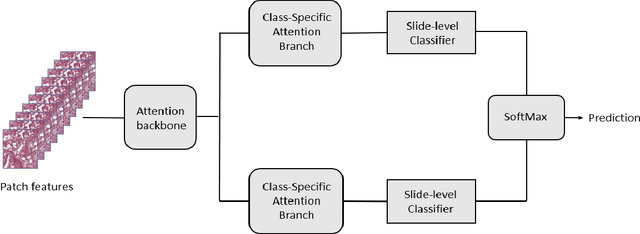
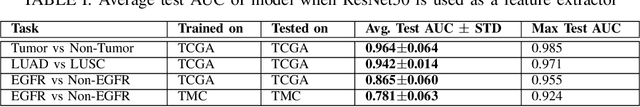
Abstract:The standard diagnostic procedures for targeted therapies in lung cancer treatment involve histological subtyping and subsequent detection of key driver mutations, such as EGFR. Even though molecular profiling can uncover the driver mutation, the process is often expensive and time-consuming. Deep learning-oriented image analysis offers a more economical alternative for discovering driver mutations directly from whole slide images (WSIs). In this work, we used customized deep learning pipelines with weak supervision to identify the morphological correlates of EGFR mutation from hematoxylin and eosin-stained WSIs, in addition to detecting tumor and histologically subtyping it. We demonstrate the effectiveness of our pipeline by conducting rigorous experiments and ablation studies on two lung cancer datasets - TCGA and a private dataset from India. With our pipeline, we achieved an average area under the curve (AUC) of 0.964 for tumor detection, and 0.942 for histological subtyping between adenocarcinoma and squamous cell carcinoma on the TCGA dataset. For EGFR detection, we achieved an average AUC of 0.864 on the TCGA dataset and 0.783 on the dataset from India. Our key learning points include the following. Firstly, there is no particular advantage of using a feature extractor layers trained on histology, if one is going to fine-tune the feature extractor on the target dataset. Secondly, selecting patches with high cellularity, presumably capturing tumor regions, is not always helpful, as the sign of a disease class may be present in the tumor-adjacent stroma.
Deep Multi-Scale U-Net Architecture and Noise-Robust Training Strategies for Histopathological Image Segmentation
May 03, 2022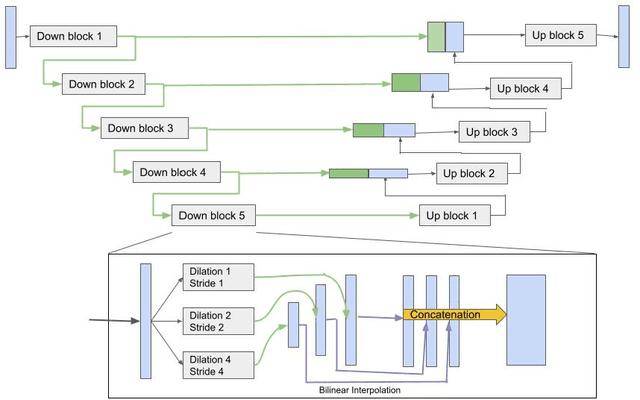
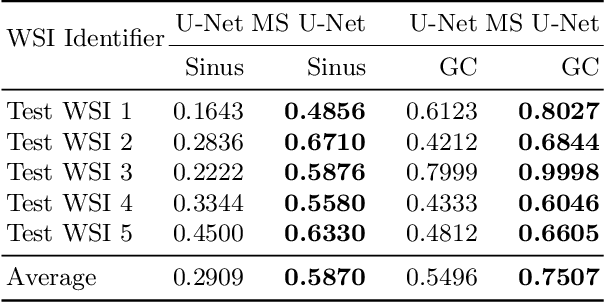

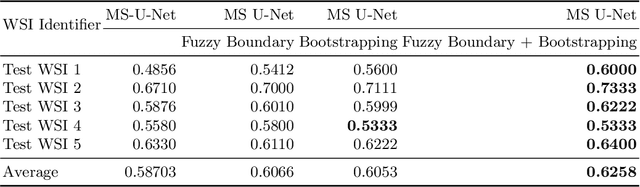
Abstract:Although the U-Net architecture has been extensively used for segmentation of medical images, we address two of its shortcomings in this work. Firstly, the accuracy of vanilla U-Net degrades when the target regions for segmentation exhibit significant variations in shape and size. Even though the U-Net already possesses some capability to analyze features at various scales, we propose to explicitly add multi-scale feature maps in each convolutional module of the U-Net encoder to improve segmentation of histology images. Secondly, the accuracy of a U-Net model also suffers when the annotations for supervised learning are noisy or incomplete. This can happen due to the inherent difficulty for a human expert to identify and delineate all instances of specific pathology very precisely and accurately. We address this challenge by introducing auxiliary confidence maps that emphasize less on the boundaries of the given target regions. Further, we utilize the bootstrapping properties of the deep network to address the missing annotation problem intelligently. In our experiments on a private dataset of breast cancer lymph nodes, where the primary task was to segment germinal centres and sinus histiocytosis, we observed substantial improvement over a U-Net baseline based on the two proposed augmentations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge