Abdur Rahman
SpatiaLab: Can Vision-Language Models Perform Spatial Reasoning in the Wild?
Feb 03, 2026Abstract:Spatial reasoning is a fundamental aspect of human cognition, yet it remains a major challenge for contemporary vision-language models (VLMs). Prior work largely relied on synthetic or LLM-generated environments with limited task designs and puzzle-like setups, failing to capture the real-world complexity, visual noise, and diverse spatial relationships that VLMs encounter. To address this, we introduce SpatiaLab, a comprehensive benchmark for evaluating VLMs' spatial reasoning in realistic, unconstrained contexts. SpatiaLab comprises 1,400 visual question-answer pairs across six major categories: Relative Positioning, Depth & Occlusion, Orientation, Size & Scale, Spatial Navigation, and 3D Geometry, each with five subcategories, yielding 30 distinct task types. Each subcategory contains at least 25 questions, and each main category includes at least 200 questions, supporting both multiple-choice and open-ended evaluation. Experiments across diverse state-of-the-art VLMs, including open- and closed-source models, reasoning-focused, and specialized spatial reasoning models, reveal a substantial gap in spatial reasoning capabilities compared with humans. In the multiple-choice setup, InternVL3.5-72B achieves 54.93% accuracy versus 87.57% for humans. In the open-ended setting, all models show a performance drop of around 10-25%, with GPT-5-mini scoring highest at 40.93% versus 64.93% for humans. These results highlight key limitations in handling complex spatial relationships, depth perception, navigation, and 3D geometry. By providing a diverse, real-world evaluation framework, SpatiaLab exposes critical challenges and opportunities for advancing VLMs' spatial reasoning, offering a benchmark to guide future research toward robust, human-aligned spatial understanding. SpatiaLab is available at: https://spatialab-reasoning.github.io/.
Echo-DND: A dual noise diffusion model for robust and precise left ventricle segmentation in echocardiography
Jun 18, 2025Abstract:Recent advancements in diffusion probabilistic models (DPMs) have revolutionized image processing, demonstrating significant potential in medical applications. Accurate segmentation of the left ventricle (LV) in echocardiograms is crucial for diagnostic procedures and necessary treatments. However, ultrasound images are notoriously noisy with low contrast and ambiguous LV boundaries, thereby complicating the segmentation process. To address these challenges, this paper introduces Echo-DND, a novel dual-noise diffusion model specifically designed for this task. Echo-DND leverages a unique combination of Gaussian and Bernoulli noises. It also incorporates a multi-scale fusion conditioning module to improve segmentation precision. Furthermore, it utilizes spatial coherence calibration to maintain spatial integrity in segmentation masks. The model's performance was rigorously validated on the CAMUS and EchoNet-Dynamic datasets. Extensive evaluations demonstrate that the proposed framework outperforms existing SOTA models. It achieves high Dice scores of 0.962 and 0.939 on these datasets, respectively. The proposed Echo-DND model establishes a new standard in echocardiogram segmentation, and its architecture holds promise for broader applicability in other medical imaging tasks, potentially improving diagnostic accuracy across various medical domains. Project page: https://abdur75648.github.io/Echo-DND
* Version of record published in Discover Applied Sciences (Springer Nature). The definitive article is available at https://doi.org/10.1007/s42452-025-07055-5
Preserving Cultural Identity with Context-Aware Translation Through Multi-Agent AI Systems
Mar 05, 2025Abstract:Language is a cornerstone of cultural identity, yet globalization and the dominance of major languages have placed nearly 3,000 languages at risk of extinction. Existing AI-driven translation models prioritize efficiency but often fail to capture cultural nuances, idiomatic expressions, and historical significance, leading to translations that marginalize linguistic diversity. To address these challenges, we propose a multi-agent AI framework designed for culturally adaptive translation in underserved language communities. Our approach leverages specialized agents for translation, interpretation, content synthesis, and bias evaluation, ensuring that linguistic accuracy and cultural relevance are preserved. Using CrewAI and LangChain, our system enhances contextual fidelity while mitigating biases through external validation. Comparative analysis shows that our framework outperforms GPT-4o, producing contextually rich and culturally embedded translations, a critical advancement for Indigenous, regional, and low-resource languages. This research underscores the potential of multi-agent AI in fostering equitable, sustainable, and culturally sensitive NLP technologies, aligning with the AI Governance, Cultural NLP, and Sustainable NLP pillars of Language Models for Underserved Communities. Our full experimental codebase is publicly available at: https://github.com/ciol-researchlab/Context-Aware_Translation_MAS
Dhoroni: Exploring Bengali Climate Change and Environmental Views with a Multi-Perspective News Dataset and Natural Language Processing
Oct 22, 2024



Abstract:Climate change poses critical challenges globally, disproportionately affecting low-income countries that often lack resources and linguistic representation on the international stage. Despite Bangladesh's status as one of the most vulnerable nations to climate impacts, research gaps persist in Bengali-language studies related to climate change and NLP. To address this disparity, we introduce Dhoroni, a novel Bengali (Bangla) climate change and environmental news dataset, comprising a 2300 annotated Bangla news articles, offering multiple perspectives such as political influence, scientific/statistical data, authenticity, stance detection, and stakeholder involvement. Furthermore, we present an in-depth exploratory analysis of Dhoroni and introduce BanglaBERT-Dhoroni family, a novel baseline model family for climate and environmental opinion detection in Bangla, fine-tuned on our dataset. This research contributes significantly to enhancing accessibility and analysis of climate discourse in Bengali (Bangla), addressing crucial communication and research gaps in climate-impacted regions like Bangladesh with 180 million people.
Activation Function Optimization Scheme for Image Classification
Sep 07, 2024Abstract:Activation function has a significant impact on the dynamics, convergence, and performance of deep neural networks. The search for a consistent and high-performing activation function has always been a pursuit during deep learning model development. Existing state-of-the-art activation functions are manually designed with human expertise except for Swish. Swish was developed using a reinforcement learning-based search strategy. In this study, we propose an evolutionary approach for optimizing activation functions specifically for image classification tasks, aiming to discover functions that outperform current state-of-the-art options. Through this optimization framework, we obtain a series of high-performing activation functions denoted as Exponential Error Linear Unit (EELU). The developed activation functions are evaluated for image classification tasks from two perspectives: (1) five state-of-the-art neural network architectures, such as ResNet50, AlexNet, VGG16, MobileNet, and Compact Convolutional Transformer which cover computationally heavy to light neural networks, and (2) eight standard datasets, including CIFAR10, Imagenette, MNIST, Fashion MNIST, Beans, Colorectal Histology, CottonWeedID15, and TinyImageNet which cover from typical machine vision benchmark, agricultural image applications to medical image applications. Finally, we statistically investigate the generalization of the resultant activation functions developed through the optimization scheme. With a Friedman test, we conclude that the optimization scheme is able to generate activation functions that outperform the existing standard ones in 92.8% cases among 28 different cases studied, and $-x\cdot erf(e^{-x})$ is found to be the best activation function for image classification generated by the optimization scheme.
MoistNet: Machine Vision-based Deep Learning Models for Wood Chip Moisture Content Measurement
Sep 07, 2024Abstract:Quick and reliable measurement of wood chip moisture content is an everlasting problem for numerous forest-reliant industries such as biofuel, pulp and paper, and bio-refineries. Moisture content is a critical attribute of wood chips due to its direct relationship with the final product quality. Conventional techniques for determining moisture content, such as oven-drying, possess some drawbacks in terms of their time-consuming nature, potential sample damage, and lack of real-time feasibility. Furthermore, alternative techniques, including NIR spectroscopy, electrical capacitance, X-rays, and microwaves, have demonstrated potential; nevertheless, they are still constrained by issues related to portability, precision, and the expense of the required equipment. Hence, there is a need for a moisture content determination method that is instant, portable, non-destructive, inexpensive, and precise. This study explores the use of deep learning and machine vision to predict moisture content classes from RGB images of wood chips. A large-scale image dataset comprising 1,600 RGB images of wood chips has been collected and annotated with ground truth labels, utilizing the results of the oven-drying technique. Two high-performing neural networks, MoistNetLite and MoistNetMax, have been developed leveraging Neural Architecture Search (NAS) and hyperparameter optimization. The developed models are evaluated and compared with state-of-the-art deep learning models. Results demonstrate that MoistNetLite achieves 87% accuracy with minimal computational overhead, while MoistNetMax exhibits exceptional precision with a 91% accuracy in wood chip moisture content class prediction. With improved accuracy and faster prediction speed, our proposed MoistNet models hold great promise for the wood chip processing industry.
V-Zen: Efficient GUI Understanding and Precise Grounding With A Novel Multimodal LLM
May 24, 2024



Abstract:In the rapidly evolving landscape of AI research and application, Multimodal Large Language Models (MLLMs) have emerged as a transformative force, adept at interpreting and integrating information from diverse modalities such as text, images, and Graphical User Interfaces (GUIs). Despite these advancements, the nuanced interaction and understanding of GUIs pose a significant challenge, limiting the potential of existing models to enhance automation levels. To bridge this gap, this paper presents V-Zen, an innovative Multimodal Large Language Model (MLLM) meticulously crafted to revolutionise the domain of GUI understanding and grounding. Equipped with dual-resolution image encoders, V-Zen establishes new benchmarks in efficient grounding and next-action prediction, thereby laying the groundwork for self-operating computer systems. Complementing V-Zen is the GUIDE dataset, an extensive collection of real-world GUI elements and task-based sequences, serving as a catalyst for specialised fine-tuning. The successful integration of V-Zen and GUIDE marks the dawn of a new era in multimodal AI research, opening the door to intelligent, autonomous computing experiences. This paper extends an invitation to the research community to join this exciting journey, shaping the future of GUI automation. In the spirit of open science, our code, data, and model will be made publicly available, paving the way for multimodal dialogue scenarios with intricate and precise interactions.
UTRNet: High-Resolution Urdu Text Recognition In Printed Documents
Jul 06, 2023Abstract:In this paper, we propose a novel approach to address the challenges of printed Urdu text recognition using high-resolution, multi-scale semantic feature extraction. Our proposed UTRNet architecture, a hybrid CNN-RNN model, demonstrates state-of-the-art performance on benchmark datasets. To address the limitations of previous works, which struggle to generalize to the intricacies of the Urdu script and the lack of sufficient annotated real-world data, we have introduced the UTRSet-Real, a large-scale annotated real-world dataset comprising over 11,000 lines and UTRSet-Synth, a synthetic dataset with 20,000 lines closely resembling real-world and made corrections to the ground truth of the existing IIITH dataset, making it a more reliable resource for future research. We also provide UrduDoc, a benchmark dataset for Urdu text line detection in scanned documents. Additionally, we have developed an online tool for end-to-end Urdu OCR from printed documents by integrating UTRNet with a text detection model. Our work not only addresses the current limitations of Urdu OCR but also paves the way for future research in this area and facilitates the continued advancement of Urdu OCR technology. The project page with source code, datasets, annotations, trained models, and online tool is available at abdur75648.github.io/UTRNet.
Fake Hilsa Fish Detection Using Machine Vision
Jan 08, 2022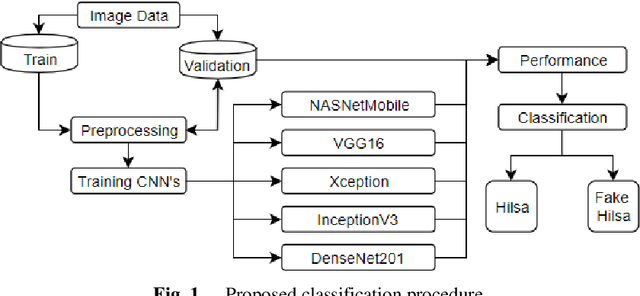
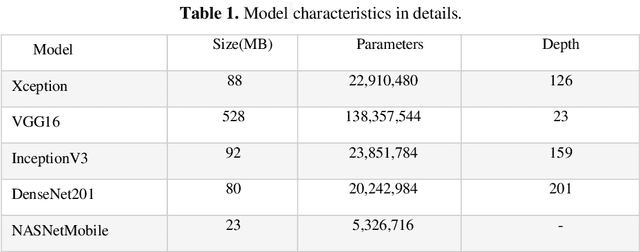
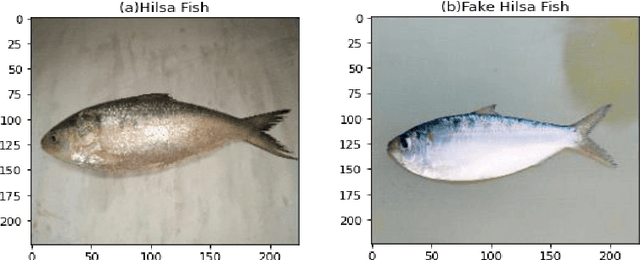
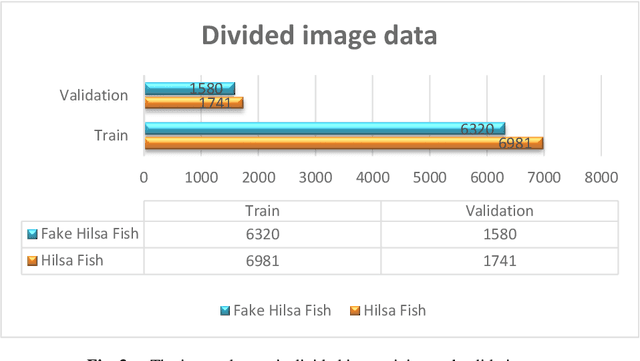
Abstract:Hilsa is the national fish of Bangladesh. Bangladesh is earning a lot of foreign currency by exporting this fish. Unfortunately, in recent days, some unscrupulous businessmen are selling fake Hilsa fishes to gain profit. The Sardines and Sardinella are the most sold in the market as Hilsa. The government agency of Bangladesh, namely Bangladesh Food Safety Authority said that these fake Hilsa fish contain high levels of cadmium and lead which are detrimental for humans. In this research, we have proposed a method that can readily identify original Hilsa fish and fake Hilsa fish. Based on the research available on online literature, we are the first to do research on identifying original Hilsa fish. We have collected more than 16,000 images of original and counterfeit Hilsa fish. To classify these images, we have used several deep learning-based models. Then, the performance has been compared between them. Among those models, DenseNet201 achieved the highest accuracy of 97.02%.
A Gradient Mapping Guided Explainable Deep Neural Network for Extracapsular Extension Identification in 3D Head and Neck Cancer Computed Tomography Images
Jan 03, 2022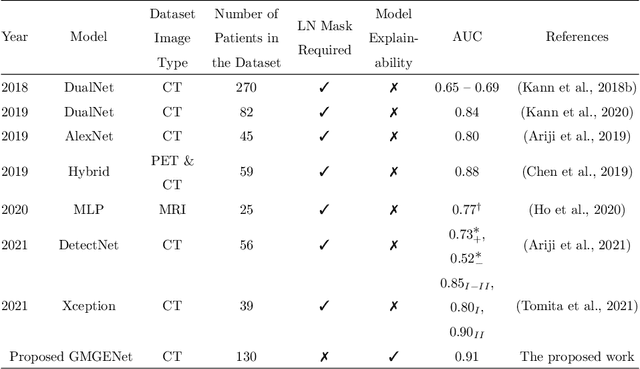
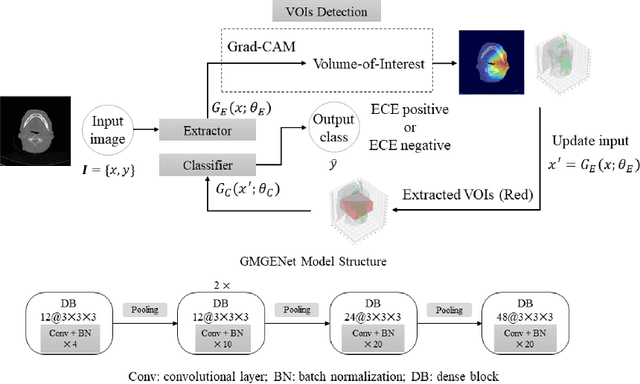
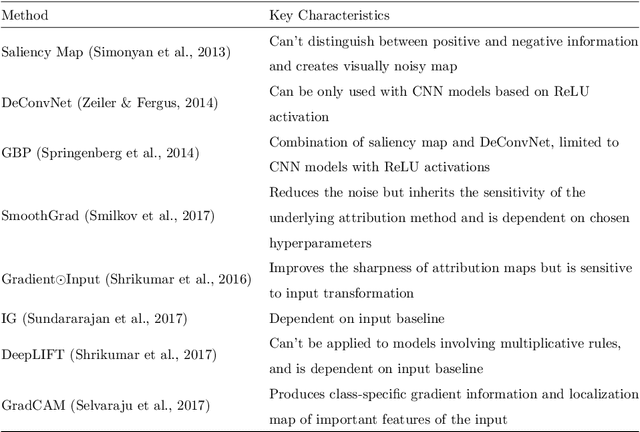
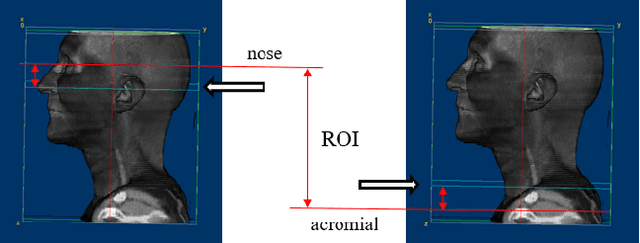
Abstract:Diagnosis and treatment management for head and neck squamous cell carcinoma (HNSCC) is guided by routine diagnostic head and neck computed tomography (CT) scans to identify tumor and lymph node features. Extracapsular extension (ECE) is a strong predictor of patients' survival outcomes with HNSCC. It is essential to detect the occurrence of ECE as it changes staging and management for the patients. Current clinical ECE detection relies on visual identification and pathologic confirmation conducted by radiologists. Machine learning (ML)-based ECE diagnosis has shown high potential in the recent years. However, manual annotation of lymph node region is a required data preprocessing step in most of the current ML-based ECE diagnosis studies. In addition, this manual annotation process is time-consuming, labor-intensive, and error-prone. Therefore, in this paper, we propose a Gradient Mapping Guided Explainable Network (GMGENet) framework to perform ECE identification automatically without requiring annotated lymph node region information. The gradient-weighted class activation mapping (Grad-CAM) technique is proposed to guide the deep learning algorithm to focus on the regions that are highly related to ECE. Informative volumes of interest (VOIs) are extracted without labeled lymph node region information. In evaluation, the proposed method is well-trained and tested using cross validation, achieving test accuracy and AUC of 90.2% and 91.1%, respectively. The presence or absence of ECE has been analyzed and correlated with gold standard histopathological findings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge