Zohaib Salahuddin
on behalf of EUCanImage working group
Robust Multicentre Detection and Classification of Colorectal Liver Metastases on CT: Application of Foundation Models
Jan 12, 2026Abstract:Colorectal liver metastases (CRLM) are a major cause of cancer-related mortality, and reliable detection on CT remains challenging in multi-centre settings. We developed a foundation model-based AI pipeline for patient-level classification and lesion-level detection of CRLM on contrast-enhanced CT, integrating uncertainty quantification and explainability. CT data from the EuCanImage consortium (n=2437) and an external TCIA cohort (n=197) were used. Among several pretrained models, UMedPT achieved the best performance and was fine-tuned with an MLP head for classification and an FCOS-based head for lesion detection. The classification model achieved an AUC of 0.90 and a sensitivity of 0.82 on the combined test set, with a sensitivity of 0.85 on the external cohort. Excluding the most uncertain 20 percent of cases improved AUC to 0.91 and balanced accuracy to 0.86. Decision curve analysis showed clinical benefit for threshold probabilities between 0.30 and 0.40. The detection model identified 69.1 percent of lesions overall, increasing from 30 percent to 98 percent across lesion size quartiles. Grad-CAM highlighted lesion-corresponding regions in high-confidence cases. These results demonstrate that foundation model-based pipelines can support robust and interpretable CRLM detection and classification across heterogeneous CT data.
Radiology Report Conditional 3D CT Generation with Multi Encoder Latent diffusion Model
Sep 18, 2025



Abstract:Text to image latent diffusion models have recently advanced medical image synthesis, but applications to 3D CT generation remain limited. Existing approaches rely on simplified prompts, neglecting the rich semantic detail in full radiology reports, which reduces text image alignment and clinical fidelity. We propose Report2CT, a radiology report conditional latent diffusion framework for synthesizing 3D chest CT volumes directly from free text radiology reports, incorporating both findings and impression sections using multiple text encoder. Report2CT integrates three pretrained medical text encoders (BiomedVLP CXR BERT, MedEmbed, and ClinicalBERT) to capture nuanced clinical context. Radiology reports and voxel spacing information condition a 3D latent diffusion model trained on 20000 CT volumes from the CT RATE dataset. Model performance was evaluated using Frechet Inception Distance (FID) for real synthetic distributional similarity and CLIP based metrics for semantic alignment, with additional qualitative and quantitative comparisons against GenerateCT model. Report2CT generated anatomically consistent CT volumes with excellent visual quality and text image alignment. Multi encoder conditioning improved CLIP scores, indicating stronger preservation of fine grained clinical details in the free text radiology reports. Classifier free guidance further enhanced alignment with only a minor trade off in FID. We ranked first in the VLM3D Challenge at MICCAI 2025 on Text Conditional CT Generation and achieved state of the art performance across all evaluation metrics. By leveraging complete radiology reports and multi encoder text conditioning, Report2CT advances 3D CT synthesis, producing clinically faithful and high quality synthetic data.
Pixels to Prognosis: Harmonized Multi-Region CT-Radiomics and Foundation-Model Signatures Across Multicentre NSCLC Data
May 23, 2025Abstract:Purpose: To evaluate the impact of harmonization and multi-region CT image feature integration on survival prediction in non-small cell lung cancer (NSCLC) patients, using handcrafted radiomics, pretrained foundation model (FM) features, and clinical data from a multicenter dataset. Methods: We analyzed CT scans and clinical data from 876 NSCLC patients (604 training, 272 test) across five centers. Features were extracted from the whole lung, tumor, mediastinal nodes, coronary arteries, and coronary artery calcium (CAC). Handcrafted radiomics and FM deep features were harmonized using ComBat, reconstruction kernel normalization (RKN), and RKN+ComBat. Regularized Cox models predicted overall survival; performance was assessed using the concordance index (C-index), 5-year time-dependent area under the curve (t-AUC), and hazard ratio (HR). SHapley Additive exPlanations (SHAP) values explained feature contributions. A consensus model used agreement across top region of interest (ROI) models to stratify patient risk. Results: TNM staging showed prognostic utility (C-index = 0.67; HR = 2.70; t-AUC = 0.85). The clinical + tumor radiomics model with ComBat achieved a C-index of 0.7552 and t-AUC of 0.8820. FM features (50-voxel cubes) combined with clinical data yielded the highest performance (C-index = 0.7616; t-AUC = 0.8866). An ensemble of all ROIs and FM features reached a C-index of 0.7142 and t-AUC of 0.7885. The consensus model, covering 78% of valid test cases, achieved a t-AUC of 0.92, sensitivity of 97.6%, and specificity of 66.7%. Conclusion: Harmonization and multi-region feature integration improve survival prediction in multicenter NSCLC data. Combining interpretable radiomics, FM features, and consensus modeling enables robust risk stratification across imaging centers.
Explainable Anatomy-Guided AI for Prostate MRI: Foundation Models and In Silico Clinical Trials for Virtual Biopsy-based Risk Assessment
May 23, 2025Abstract:We present a fully automated, anatomically guided deep learning pipeline for prostate cancer (PCa) risk stratification using routine MRI. The pipeline integrates three key components: an nnU-Net module for segmenting the prostate gland and its zones on axial T2-weighted MRI; a classification module based on the UMedPT Swin Transformer foundation model, fine-tuned on 3D patches with optional anatomical priors and clinical data; and a VAE-GAN framework for generating counterfactual heatmaps that localize decision-driving image regions. The system was developed using 1,500 PI-CAI cases for segmentation and 617 biparametric MRIs with metadata from the CHAIMELEON challenge for classification (split into 70% training, 10% validation, and 20% testing). Segmentation achieved mean Dice scores of 0.95 (gland), 0.94 (peripheral zone), and 0.92 (transition zone). Incorporating gland priors improved AUC from 0.69 to 0.72, with a three-scale ensemble achieving top performance (AUC = 0.79, composite score = 0.76), outperforming the 2024 CHAIMELEON challenge winners. Counterfactual heatmaps reliably highlighted lesions within segmented regions, enhancing model interpretability. In a prospective multi-center in-silico trial with 20 clinicians, AI assistance increased diagnostic accuracy from 0.72 to 0.77 and Cohen's kappa from 0.43 to 0.53, while reducing review time per case by 40%. These results demonstrate that anatomy-aware foundation models with counterfactual explainability can enable accurate, interpretable, and efficient PCa risk assessment, supporting their potential use as virtual biopsies in clinical practice.
A Foundation Model Framework for Multi-View MRI Classification of Extramural Vascular Invasion and Mesorectal Fascia Invasion in Rectal Cancer
May 23, 2025Abstract:Background: Accurate MRI-based identification of extramural vascular invasion (EVI) and mesorectal fascia invasion (MFI) is pivotal for risk-stratified management of rectal cancer, yet visual assessment is subjective and vulnerable to inter-institutional variability. Purpose: To develop and externally evaluate a multicenter, foundation-model-driven framework that automatically classifies EVI and MFI on axial and sagittal T2-weighted MRI. Methods: This retrospective study used 331 pre-treatment rectal cancer MRI examinations from three European hospitals. After TotalSegmentator-guided rectal patch extraction, a self-supervised frequency-domain harmonization pipeline was trained to minimize scanner-related contrast shifts. Four classifiers were compared: ResNet50, SeResNet, the universal biomedical pretrained transformer (UMedPT) with a lightweight MLP head, and a logistic-regression variant using frozen UMedPT features (UMedPT_LR). Results: UMedPT_LR achieved the best EVI detection when axial and sagittal features were fused (AUC = 0.82; sensitivity = 0.75; F1 score = 0.73), surpassing the Chaimeleon Grand-Challenge winner (AUC = 0.74). The highest MFI performance was attained by UMedPT on axial harmonized images (AUC = 0.77), surpassing the Chaimeleon Grand-Challenge winner (AUC = 0.75). Frequency-domain harmonization improved MFI classification but variably affected EVI performance. Conventional CNNs (ResNet50, SeResNet) underperformed, especially in F1 score and balanced accuracy. Conclusion: These findings demonstrate that combining foundation model features, harmonization, and multi-view fusion significantly enhances diagnostic performance in rectal MRI.
Counterfactuals and Uncertainty-Based Explainable Paradigm for the Automated Detection and Segmentation of Renal Cysts in Computed Tomography Images: A Multi-Center Study
Aug 07, 2024Abstract:Routine computed tomography (CT) scans often detect a wide range of renal cysts, some of which may be malignant. Early and precise localization of these cysts can significantly aid quantitative image analysis. Current segmentation methods, however, do not offer sufficient interpretability at the feature and pixel levels, emphasizing the necessity for an explainable framework that can detect and rectify model inaccuracies. We developed an interpretable segmentation framework and validated it on a multi-centric dataset. A Variational Autoencoder Generative Adversarial Network (VAE-GAN) was employed to learn the latent representation of 3D input patches and reconstruct input images. Modifications in the latent representation using the gradient of the segmentation model generated counterfactual explanations for varying dice similarity coefficients (DSC). Radiomics features extracted from these counterfactual images, using a ground truth cyst mask, were analyzed to determine their correlation with segmentation performance. The DSCs for the original and VAE-GAN reconstructed images for counterfactual image generation showed no significant differences. Counterfactual explanations highlighted how variations in cyst image features influence segmentation outcomes and showed model discrepancies. Radiomics features correlating positively and negatively with dice scores were identified. The uncertainty of the predicted segmentation masks was estimated using posterior sampling of the weight space. The combination of counterfactual explanations and uncertainty maps provided a deeper understanding of the image features within the segmented renal cysts that lead to high uncertainty. The proposed segmentation framework not only achieved high segmentation accuracy but also increased interpretability regarding how image features impact segmentation performance.
Methodological Explainability Evaluation of an Interpretable Deep Learning Model for Post-Hepatectomy Liver Failure Prediction Incorporating Counterfactual Explanations and Layerwise Relevance Propagation: A Prospective In Silico Trial
Aug 07, 2024



Abstract:Artificial intelligence (AI)-based decision support systems have demonstrated value in predicting post-hepatectomy liver failure (PHLF) in hepatocellular carcinoma (HCC). However, they often lack transparency, and the impact of model explanations on clinicians' decisions has not been thoroughly evaluated. Building on prior research, we developed a variational autoencoder-multilayer perceptron (VAE-MLP) model for preoperative PHLF prediction. This model integrated counterfactuals and layerwise relevance propagation (LRP) to provide insights into its decision-making mechanism. Additionally, we proposed a methodological framework for evaluating the explainability of AI systems. This framework includes qualitative and quantitative assessments of explanations against recognized biomarkers, usability evaluations, and an in silico clinical trial. Our evaluations demonstrated that the model's explanation correlated with established biomarkers and exhibited high usability at both the case and system levels. Furthermore, results from the three-track in silico clinical trial showed that clinicians' prediction accuracy and confidence increased when AI explanations were provided.
Precision-medicine-toolbox: An open-source python package for facilitation of quantitative medical imaging and radiomics analysis
Feb 28, 2022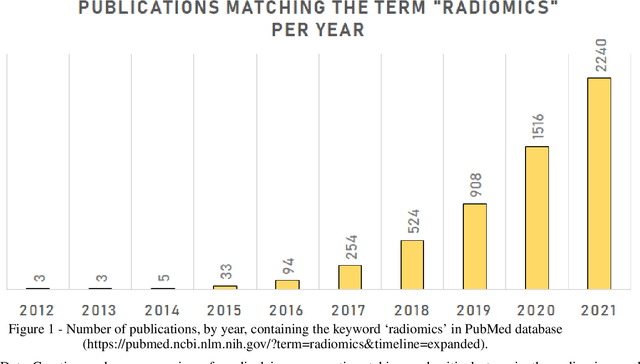
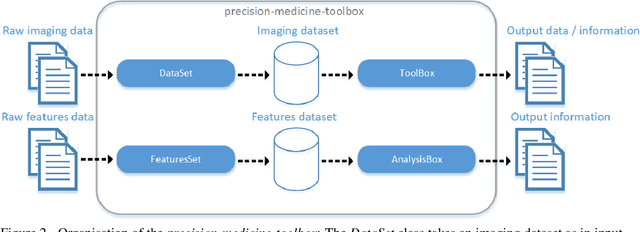
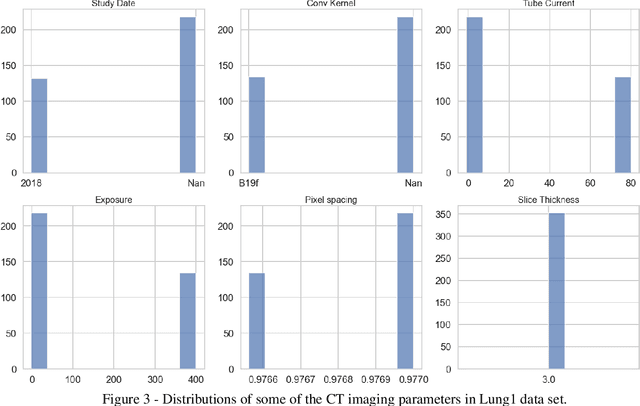
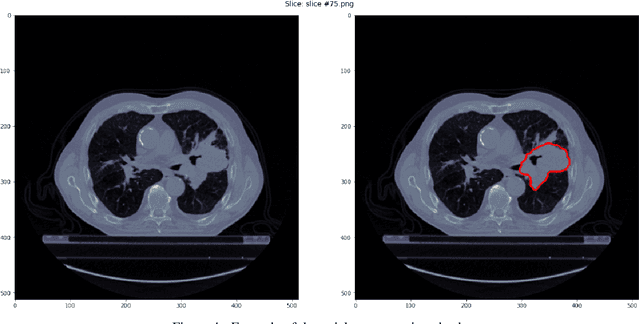
Abstract:Medical image analysis plays a key role in precision medicine as it allows the clinicians to identify anatomical abnormalities and it is routinely used in clinical assessment. Data curation and pre-processing of medical images are critical steps in the quantitative medical image analysis that can have a significant impact on the resulting model performance. In this paper, we introduce a precision-medicine-toolbox that allows researchers to perform data curation, image pre-processing and handcrafted radiomics extraction (via Pyradiomics) and feature exploration tasks with Python. With this open-source solution, we aim to address the data preparation and exploration problem, bridge the gap between the currently existing packages, and improve the reproducibility of quantitative medical imaging research.
Transparency of Deep Neural Networks for Medical Image Analysis: A Review of Interpretability Methods
Nov 01, 2021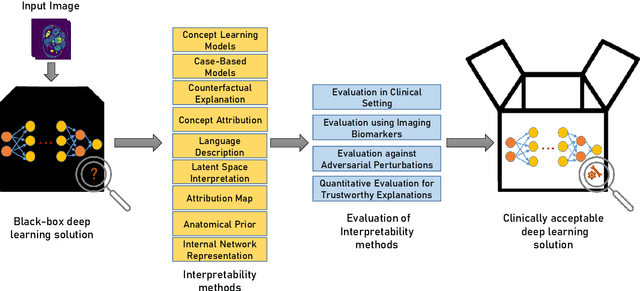
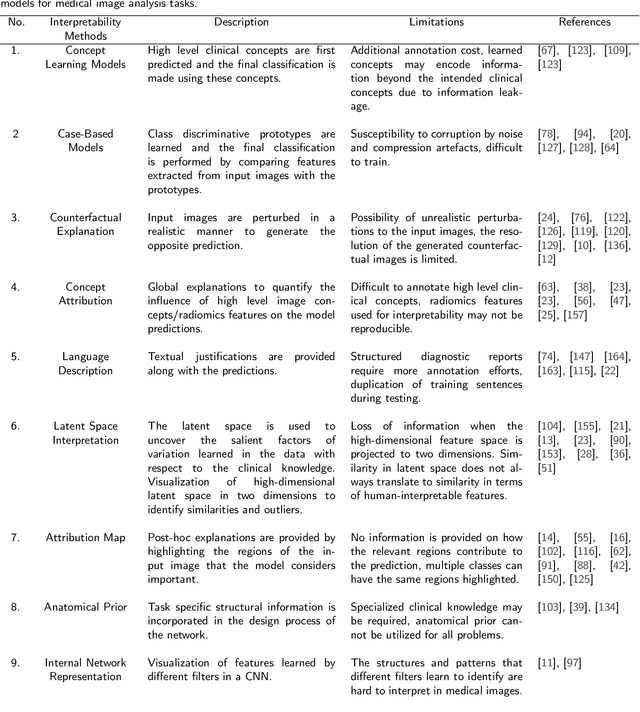
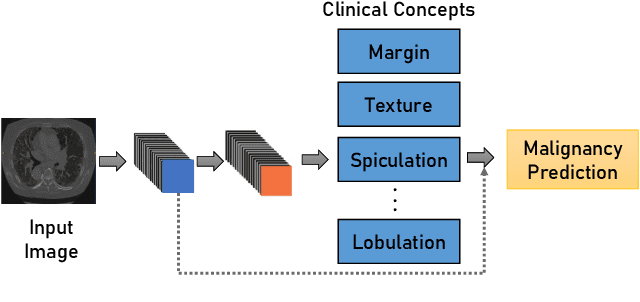
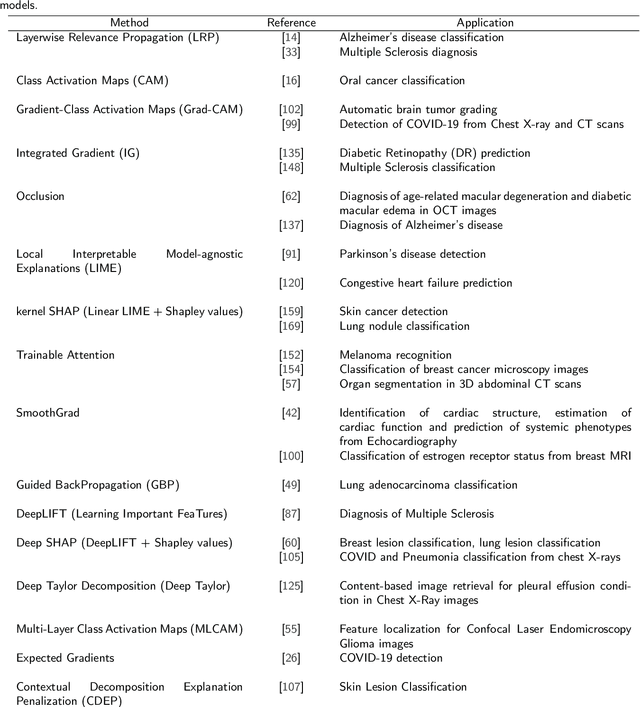
Abstract:Artificial Intelligence has emerged as a useful aid in numerous clinical applications for diagnosis and treatment decisions. Deep neural networks have shown same or better performance than clinicians in many tasks owing to the rapid increase in the available data and computational power. In order to conform to the principles of trustworthy AI, it is essential that the AI system be transparent, robust, fair and ensure accountability. Current deep neural solutions are referred to as black-boxes due to a lack of understanding of the specifics concerning the decision making process. Therefore, there is a need to ensure interpretability of deep neural networks before they can be incorporated in the routine clinical workflow. In this narrative review, we utilized systematic keyword searches and domain expertise to identify nine different types of interpretability methods that have been used for understanding deep learning models for medical image analysis applications based on the type of generated explanations and technical similarities. Furthermore, we report the progress made towards evaluating the explanations produced by various interpretability methods. Finally we discuss limitations, provide guidelines for using interpretability methods and future directions concerning the interpretability of deep neural networks for medical imaging analysis.
FUTURE-AI: Guiding Principles and Consensus Recommendations for Trustworthy Artificial Intelligence in Medical Imaging
Sep 29, 2021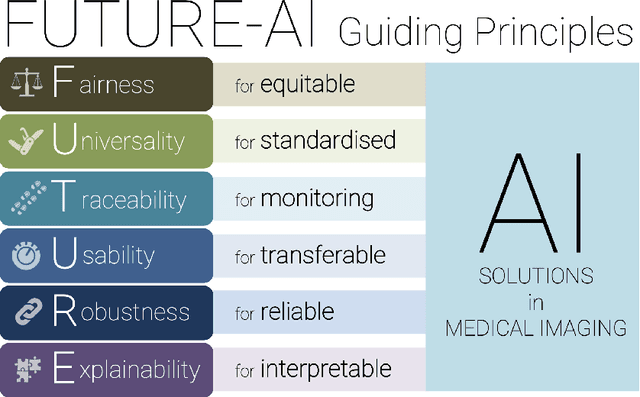
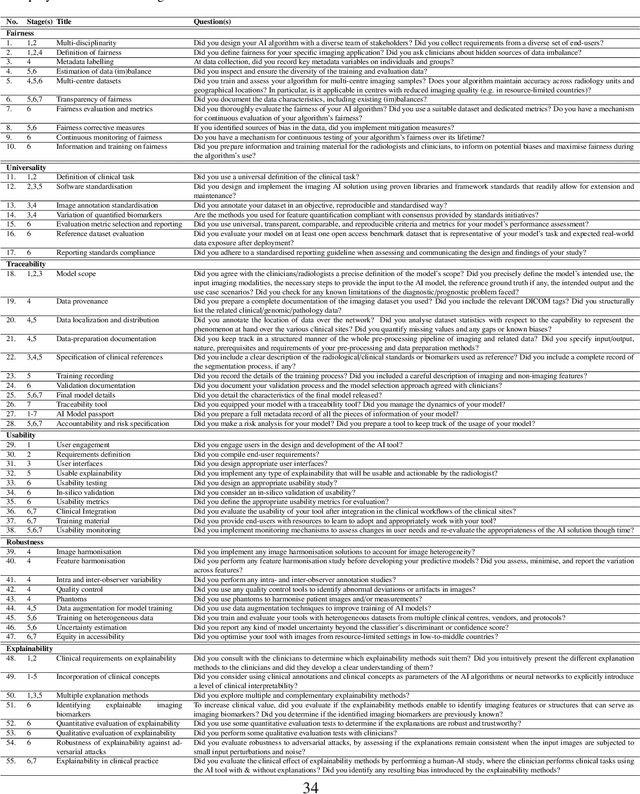
Abstract:The recent advancements in artificial intelligence (AI) combined with the extensive amount of data generated by today's clinical systems, has led to the development of imaging AI solutions across the whole value chain of medical imaging, including image reconstruction, medical image segmentation, image-based diagnosis and treatment planning. Notwithstanding the successes and future potential of AI in medical imaging, many stakeholders are concerned of the potential risks and ethical implications of imaging AI solutions, which are perceived as complex, opaque, and difficult to comprehend, utilise, and trust in critical clinical applications. Despite these concerns and risks, there are currently no concrete guidelines and best practices for guiding future AI developments in medical imaging towards increased trust, safety and adoption. To bridge this gap, this paper introduces a careful selection of guiding principles drawn from the accumulated experiences, consensus, and best practices from five large European projects on AI in Health Imaging. These guiding principles are named FUTURE-AI and its building blocks consist of (i) Fairness, (ii) Universality, (iii) Traceability, (iv) Usability, (v) Robustness and (vi) Explainability. In a step-by-step approach, these guidelines are further translated into a framework of concrete recommendations for specifying, developing, evaluating, and deploying technically, clinically and ethically trustworthy AI solutions into clinical practice.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge