Yujin Oh
LLM-driven Multimodal Target Volume Contouring in Radiation Oncology
Nov 03, 2023Abstract:Target volume contouring for radiation therapy is considered significantly more challenging than the normal organ segmentation tasks as it necessitates the utilization of both image and text-based clinical information. Inspired by the recent advancement of large language models (LLMs) that can facilitate the integration of the textural information and images, here we present a novel LLM-driven multi-modal AI that utilizes the clinical text information and is applicable to the challenging task of target volume contouring for radiation therapy, and validate it within the context of breast cancer radiation therapy target volume contouring. Using external validation and data-insufficient environments, which attributes highly conducive to real-world applications, we demonstrate that the proposed model exhibits markedly improved performance compared to conventional vision-only AI models, particularly exhibiting robust generalization performance and data-efficiency. To our best knowledge, this is the first LLM-driven multimodal AI model that integrates the clinical text information into target volume delineation for radiation oncology.
C-DARL: Contrastive diffusion adversarial representation learning for label-free blood vessel segmentation
Jul 31, 2023Abstract:Blood vessel segmentation in medical imaging is one of the essential steps for vascular disease diagnosis and interventional planning in a broad spectrum of clinical scenarios in image-based medicine and interventional medicine. Unfortunately, manual annotation of the vessel masks is challenging and resource-intensive due to subtle branches and complex structures. To overcome this issue, this paper presents a self-supervised vessel segmentation method, dubbed the contrastive diffusion adversarial representation learning (C-DARL) model. Our model is composed of a diffusion module and a generation module that learns the distribution of multi-domain blood vessel data by generating synthetic vessel images from diffusion latent. Moreover, we employ contrastive learning through a mask-based contrastive loss so that the model can learn more realistic vessel representations. To validate the efficacy, C-DARL is trained using various vessel datasets, including coronary angiograms, abdominal digital subtraction angiograms, and retinal imaging. Experimental results confirm that our model achieves performance improvement over baseline methods with noise robustness, suggesting the effectiveness of C-DARL for vessel segmentation.
ZegOT: Zero-shot Segmentation Through Optimal Transport of Text Prompts
Jan 28, 2023



Abstract:Recent success of large-scale Contrastive Language-Image Pre-training (CLIP) has led to great promise in zero-shot semantic segmentation by transferring image-text aligned knowledge to pixel-level classification. However, existing methods usually require an additional image encoder or retraining/tuning the CLIP module. Here, we present a cost-effective strategy using text-prompt learning that keeps the entire CLIP module frozen while fully leveraging its rich information. Specifically, we propose a novel Zero-shot segmentation with Optimal Transport (ZegOT) method that matches multiple text prompts with frozen image embeddings through optimal transport, which allows each text prompt to efficiently focus on specific semantic attributes. Additionally, we propose Deep Local Feature Alignment (DLFA) that deeply aligns the text prompts with intermediate local feature of the frozen image encoder layers, which significantly boosts the zero-shot segmentation performance. Through extensive experiments on benchmark datasets, we show that our method achieves the state-of-the-art (SOTA) performance with only x7 lighter parameters compared to previous SOTA approaches.
Diffusion Adversarial Representation Learning for Self-supervised Vessel Segmentation
Sep 29, 2022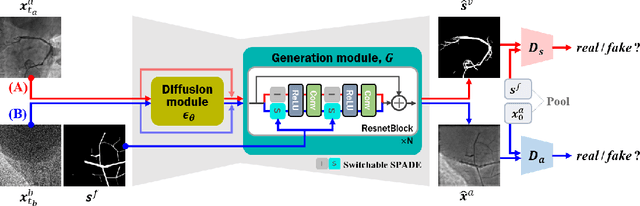
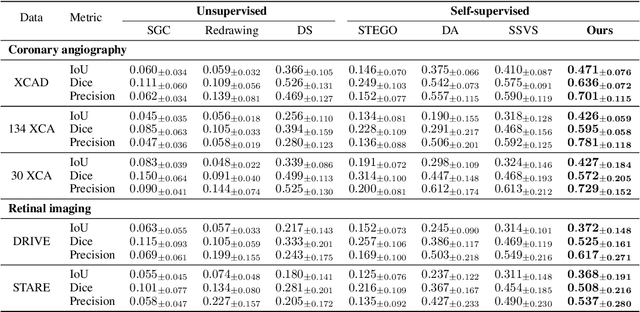
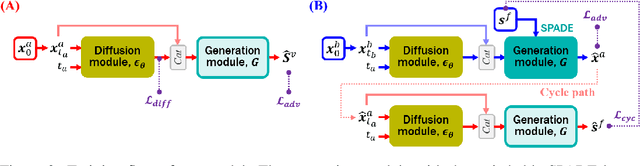
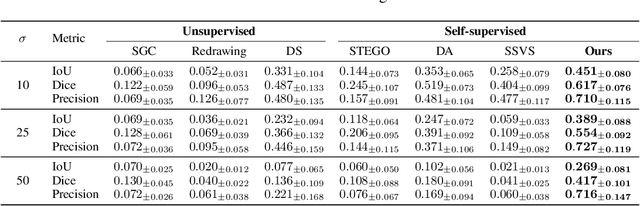
Abstract:Vessel segmentation in medical images is one of the important tasks in the diagnosis of vascular diseases and therapy planning. Although learning-based segmentation approaches have been extensively studied, a large amount of ground-truth labels are required in supervised methods and confusing background structures make neural networks hard to segment vessels in an unsupervised manner. To address this, here we introduce a novel diffusion adversarial representation learning (DARL) model that leverages a denoising diffusion probabilistic model with adversarial learning, and apply it for vessel segmentation. In particular, for self-supervised vessel segmentation, DARL learns background image distribution using a diffusion module, which lets a generation module effectively provide vessel representations. Also, by adversarial learning based on the proposed switchable spatially-adaptive denormalization, our model estimates synthetic fake vessel images as well as vessel segmentation masks, which further makes the model capture vessel-relevant semantic information. Once the proposed model is trained, the model generates segmentation masks by one step and can be applied to general vascular structure segmentation of coronary angiography and retinal images. Experimental results on various datasets show that our method significantly outperforms existing unsupervised and self-supervised methods in vessel segmentation.
A hybrid 2-stage vision transformer for AI-assisted 5 class pathologic diagnosis of gastric endoscopic biopsies
Feb 17, 2022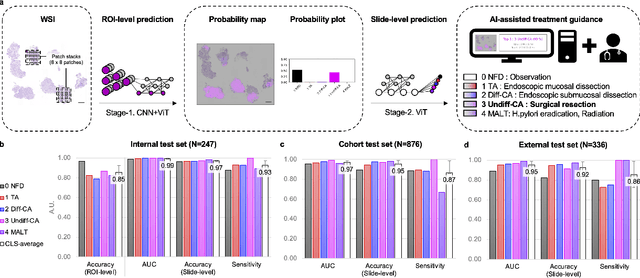
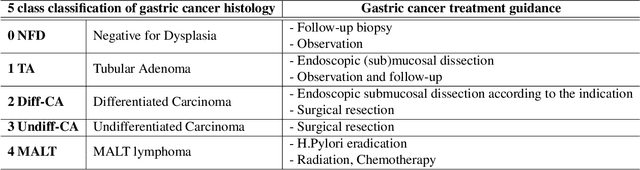
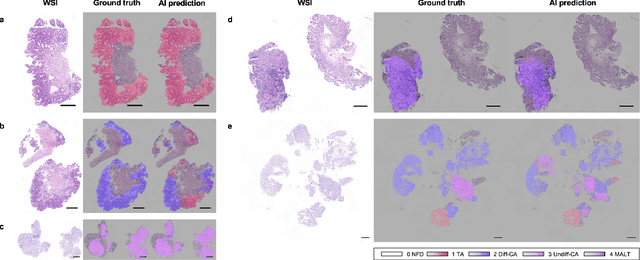
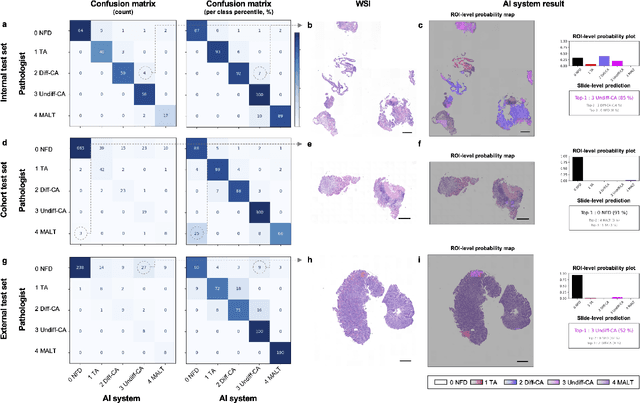
Abstract:Gastric endoscopic screening is an effective way to decide appropriate gastric cancer (GC) treatment at an early stage, reducing GC-associated mortality rate. Although artificial intelligence (AI) has brought a great promise to assist pathologist to screen digitalized whole slide images, automatic classification systems for guiding proper GC treatment based on clinical guideline are still lacking. Here, we propose an AI system classifying 5 classes of GC histology, which can be perfectly matched to general treatment guidance. The AI system, mimicking the way pathologist understand slides through multi-scale self-attention mechanism using a 2-stage Vision Transformer, demonstrates clinical capability by achieving diagnostic sensitivity of above 85% for both internal and external cohort analysis. Furthermore, AI-assisted pathologists showed significantly improved diagnostic sensitivity by 10% within 18% saved screening time compared to human pathologists. Our AI system has a great potential for providing presumptive pathologic opinion for deciding proper treatment for early GC patients.
AI can evolve without labels: self-evolving vision transformer for chest X-ray diagnosis through knowledge distillation
Feb 13, 2022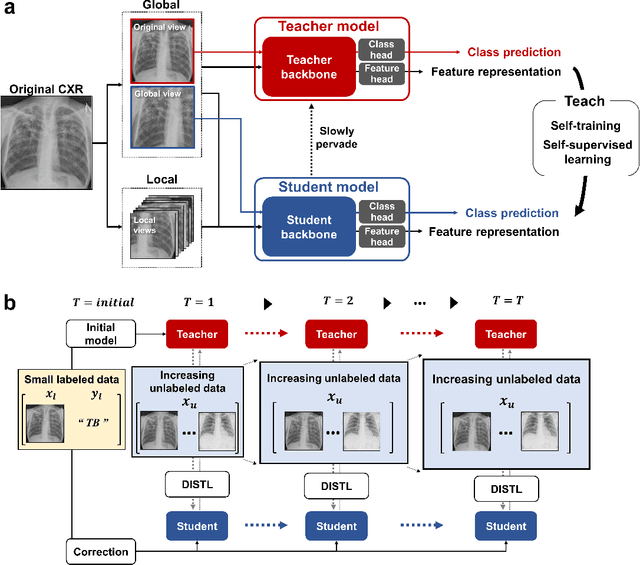
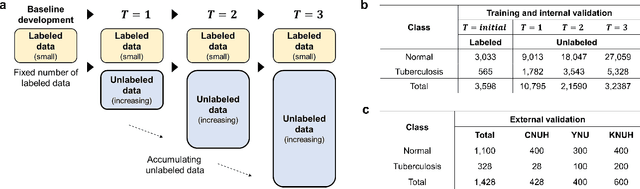
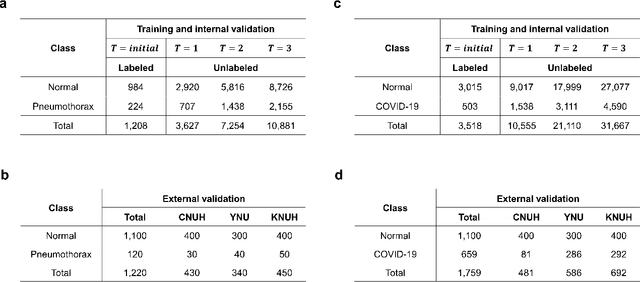
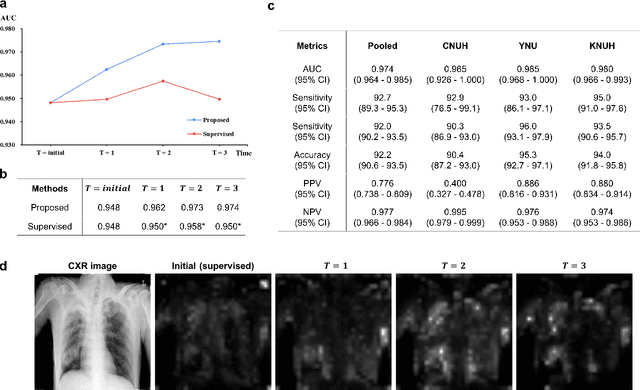
Abstract:Although deep learning-based computer-aided diagnosis systems have recently achieved expert-level performance, developing a robust deep learning model requires large, high-quality data with manual annotation, which is expensive to obtain. This situation poses the problem that the chest x-rays collected annually in hospitals cannot be used due to the lack of manual labeling by experts, especially in deprived areas. To address this, here we present a novel deep learning framework that uses knowledge distillation through self-supervised learning and self-training, which shows that the performance of the original model trained with a small number of labels can be gradually improved with more unlabeled data. Experimental results show that the proposed framework maintains impressive robustness against a real-world environment and has general applicability to several diagnostic tasks such as tuberculosis, pneumothorax, and COVID-19. Notably, we demonstrated that our model performs even better than those trained with the same amount of labeled data. The proposed framework has a great potential for medical imaging, where plenty of data is accumulated every year, but ground truth annotations are expensive to obtain.
Unifying domain adaptation and self-supervised learning for CXR segmentation via AdaIN-based knowledge distillation
Apr 23, 2021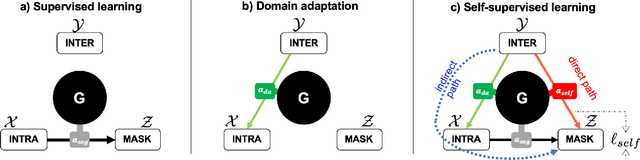
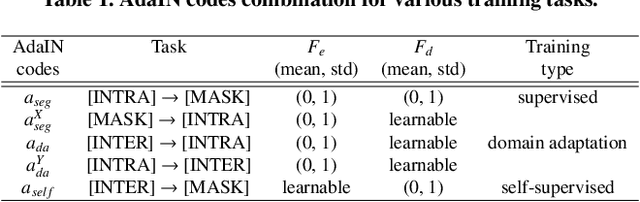
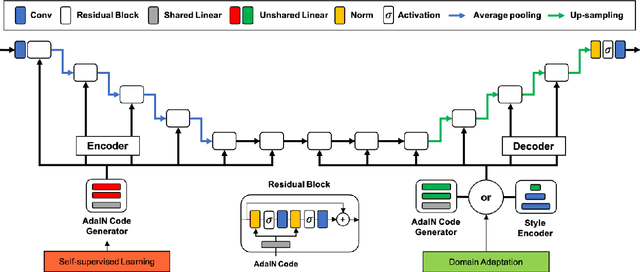
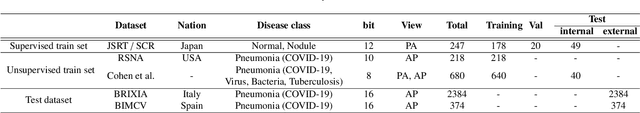
Abstract:As the segmentation labels are scarce, extensive researches have been conducted to train segmentation networks without labels or with only limited labels. In particular, domain adaptation, self-supervised learning, and teacher-student architecture have been introduced to distill knowledge from various tasks to improve the segmentation performance. However, these approaches appear different from each other, so it is not clear how these seemingly different approaches can be combined for better performance. Inspired by the recent StarGANv2 for multi-domain image translation, here we propose a novel segmentation framework via AdaIN-based knowledge distillation, where a single generator with AdaIN layers is trained along with the AdaIN code generator and style encoder so that the generator can perform both domain adaptation and segmentation. Specifically, our framework is designed to deal with difficult situations in chest X-ray (CXR) segmentation tasks where segmentation masks are only available for normal CXR data, but the trained model should be applied for both normal and abnormal CXR images. Since a single generator is used for abnormal to normal domain conversion and segmentation by simply changing the AdaIN codes, the generator can synergistically learn the common features to improve segmentation performance. Experimental results using CXR data confirm that the trained network can achieve the state-of-the art segmentation performance for both normal and abnormal CXR images.
Vision Transformer using Low-level Chest X-ray Feature Corpus for COVID-19 Diagnosis and Severity Quantification
Apr 15, 2021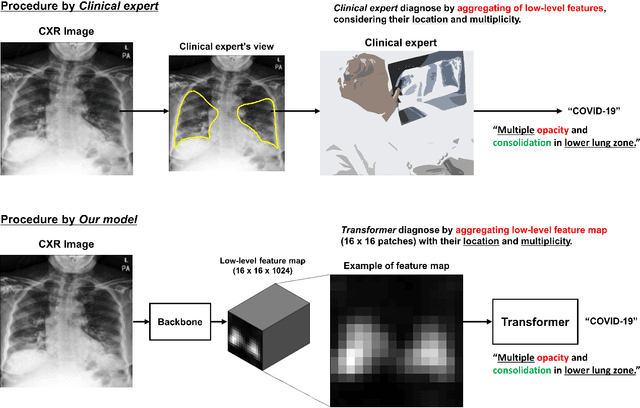
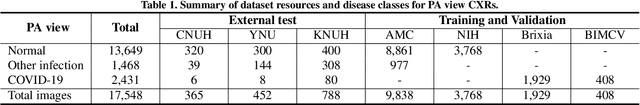
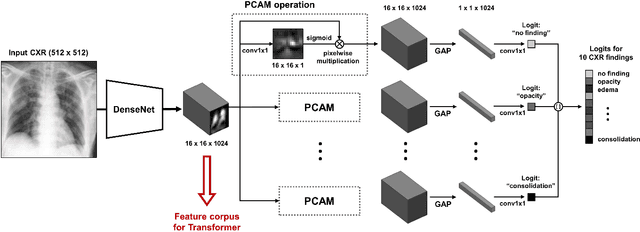
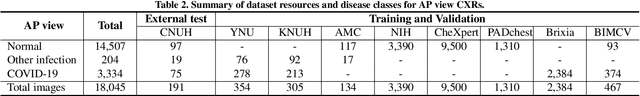
Abstract:Developing a robust algorithm to diagnose and quantify the severity of COVID-19 using Chest X-ray (CXR) requires a large number of well-curated COVID-19 datasets, which is difficult to collect under the global COVID-19 pandemic. On the other hand, CXR data with other findings are abundant. This situation is ideally suited for the Vision Transformer (ViT) architecture, where a lot of unlabeled data can be used through structural modeling by the self-attention mechanism. However, the use of existing ViT is not optimal, since feature embedding through direct patch flattening or ResNet backbone in the standard ViT is not intended for CXR. To address this problem, here we propose a novel Vision Transformer that utilizes low-level CXR feature corpus obtained from a backbone network that extracts common CXR findings. Specifically, the backbone network is first trained with large public datasets to detect common abnormal findings such as consolidation, opacity, edema, etc. Then, the embedded features from the backbone network are used as corpora for a Transformer model for the diagnosis and the severity quantification of COVID-19. We evaluate our model on various external test datasets from totally different institutions to evaluate the generalization capability. The experimental results confirm that our model can achieve the state-of-the-art performance in both diagnosis and severity quantification tasks with superior generalization capability, which are sine qua non of widespread deployment.
Severity Quantification and Lesion Localization of COVID-19 on CXR using Vision Transformer
Mar 12, 2021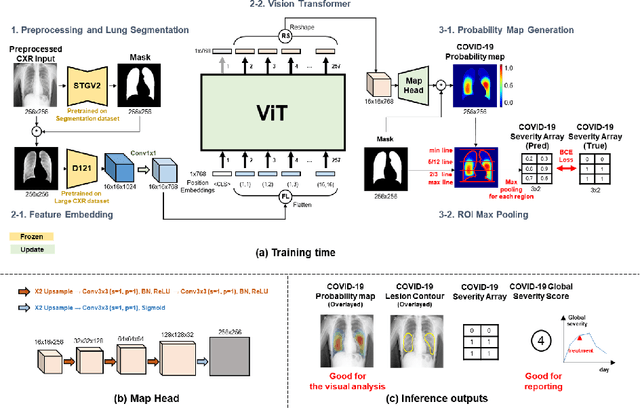
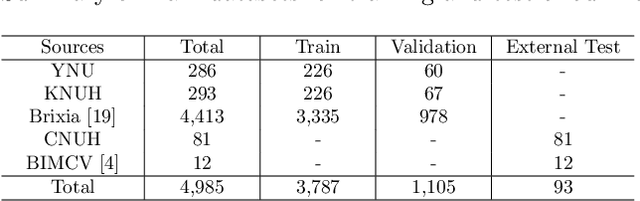
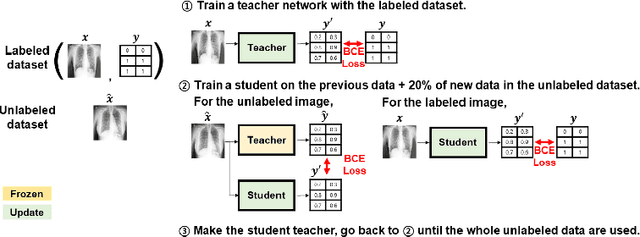
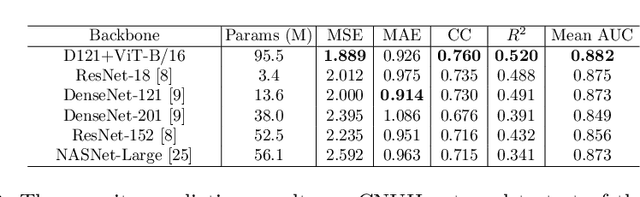
Abstract:Under the global pandemic of COVID-19, building an automated framework that quantifies the severity of COVID-19 and localizes the relevant lesion on chest X-ray images has become increasingly important. Although pixel-level lesion severity labels, e.g. lesion segmentation, can be the most excellent target to build a robust model, collecting enough data with such labels is difficult due to time and labor-intensive annotation tasks. Instead, array-based severity labeling that assigns integer scores on six subdivisions of lungs can be an alternative choice enabling the quick labeling. Several groups proposed deep learning algorithms that quantify the severity of COVID-19 using the array-based COVID-19 labels and localize the lesions with explainability maps. To further improve the accuracy and interpretability, here we propose a novel Vision Transformer tailored for both quantification of the severity and clinically applicable localization of the COVID-19 related lesions. Our model is trained in a weakly-supervised manner to generate the full probability maps from weak array-based labels. Furthermore, a novel progressive self-training method enables us to build a model with a small labeled dataset. The quantitative and qualitative analysis on the external testset demonstrates that our method shows comparable performance with radiologists for both tasks with stability in a real-world application.
Vision Transformer for COVID-19 CXR Diagnosis using Chest X-ray Feature Corpus
Mar 12, 2021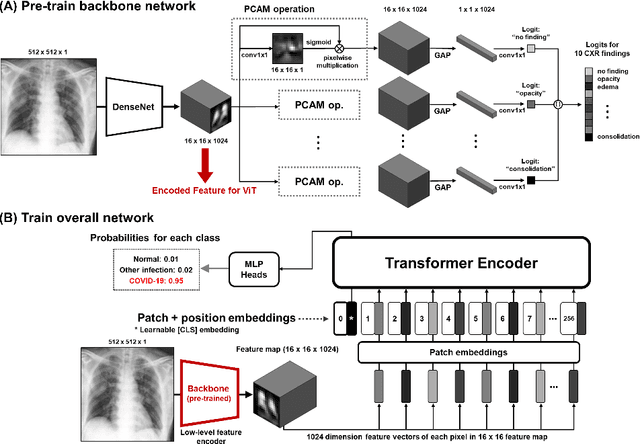
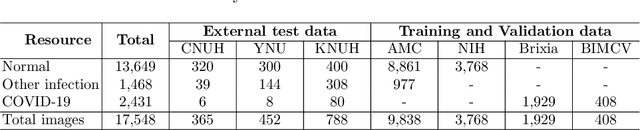
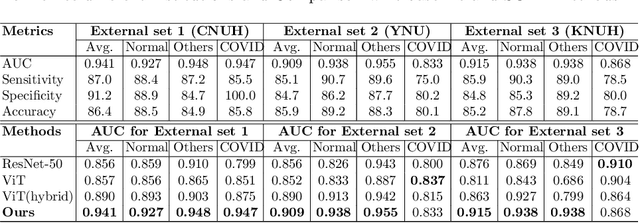
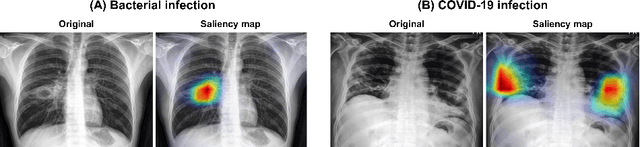
Abstract:Under the global COVID-19 crisis, developing robust diagnosis algorithm for COVID-19 using CXR is hampered by the lack of the well-curated COVID-19 data set, although CXR data with other disease are abundant. This situation is suitable for vision transformer architecture that can exploit the abundant unlabeled data using pre-training. However, the direct use of existing vision transformer that uses the corpus generated by the ResNet is not optimal for correct feature embedding. To mitigate this problem, we propose a novel vision Transformer by using the low-level CXR feature corpus that are obtained to extract the abnormal CXR features. Specifically, the backbone network is trained using large public datasets to obtain the abnormal features in routine diagnosis such as consolidation, glass-grass opacity (GGO), etc. Then, the embedded features from the backbone network are used as corpus for vision transformer training. We examine our model on various external test datasets acquired from totally different institutions to assess the generalization ability. Our experiments demonstrate that our method achieved the state-of-art performance and has better generalization capability, which are crucial for a widespread deployment.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge