Joon Beom Seo
AI can evolve without labels: self-evolving vision transformer for chest X-ray diagnosis through knowledge distillation
Feb 13, 2022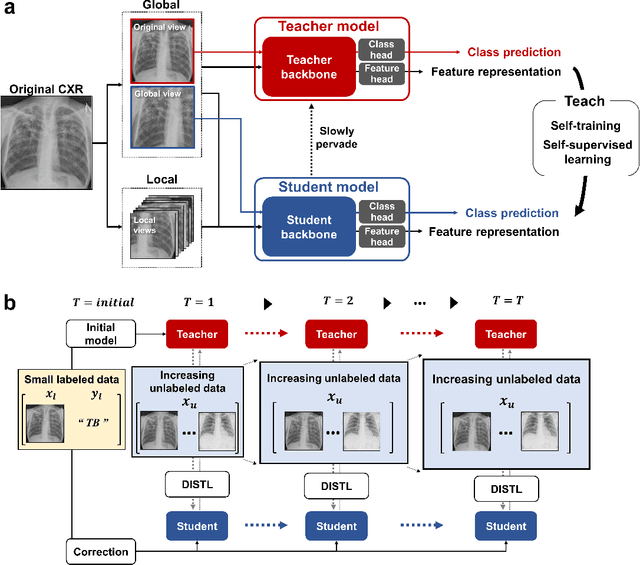
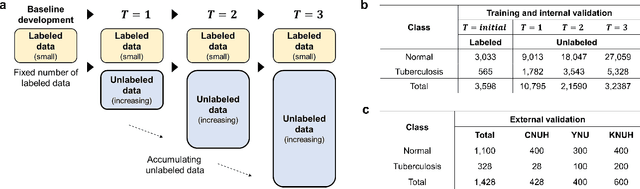
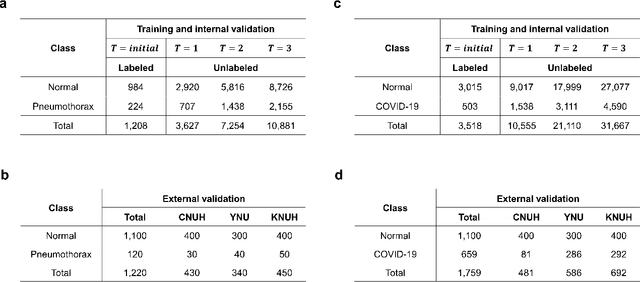
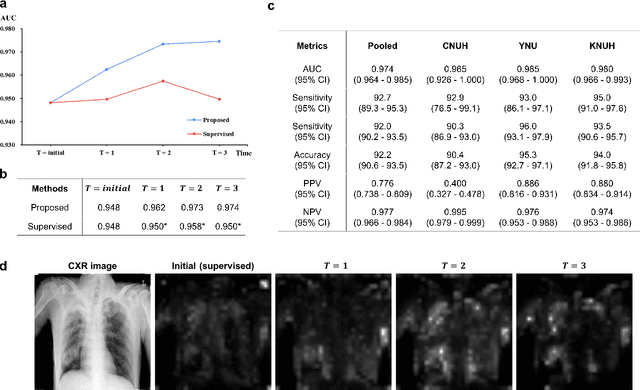
Abstract:Although deep learning-based computer-aided diagnosis systems have recently achieved expert-level performance, developing a robust deep learning model requires large, high-quality data with manual annotation, which is expensive to obtain. This situation poses the problem that the chest x-rays collected annually in hospitals cannot be used due to the lack of manual labeling by experts, especially in deprived areas. To address this, here we present a novel deep learning framework that uses knowledge distillation through self-supervised learning and self-training, which shows that the performance of the original model trained with a small number of labels can be gradually improved with more unlabeled data. Experimental results show that the proposed framework maintains impressive robustness against a real-world environment and has general applicability to several diagnostic tasks such as tuberculosis, pneumothorax, and COVID-19. Notably, we demonstrated that our model performs even better than those trained with the same amount of labeled data. The proposed framework has a great potential for medical imaging, where plenty of data is accumulated every year, but ground truth annotations are expensive to obtain.
Vision Transformer using Low-level Chest X-ray Feature Corpus for COVID-19 Diagnosis and Severity Quantification
Apr 15, 2021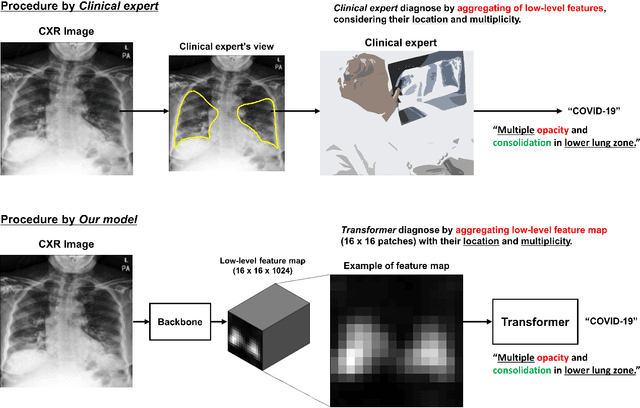
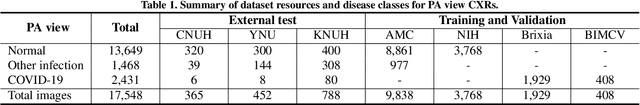
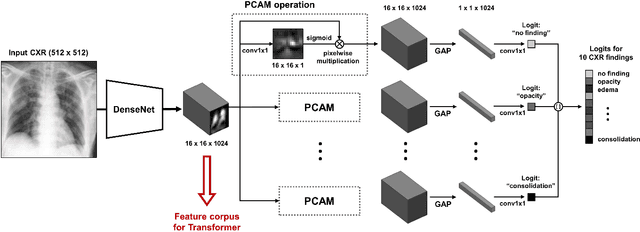
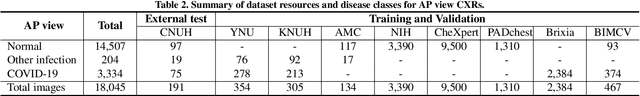
Abstract:Developing a robust algorithm to diagnose and quantify the severity of COVID-19 using Chest X-ray (CXR) requires a large number of well-curated COVID-19 datasets, which is difficult to collect under the global COVID-19 pandemic. On the other hand, CXR data with other findings are abundant. This situation is ideally suited for the Vision Transformer (ViT) architecture, where a lot of unlabeled data can be used through structural modeling by the self-attention mechanism. However, the use of existing ViT is not optimal, since feature embedding through direct patch flattening or ResNet backbone in the standard ViT is not intended for CXR. To address this problem, here we propose a novel Vision Transformer that utilizes low-level CXR feature corpus obtained from a backbone network that extracts common CXR findings. Specifically, the backbone network is first trained with large public datasets to detect common abnormal findings such as consolidation, opacity, edema, etc. Then, the embedded features from the backbone network are used as corpora for a Transformer model for the diagnosis and the severity quantification of COVID-19. We evaluate our model on various external test datasets from totally different institutions to evaluate the generalization capability. The experimental results confirm that our model can achieve the state-of-the-art performance in both diagnosis and severity quantification tasks with superior generalization capability, which are sine qua non of widespread deployment.
Severity Quantification and Lesion Localization of COVID-19 on CXR using Vision Transformer
Mar 12, 2021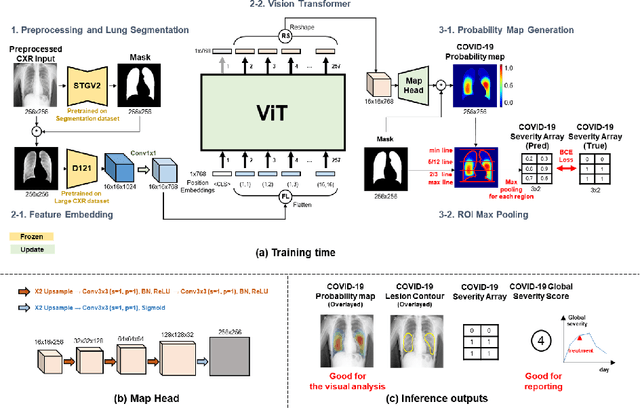
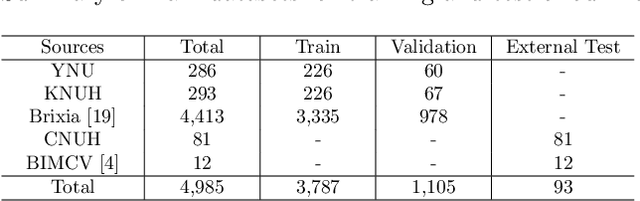
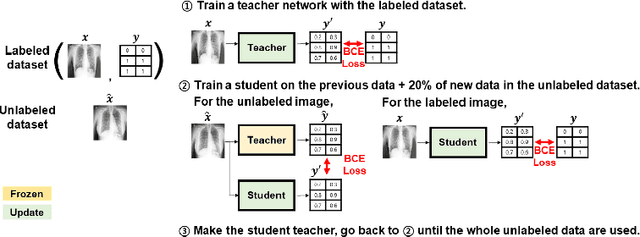
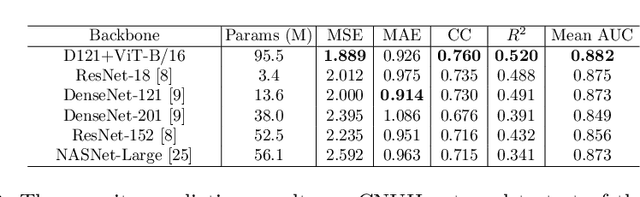
Abstract:Under the global pandemic of COVID-19, building an automated framework that quantifies the severity of COVID-19 and localizes the relevant lesion on chest X-ray images has become increasingly important. Although pixel-level lesion severity labels, e.g. lesion segmentation, can be the most excellent target to build a robust model, collecting enough data with such labels is difficult due to time and labor-intensive annotation tasks. Instead, array-based severity labeling that assigns integer scores on six subdivisions of lungs can be an alternative choice enabling the quick labeling. Several groups proposed deep learning algorithms that quantify the severity of COVID-19 using the array-based COVID-19 labels and localize the lesions with explainability maps. To further improve the accuracy and interpretability, here we propose a novel Vision Transformer tailored for both quantification of the severity and clinically applicable localization of the COVID-19 related lesions. Our model is trained in a weakly-supervised manner to generate the full probability maps from weak array-based labels. Furthermore, a novel progressive self-training method enables us to build a model with a small labeled dataset. The quantitative and qualitative analysis on the external testset demonstrates that our method shows comparable performance with radiologists for both tasks with stability in a real-world application.
Vision Transformer for COVID-19 CXR Diagnosis using Chest X-ray Feature Corpus
Mar 12, 2021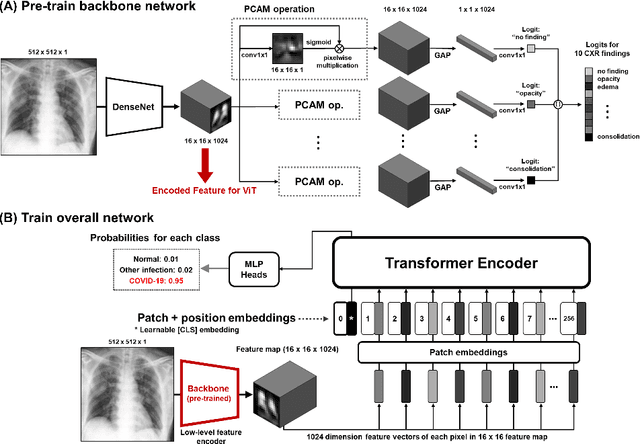
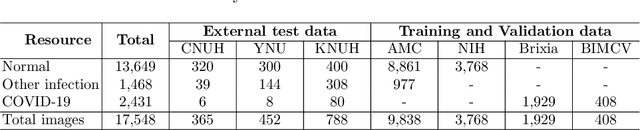
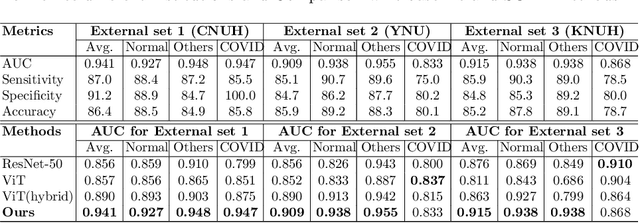
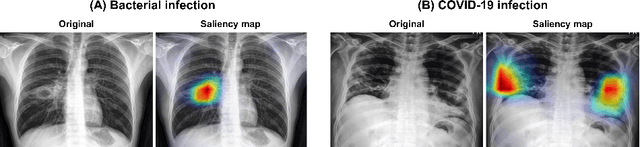
Abstract:Under the global COVID-19 crisis, developing robust diagnosis algorithm for COVID-19 using CXR is hampered by the lack of the well-curated COVID-19 data set, although CXR data with other disease are abundant. This situation is suitable for vision transformer architecture that can exploit the abundant unlabeled data using pre-training. However, the direct use of existing vision transformer that uses the corpus generated by the ResNet is not optimal for correct feature embedding. To mitigate this problem, we propose a novel vision Transformer by using the low-level CXR feature corpus that are obtained to extract the abnormal CXR features. Specifically, the backbone network is trained using large public datasets to obtain the abnormal features in routine diagnosis such as consolidation, glass-grass opacity (GGO), etc. Then, the embedded features from the backbone network are used as corpus for vision transformer training. We examine our model on various external test datasets acquired from totally different institutions to assess the generalization ability. Our experiments demonstrate that our method achieved the state-of-art performance and has better generalization capability, which are crucial for a widespread deployment.
False Positive Reduction by Actively Mining Negative Samples for Pulmonary Nodule Detection in Chest Radiographs
Jul 26, 2018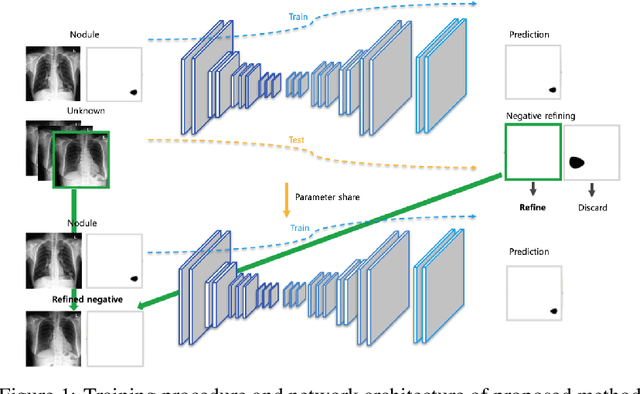
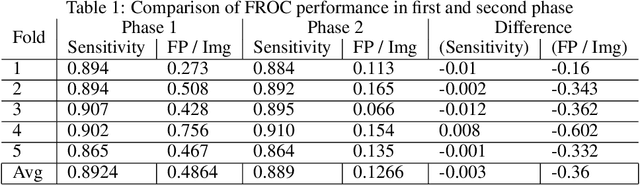
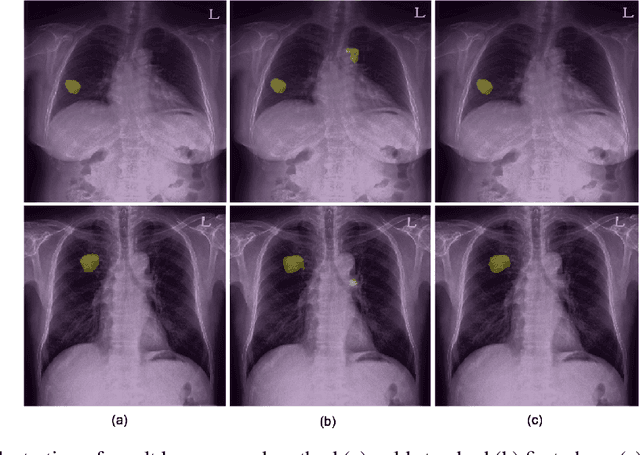
Abstract:Generating large quantities of quality labeled data in medical imaging is very time consuming and expensive. The performance of supervised algorithms for various tasks on imaging has improved drastically over the years, however the availability of data to train these algorithms have become one of the main bottlenecks for implementation. To address this, we propose a semi-supervised learning method where pseudo-negative labels from unlabeled data are used to further refine the performance of a pulmonary nodule detection network in chest radiographs. After training with the proposed network, the false positive rate was reduced to 0.1266 from 0.4864 while maintaining sensitivity at 0.89.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge