Yonggang Shi
NeuroAgent: LLM Agents for Multimodal Neuroimaging Analysis and Research
May 07, 2026Abstract:Multimodal neuroimaging analysis often involves complex, modality-specific preprocessing workflows that require careful configuration, quality control, and coordination across heterogeneous toolchains. Beyond preprocessing, downstream statistical analysis and disease classification commonly require task-specific code, evaluation protocols, and data-format conventions, creating additional barriers between raw acquisitions and reproducible scientific analysis. We present NeuroAgent, an LLM-driven agentic framework that automates key preprocessing and analysis steps for heterogeneous neuroimaging data, including sMRI, fMRI, dMRI, and PET, and supports interactive downstream analysis through natural-language queries. NeuroAgent employs a hierarchical multi-agent architecture with a feedback-driven Generate-Execute-Validate engine: agents autonomously generate executable preprocessing code, detect and recover from runtime errors, and validate output integrity. We evaluate the system on 1,470 subjects pooled across all ADNI phases (CN=1,000, AD=470), where all subjects have sMRI and tabular data, with subsets also having Tau-PET (n=469), fMRI (n=278), and DTI ($n=620$). Pipeline ablation studies across multiple LLM backends show that capable models reach up to 100% intent-parsing accuracy, with the strongest backend (Qwen3.5-27B) reaching 84.8% end-to-end preprocessing step correctness. Automated recovery limits manual intervention to edge cases where human review is required via the Human-In-The-Loop interface. For Alzheimer's Disease classification using automatically preprocessed multimodal data, our agent ensemble achieves an AUC of 0.9518 with four modalities, outperforming all single-modality baselines. These results show that NeuroAgent can reduce the manual effort required for neuroimaging preprocessing and enable end-to-end automated analysis pipelines for neuroimaging research.
ssProp: Energy-Efficient Training for Convolutional Neural Networks with Scheduled Sparse Back Propagation
Aug 22, 2024

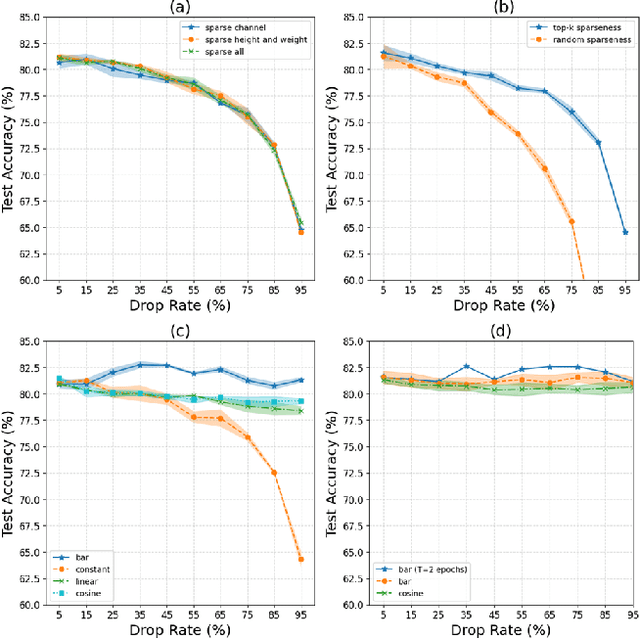

Abstract:Recently, deep learning has made remarkable strides, especially with generative modeling, such as large language models and probabilistic diffusion models. However, training these models often involves significant computational resources, requiring billions of petaFLOPs. This high resource consumption results in substantial energy usage and a large carbon footprint, raising critical environmental concerns. Back-propagation (BP) is a major source of computational expense during training deep learning models. To advance research on energy-efficient training and allow for sparse learning on any machine and device, we propose a general, energy-efficient convolution module that can be seamlessly integrated into any deep learning architecture. Specifically, we introduce channel-wise sparsity with additional gradient selection schedulers during backward based on the assumption that BP is often dense and inefficient, which can lead to over-fitting and high computational consumption. Our experiments demonstrate that our approach reduces 40\% computations while potentially improving model performance, validated on image classification and generation tasks. This reduction can lead to significant energy savings and a lower carbon footprint during the research and development phases of large-scale AI systems. Additionally, our method mitigates over-fitting in a manner distinct from Dropout, allowing it to be combined with Dropout to further enhance model performance and reduce computational resource usage. Extensive experiments validate that our method generalizes to a variety of datasets and tasks and is compatible with a wide range of deep learning architectures and modules. Code is publicly available at https://github.com/lujiazho/ssProp.
Diffusion Model-based FOD Restoration from High Distortion in dMRI
Jun 19, 2024



Abstract:Fiber orientation distributions (FODs) is a popular model to represent the diffusion MRI (dMRI) data. However, imaging artifacts such as susceptibility-induced distortion in dMRI can cause signal loss and lead to the corrupted reconstruction of FODs, which prohibits successful fiber tracking and connectivity analysis in affected brain regions such as the brain stem. Generative models, such as the diffusion models, have been successfully applied in various image restoration tasks. However, their application on FOD images poses unique challenges since FODs are 4-dimensional data represented by spherical harmonics (SPHARM) with the 4-th dimension exhibiting order-related dependency. In this paper, we propose a novel diffusion model for FOD restoration that can recover the signal loss caused by distortion artifacts. We use volume-order encoding to enhance the ability of the diffusion model to generate individual FOD volumes at all SPHARM orders. Moreover, we add cross-attention features extracted across all SPHARM orders in generating every individual FOD volume to capture the order-related dependency across FOD volumes. We also condition the diffusion model with low-distortion FODs surrounding high-distortion areas to maintain the geometric coherence of the generated FODs. We trained and tested our model using data from the UK Biobank (n = 1315). On a test set with ground truth (n = 43), we demonstrate the high accuracy of the generated FODs in terms of root mean square errors of FOD volumes and angular errors of FOD peaks. We also apply our method to a test set with large distortion in the brain stem area (n = 1172) and demonstrate the efficacy of our method in restoring the FOD integrity and, hence, greatly improving tractography performance in affected brain regions.
TauAD: MRI-free Tau Anomaly Detection in PET Imaging via Conditioned Diffusion Models
May 21, 2024



Abstract:The emergence of tau PET imaging over the last decade has enabled Alzheimer's disease (AD) researchers to examine tau pathology in vivo and more effectively characterize the disease trajectories of AD. Current tau PET analysis methods, however, typically perform inferences on large cortical ROIs and are limited in the detection of localized tau pathology that varies across subjects. Furthermore, a high-resolution MRI is required to carry out conventional tau PET analysis, which is not commonly acquired in clinical practices and may not be acquired for many elderly patients with dementia due to strong motion artifacts, claustrophobia, or certain metal implants. In this work, we propose a novel conditional diffusion model to perform MRI-free anomaly detection from tau PET imaging data. By including individualized conditions and two complementary loss maps from pseudo-healthy and pseudo-unhealthy reconstructions, our model computes an anomaly map across the entire brain area that allows simply training a support vector machine (SVM) for classifying disease severity. We train our model on ADNI subjects (n=534) and evaluate its performance on a separate dataset from the preclinical subjects of the A4 clinical trial (n=447). We demonstrate that our method outperforms baseline generative models and the conventional Z-score-based method in anomaly localization without mis-detecting off-target bindings in sub-cortical and out-of-brain areas. By classifying the A4 subjects according to their anomaly map using the SVM trained on ADNI data, we show that our method can successfully group preclinical subjects with significantly different cognitive functions, which further demonstrates the effectiveness of our method in capturing biologically relevant anomaly in tau PET imaging.
Anatomy-Guided Surface Diffusion Model for Alzheimer's Disease Normative Modeling
Mar 07, 2024



Abstract:Normative modeling has emerged as a pivotal approach for characterizing heterogeneity and individual variance in neurodegenerative diseases, notably Alzheimer's disease(AD). One of the challenges of cortical normative modeling is the anatomical structure mismatch due to folding pattern variability. Traditionally, registration is applied to address this issue and recently many studies have utilized deep generative models to generate anatomically align samples for analyzing disease progression; however, these models are predominantly applied to volume-based data, which often falls short in capturing intricate morphological changes on the brain cortex. As an alternative, surface-based analysis has been proven to be more sensitive in disease modeling such as AD, yet, like volume-based data, it also suffers from the mismatch problem. To address these limitations, we proposed a novel generative normative modeling framework by transferring the conditional diffusion generative model to the spherical non-Euclidean domain. Additionally, this approach generates normal feature map distributions by explicitly conditioning on individual anatomical segmentation to ensure better geometrical alignment which helps to reduce anatomical variance between subjects in analysis. We find that our model can generate samples that are better anatomically aligned than registered reference data and through ablation study and normative assessment experiments, the samples are able to better measure individual differences from the normal distribution and increase sensitivity in differentiating cognitively normal (CN), mild cognitive impairment (MCI), and Alzheimer's disease (AD) patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge