Yikuan Li
Hi-BEHRT: Hierarchical Transformer-based model for accurate prediction of clinical events using multimodal longitudinal electronic health records
Jun 21, 2021
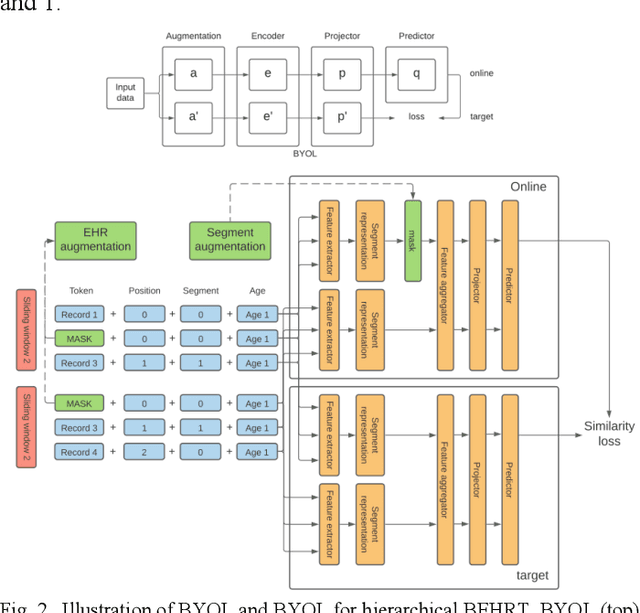
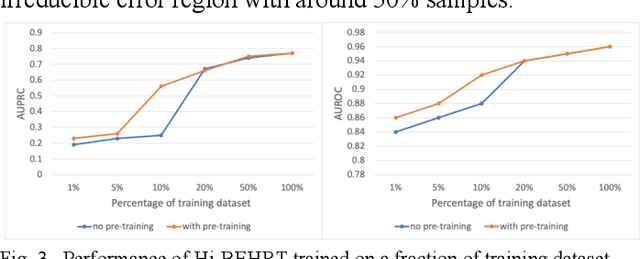
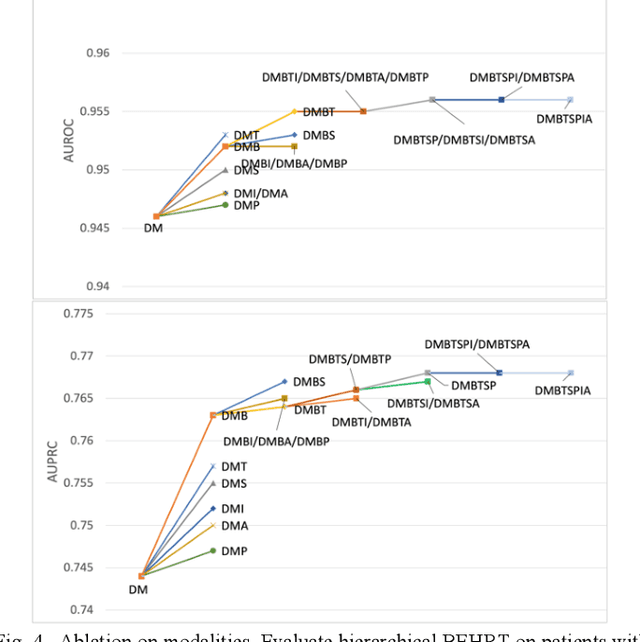
Abstract:Electronic health records represent a holistic overview of patients' trajectories. Their increasing availability has fueled new hopes to leverage them and develop accurate risk prediction models for a wide range of diseases. Given the complex interrelationships of medical records and patient outcomes, deep learning models have shown clear merits in achieving this goal. However, a key limitation of these models remains their capacity in processing long sequences. Capturing the whole history of medical encounters is expected to lead to more accurate predictions, but the inclusion of records collected for decades and from multiple resources can inevitably exceed the receptive field of the existing deep learning architectures. This can result in missing crucial, long-term dependencies. To address this gap, we present Hi-BEHRT, a hierarchical Transformer-based model that can significantly expand the receptive field of Transformers and extract associations from much longer sequences. Using a multimodal large-scale linked longitudinal electronic health records, the Hi-BEHRT exceeds the state-of-the-art BEHRT 1% to 5% for area under the receiver operating characteristic (AUROC) curve and 3% to 6% for area under the precision recall (AUPRC) curve on average, and 3% to 6% (AUROC) and 3% to 11% (AUPRC) for patients with long medical history for 5-year heart failure, diabetes, chronic kidney disease, and stroke risk prediction. Additionally, because pretraining for hierarchical Transformer is not well-established, we provide an effective end-to-end contrastive pre-training strategy for Hi-BEHRT using EHR, improving its transferability on predicting clinical events with relatively small training dataset.
Risk factor identification for incident heart failure using neural network distillation and variable selection
Mar 01, 2021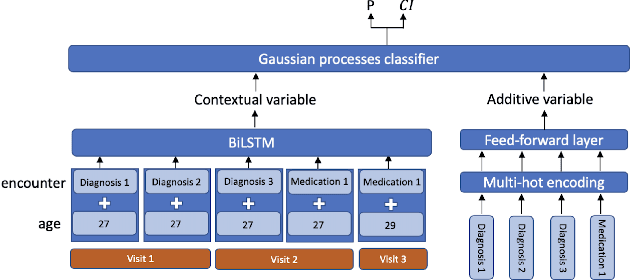
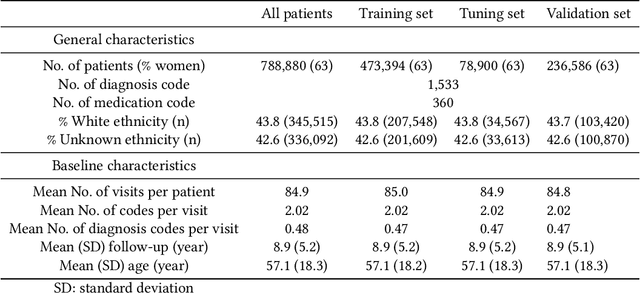
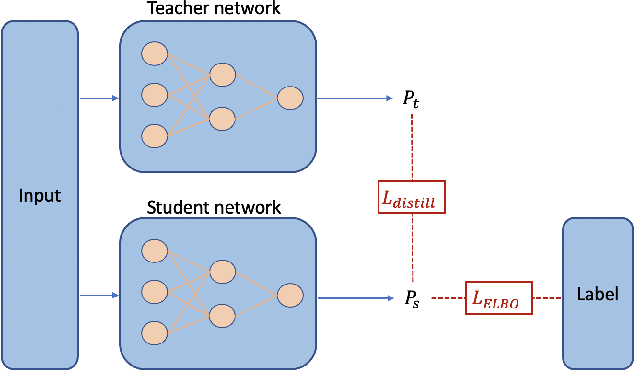
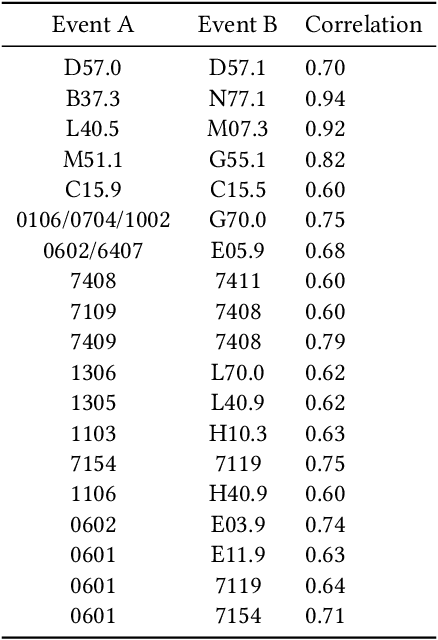
Abstract:Recent evidence shows that deep learning models trained on electronic health records from millions of patients can deliver substantially more accurate predictions of risk compared to their statistical counterparts. While this provides an important opportunity for improving clinical decision-making, the lack of interpretability is a major barrier to the incorporation of these black-box models in routine care, limiting their trustworthiness and preventing further hypothesis-testing investigations. In this study, we propose two methods, namely, model distillation and variable selection, to untangle hidden patterns learned by an established deep learning model (BEHRT) for risk association identification. Due to the clinical importance and diversity of heart failure as a phenotype, it was used to showcase the merits of the proposed methods. A cohort with 788,880 (8.3% incident heart failure) patients was considered for the study. Model distillation identified 598 and 379 diseases that were associated and dissociated with heart failure at the population level, respectively. While the associations were broadly consistent with prior knowledge, our method also highlighted several less appreciated links that are worth further investigation. In addition to these important population-level insights, we developed an approach to individual-level interpretation to take account of varying manifestation of heart failure in clinical practice. This was achieved through variable selection by detecting a minimal set of encounters that can maximally preserve the accuracy of prediction for individuals. Our proposed work provides a discovery-enabling tool to identify risk factors in both population and individual levels from a data-driven perspective. This helps to generate new hypotheses and guides further investigations on causal links.
An explainable Transformer-based deep learning model for the prediction of incident heart failure
Jan 27, 2021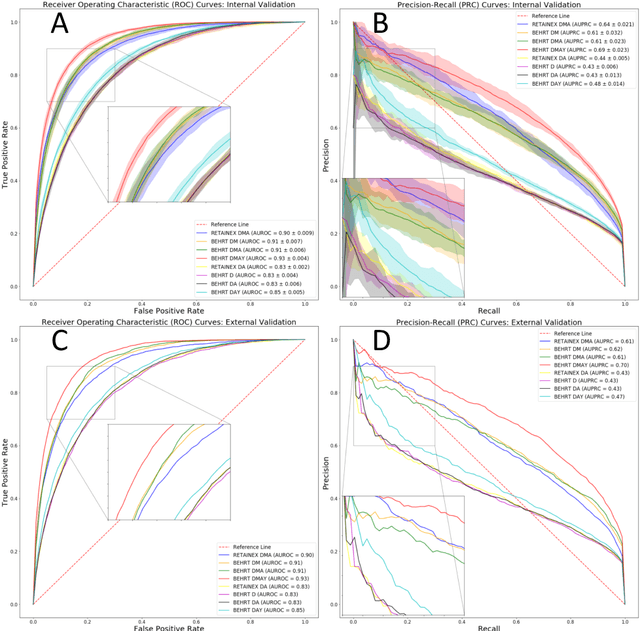
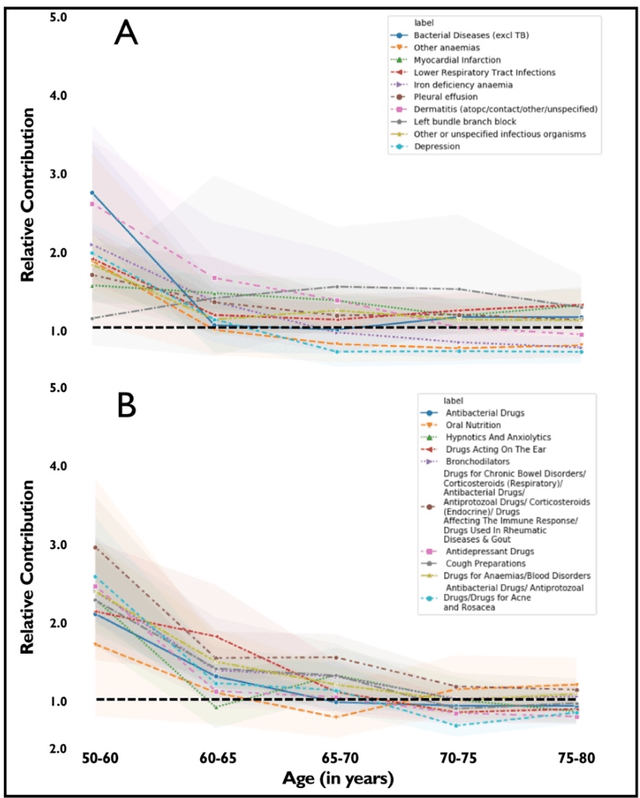
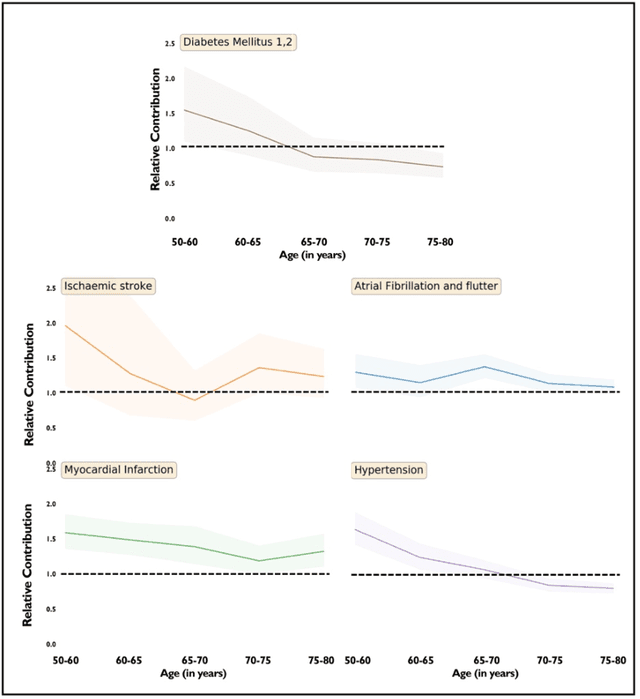
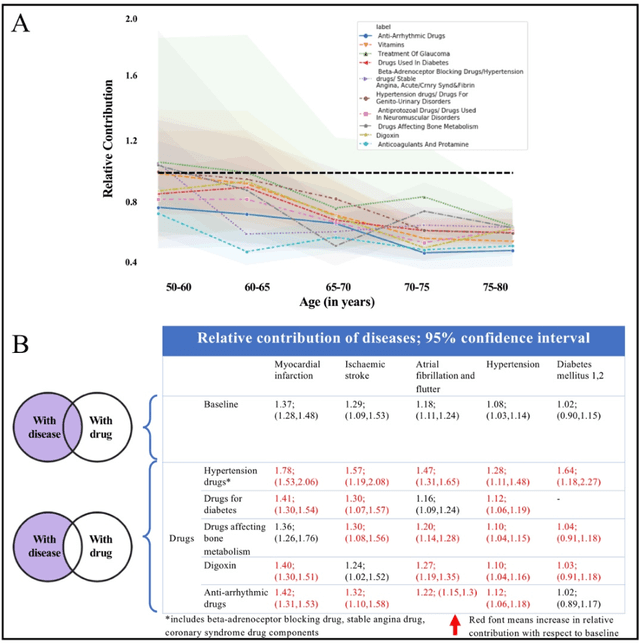
Abstract:Predicting the incidence of complex chronic conditions such as heart failure is challenging. Deep learning models applied to rich electronic health records may improve prediction but remain unexplainable hampering their wider use in medical practice. We developed a novel Transformer deep-learning model for more accurate and yet explainable prediction of incident heart failure involving 100,071 patients from longitudinal linked electronic health records across the UK. On internal 5-fold cross validation and held-out external validation, our model achieved 0.93 and 0.93 area under the receiver operator curve and 0.69 and 0.70 area under the precision-recall curve, respectively and outperformed existing deep learning models. Predictor groups included all community and hospital diagnoses and medications contextualised within the age and calendar year for each patient's clinical encounter. The importance of contextualised medical information was revealed in a number of sensitivity analyses, and our perturbation method provided a way of identifying factors contributing to risk. Many of the identified risk factors were consistent with existing knowledge from clinical and epidemiological research but several new associations were revealed which had not been considered in expert-driven risk prediction models.
A Comparison of Pre-trained Vision-and-Language Models for Multimodal Representation Learning across Medical Images and Reports
Sep 03, 2020



Abstract:Joint image-text embedding extracted from medical images and associated contextual reports is the bedrock for most biomedical vision-and-language (V+L) tasks, including medical visual question answering, clinical image-text retrieval, clinical report auto-generation. In this study, we adopt four pre-trained V+L models: LXMERT, VisualBERT, UNIER and PixelBERT to learn multimodal representation from MIMIC-CXR radiographs and associated reports. The extrinsic evaluation on OpenI dataset shows that in comparison to the pioneering CNN-RNN model, the joint embedding learned by pre-trained V+L models demonstrate performance improvement in the thoracic findings classification task. We conduct an ablation study to analyze the contribution of certain model components and validate the advantage of joint embedding over text-only embedding. We also visualize attention maps to illustrate the attention mechanism of V+L models.
Deep Bayesian Gaussian Processes for Uncertainty Estimation in Electronic Health Records
Mar 23, 2020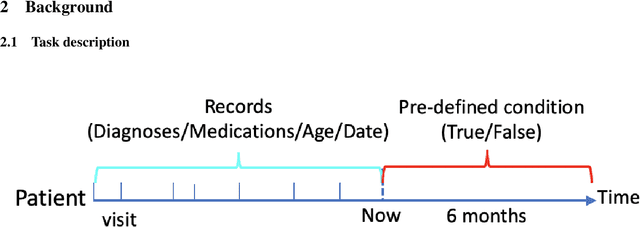
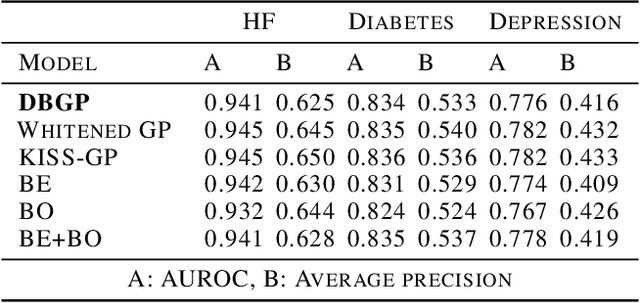
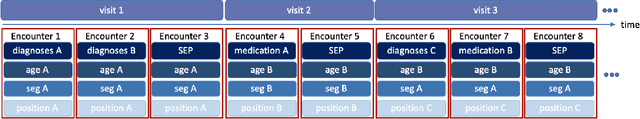
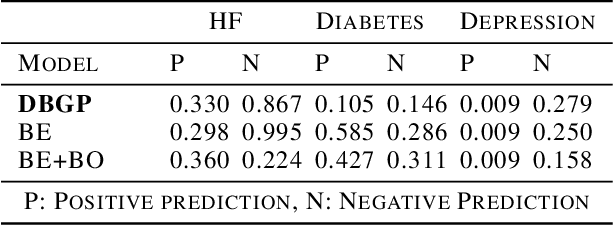
Abstract:One major impediment to the wider use of deep learning for clinical decision making is the difficulty of assigning a level of confidence to model predictions. Currently, deep Bayesian neural networks and sparse Gaussian processes are the main two scalable uncertainty estimation methods. However, deep Bayesian neural network suffers from lack of expressiveness, and more expressive models such as deep kernel learning, which is an extension of sparse Gaussian process, captures only the uncertainty from the higher level latent space. Therefore, the deep learning model under it lacks interpretability and ignores uncertainty from the raw data. In this paper, we merge features of the deep Bayesian learning framework with deep kernel learning to leverage the strengths of both methods for more comprehensive uncertainty estimation. Through a series of experiments on predicting the first incidence of heart failure, diabetes and depression applied to large-scale electronic medical records, we demonstrate that our method is better at capturing uncertainty than both Gaussian processes and deep Bayesian neural networks in terms of indicating data insufficiency and distinguishing true positive and false positive predictions, with a comparable generalisation performance. Furthermore, by assessing the accuracy and area under the receiver operating characteristic curve over the predictive probability, we show that our method is less susceptible to making overconfident predictions, especially for the minority class in imbalanced datasets. Finally, we demonstrate how uncertainty information derived by the model can inform risk factor analysis towards model interpretability.
BEHRT: Transformer for Electronic Health Records
Jul 22, 2019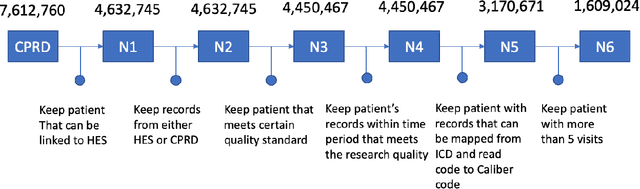

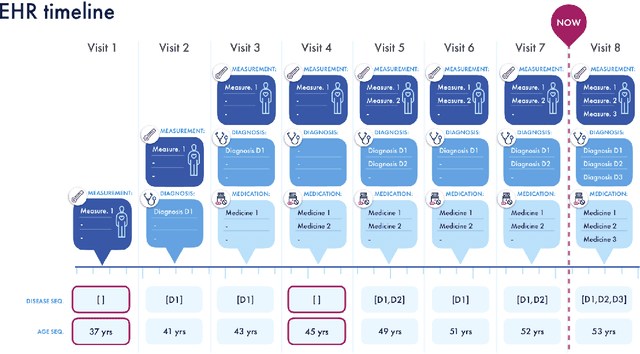
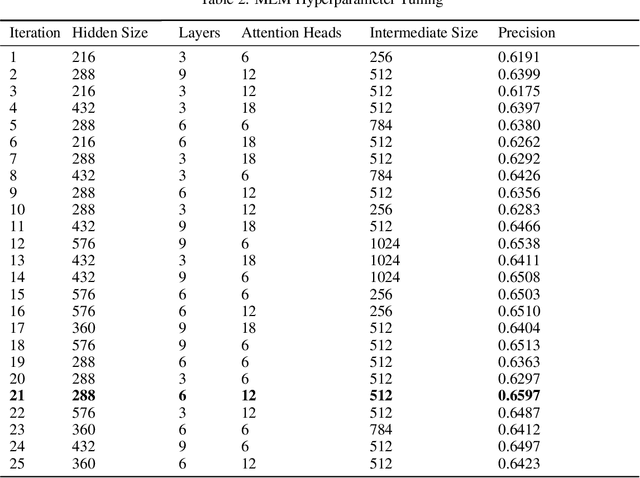
Abstract:Today, despite decades of developments in medicine and the growing interest in precision healthcare, vast majority of diagnoses happen once patients begin to show noticeable signs of illness. Early indication and detection of diseases, however, can provide patients and carers with the chance of early intervention, better disease management, and efficient allocation of healthcare resources. The latest developments in machine learning (more specifically, deep learning) provides a great opportunity to address this unmet need. In this study, we introduce BEHRT: A deep neural sequence transduction model for EHR (electronic health records), capable of multitask prediction and disease trajectory mapping. When trained and evaluated on the data from nearly 1.6 million individuals, BEHRT shows a striking absolute improvement of 8.0-10.8%, in terms of Average Precision Score, compared to the existing state-of-the-art deep EHR models (in terms of average precision, when predicting for the onset of 301 conditions). In addition to its superior prediction power, BEHRT provides a personalised view of disease trajectories through its attention mechanism; its flexible architecture enables it to incorporate multiple heterogeneous concepts (e.g., diagnosis, medication, measurements, and more) to improve the accuracy of its predictions; and its (pre-)training results in disease and patient representations that can help us get a step closer to interpretable predictions.
Learning Multimorbidity Patterns from Electronic Health Records Using Non-negative Matrix Factorisation
Jul 19, 2019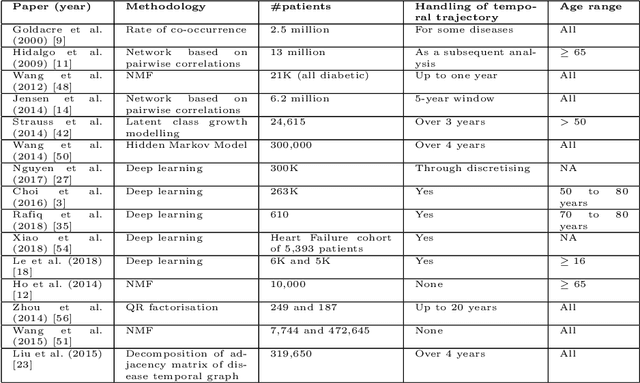
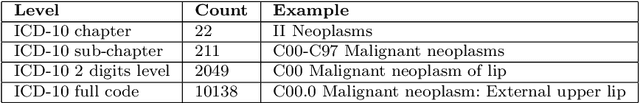
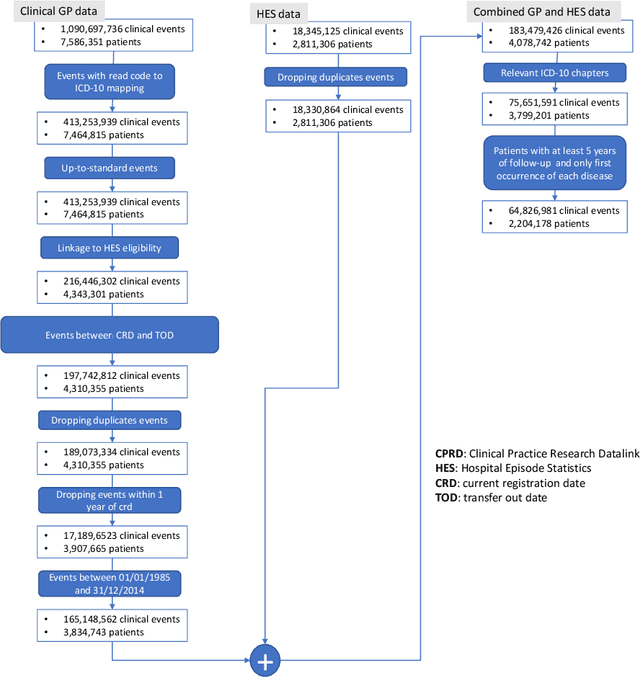

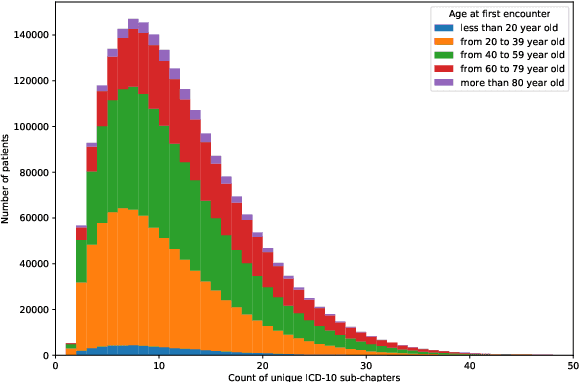
Abstract:Multimorbidity, or the presence of several medical conditions in the same individual, have been increasing in the population both in absolute and relative terms. However, multimorbidity remains poorly understood, and the evidence from existing research to describe its burden, determinants and consequences have been limited. Many of these studies are often cross-sectional and do not explicitly account for multimorbidity patterns' evolution over time. Some studies were based on small datasets, used arbitrary or narrow age range, or lacked appropriate clinical validations. In this study, we applied Non-negative Matrix Factorisation (NMF) in a novel way to one of the largest electronic health records (EHR) databases in the world (with 4 million patients), for simultaneously modelling disease clusters and their role in one's multimorbidity over time. Furthermore, we demonstrated how the temporal characteristics that our model associates with each disease cluster can help mine disease trajectories/networks and generate new hypotheses for the formation of multimorbidity clusters as a function of time/ageing. Our results suggest that our method's ability to learn the underlying dynamics of diseases can provide the field with a novel data-driven / exploratory way of learning the patterns of multimorbidity and their interactions over time.
Performance Measurement for Deep Bayesian Neural Network
Mar 22, 2019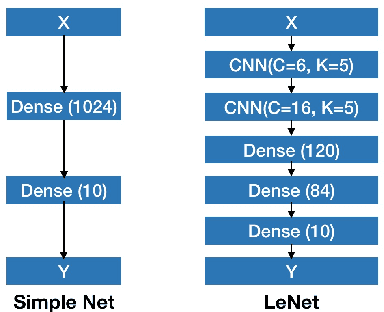
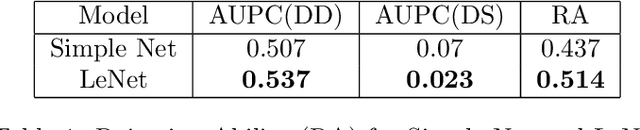
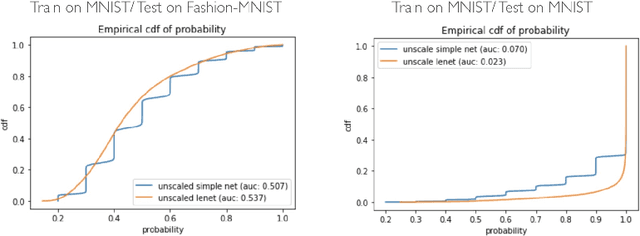
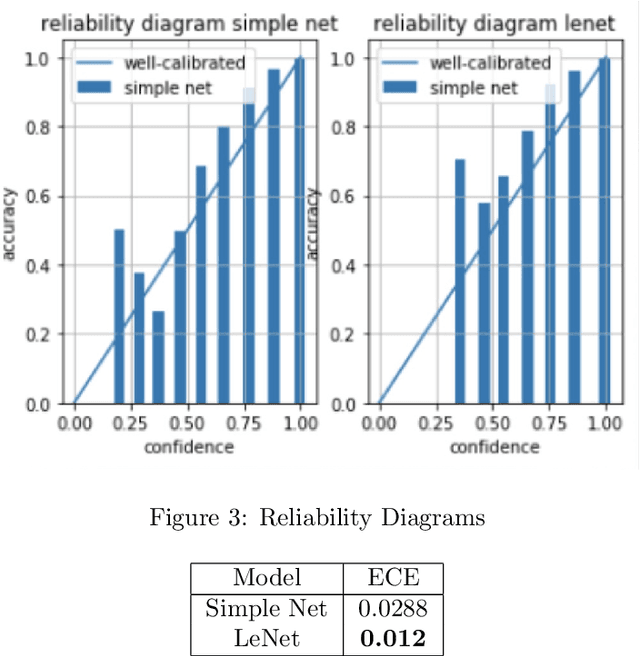
Abstract:Deep Bayesian neural network has aroused a great attention in recent years since it combines the benefits of deep neural network and probability theory. Because of this, the network can make predictions and quantify the uncertainty of the predictions at the same time, which is important in many life-threatening areas. However, most of the recent researches are mainly focusing on making the Bayesian neural network easier to train, and proposing methods to estimate the uncertainty. I notice there are very few works that properly discuss the ways to measure the performance of the Bayesian neural network. Although accuracy and average uncertainty are commonly used for now, they are too general to provide any insight information about the model. In this paper, we would like to introduce more specific criteria and propose several metrics to measure the model performance from different perspectives, which include model calibration measurement, data rejection ability and uncertainty divergence for samples from the same and different distributions.
Early Prediction of Acute Kidney Injury in Critical Care Setting Using Clinical Notes
Nov 09, 2018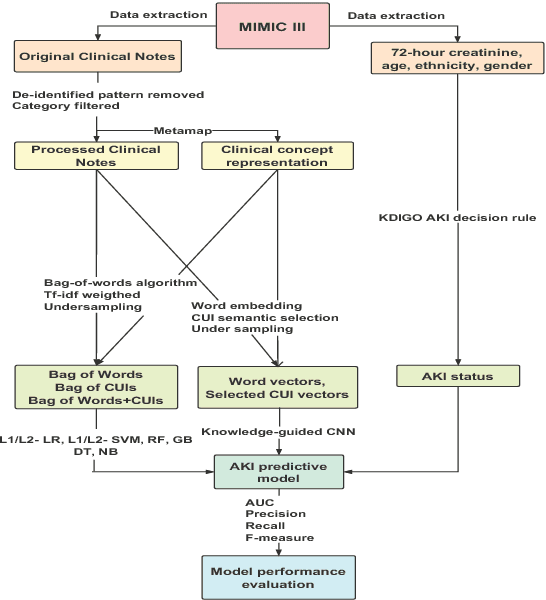
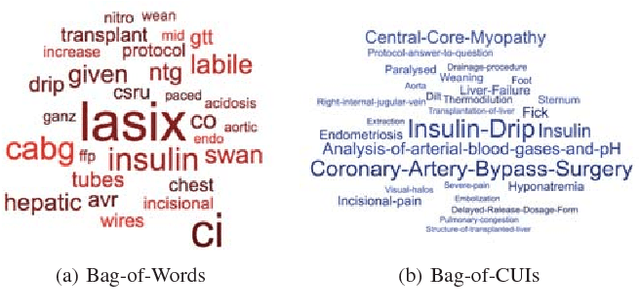

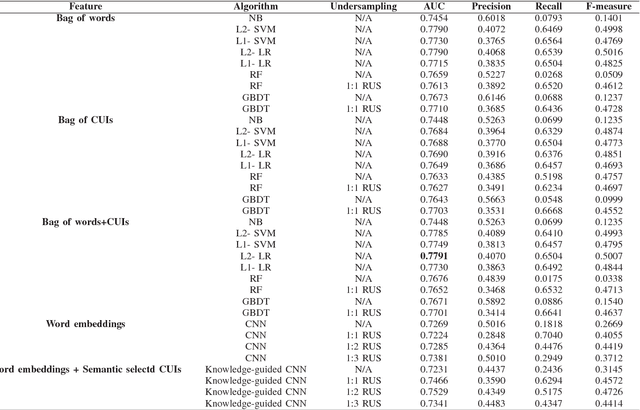
Abstract:Acute kidney injury (AKI) in critically ill patients is associated with significant morbidity and mortality. Development of novel methods to identify patients with AKI earlier will allow for testing of novel strategies to prevent or reduce the complications of AKI. We developed data-driven prediction models to estimate the risk of new AKI onset. We generated models from clinical notes within the first 24 hours following intensive care unit (ICU) admission extracted from Medical Information Mart for Intensive Care III (MIMIC-III). From the clinical notes, we generated clinically meaningful word and concept representations and embeddings, respectively. Five supervised learning classifiers and knowledge-guided deep learning architecture were used to construct prediction models. The best configuration yielded a competitive AUC of 0.779. Our work suggests that natural language processing of clinical notes can be applied to assist clinicians in identifying the risk of incident AKI onset in critically ill patients upon admission to the ICU.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge