Xiaomei Zhao
Automatically Extract the Semi-transparent Motion-blurred Hand from a Single Image
Jun 27, 2019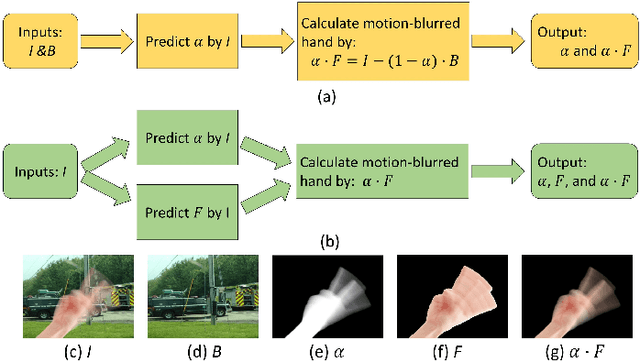
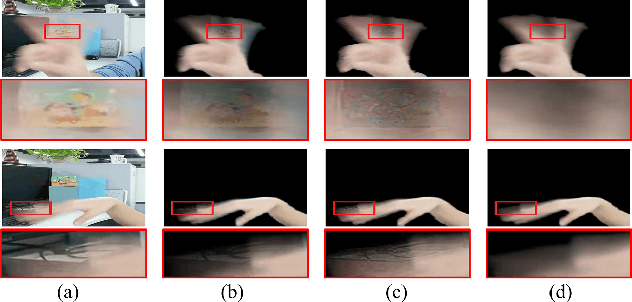
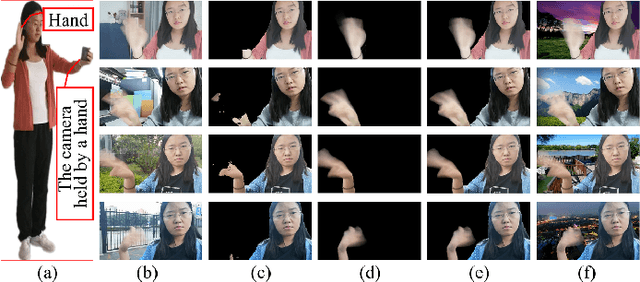
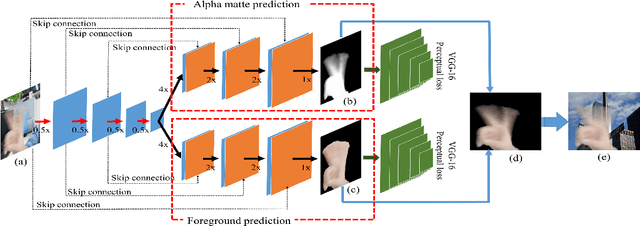


Abstract:When we use video chat, video game, or other video applications, motion-blurred hands often appear. Accurately extracting these hands is very useful for video editing and behavior analysis. However, existing motion-blurred object extraction methods either need user interactions, such as user supplied trimaps and scribbles, or need additional information, such as background images. In this paper, a novel method which can automatically extract the semi-transparent motion-blurred hand just according to the original RGB image is proposed. The proposed method separates the extraction task into two subtasks: alpha matte prediction and foreground prediction. These two subtasks are implemented by Xception based encoder-decoder networks. The extracted motion-blurred hand images can be calculated by multiplying the predicted alpha mattes and foreground images. Experiments on synthetic and real datasets show that the proposed method has promising performance.
A deep learning model integrating FCNNs and CRFs for brain tumor segmentation
Nov 10, 2017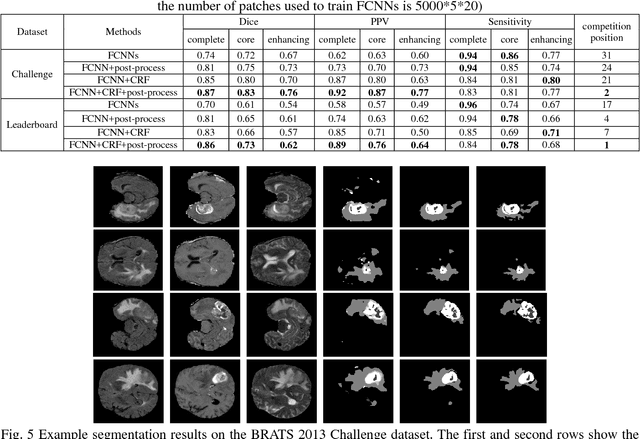
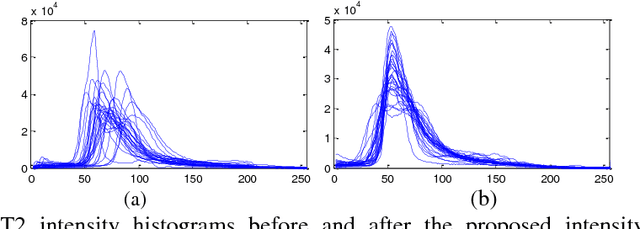
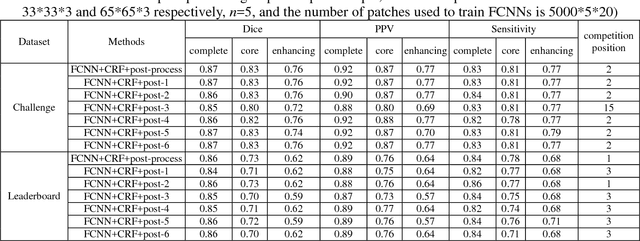
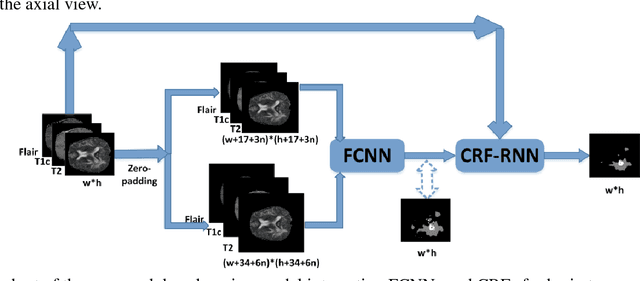
Abstract:Accurate and reliable brain tumor segmentation is a critical component in cancer diagnosis, treatment planning, and treatment outcome evaluation. Build upon successful deep learning techniques, a novel brain tumor segmentation method is developed by integrating fully convolutional neural networks (FCNNs) and Conditional Random Fields (CRFs) in a unified framework to obtain segmentation results with appearance and spatial consistency. We train a deep learning based segmentation model using 2D image patches and image slices in following steps: 1) training FCNNs using image patches; 2) training CRFs as Recurrent Neural Networks (CRF-RNN) using image slices with parameters of FCNNs fixed; and 3) fine-tuning the FCNNs and the CRF-RNN using image slices. Particularly, we train 3 segmentation models using 2D image patches and slices obtained in axial, coronal and sagittal views respectively, and combine them to segment brain tumors using a voting based fusion strategy. Our method could segment brain images slice-by-slice, much faster than those based on image patches. We have evaluated our method based on imaging data provided by the Multimodal Brain Tumor Image Segmentation Challenge (BRATS) 2013, BRATS 2015 and BRATS 2016. The experimental results have demonstrated that our method could build a segmentation model with Flair, T1c, and T2 scans and achieve competitive performance as those built with Flair, T1, T1c, and T2 scans.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge