Vincent Fortuin
A Bayesian Approach to Invariant Deep Neural Networks
Jul 20, 2021
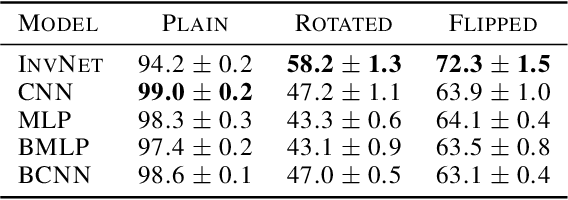
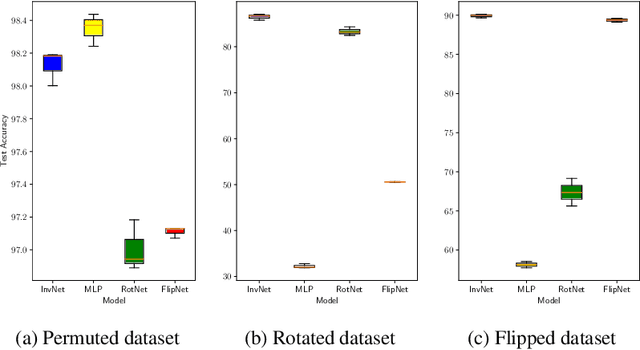
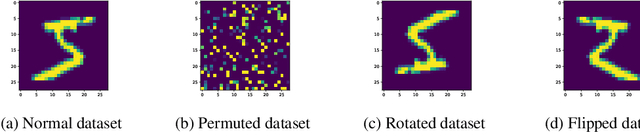
Abstract:We propose a novel Bayesian neural network architecture that can learn invariances from data alone by inferring a posterior distribution over different weight-sharing schemes. We show that our model outperforms other non-invariant architectures, when trained on datasets that contain specific invariances. The same holds true when no data augmentation is performed.
Repulsive Deep Ensembles are Bayesian
Jun 22, 2021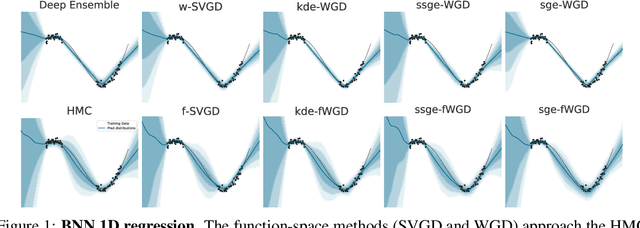
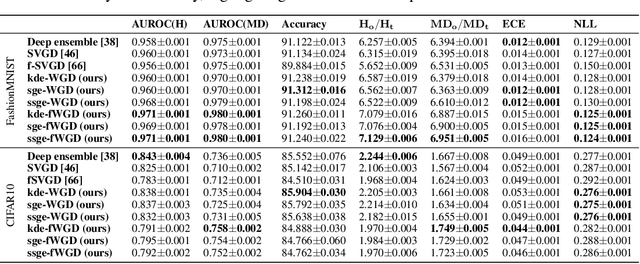
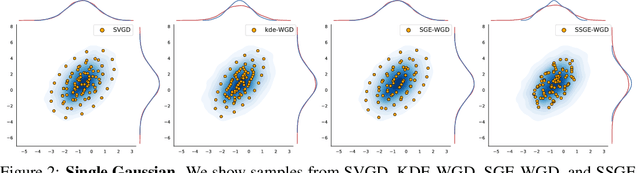
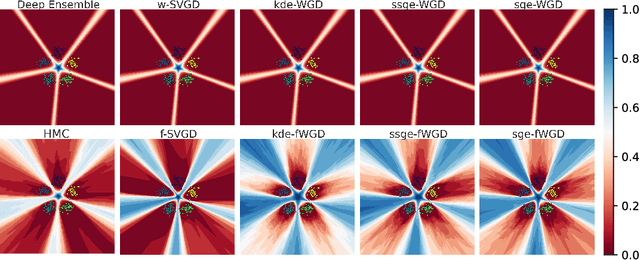
Abstract:Deep ensembles have recently gained popularity in the deep learning community for their conceptual simplicity and efficiency. However, maintaining functional diversity between ensemble members that are independently trained with gradient descent is challenging. This can lead to pathologies when adding more ensemble members, such as a saturation of the ensemble performance, which converges to the performance of a single model. Moreover, this does not only affect the quality of its predictions, but even more so the uncertainty estimates of the ensemble, and thus its performance on out-of-distribution data. We hypothesize that this limitation can be overcome by discouraging different ensemble members from collapsing to the same function. To this end, we introduce a kernelized repulsive term in the update rule of the deep ensembles. We show that this simple modification not only enforces and maintains diversity among the members but, even more importantly, transforms the maximum a posteriori inference into proper Bayesian inference. Namely, we show that the training dynamics of our proposed repulsive ensembles follow a Wasserstein gradient flow of the KL divergence with the true posterior. We study repulsive terms in weight and function space and empirically compare their performance to standard ensembles and Bayesian baselines on synthetic and real-world prediction tasks.
On Stein Variational Neural Network Ensembles
Jun 22, 2021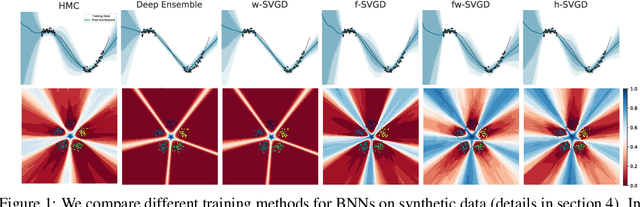
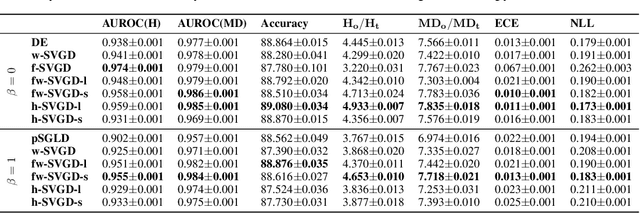
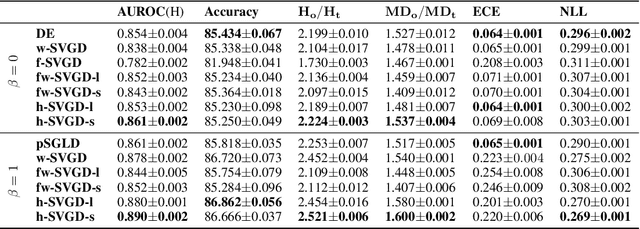
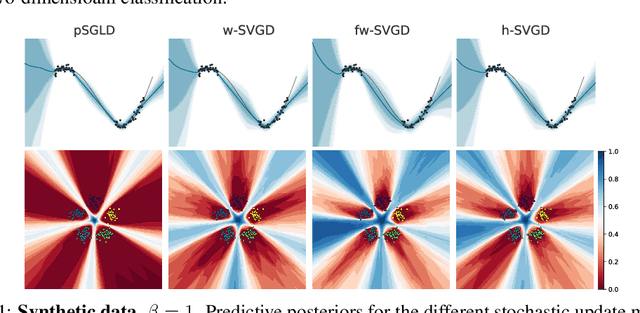
Abstract:Ensembles of deep neural networks have achieved great success recently, but they do not offer a proper Bayesian justification. Moreover, while they allow for averaging of predictions over several hypotheses, they do not provide any guarantees for their diversity, leading to redundant solutions in function space. In contrast, particle-based inference methods, such as Stein variational gradient descent (SVGD), offer a Bayesian framework, but rely on the choice of a kernel to measure the similarity between ensemble members. In this work, we study different SVGD methods operating in the weight space, function space, and in a hybrid setting. We compare the SVGD approaches to other ensembling-based methods in terms of their theoretical properties and assess their empirical performance on synthetic and real-world tasks. We find that SVGD using functional and hybrid kernels can overcome the limitations of deep ensembles. It improves on functional diversity and uncertainty estimation and approaches the true Bayesian posterior more closely. Moreover, we show that using stochastic SVGD updates, as opposed to the standard deterministic ones, can further improve the performance.
Data augmentation in Bayesian neural networks and the cold posterior effect
Jun 10, 2021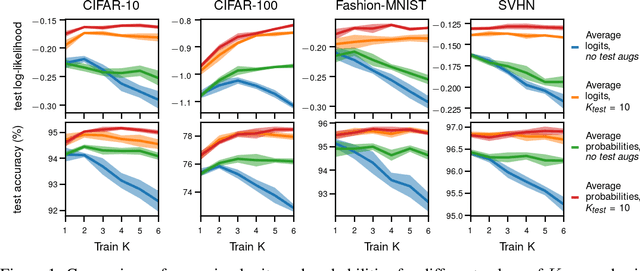
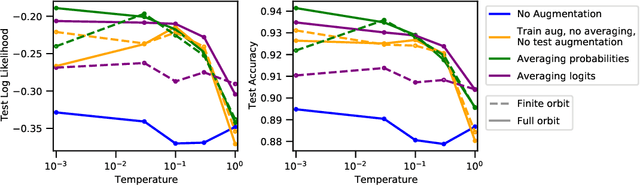
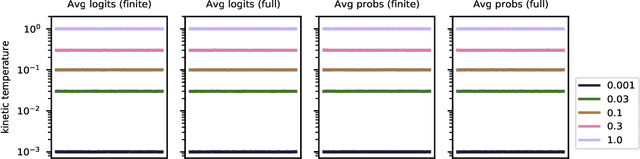
Abstract:Data augmentation is a highly effective approach for improving performance in deep neural networks. The standard view is that it creates an enlarged dataset by adding synthetic data, which raises a problem when combining it with Bayesian inference: how much data are we really conditioning on? This question is particularly relevant to recent observations linking data augmentation to the cold posterior effect. We investigate various principled ways of finding a log-likelihood for augmented datasets. Our approach prescribes augmenting the same underlying image multiple times, both at test and train-time, and averaging either the logits or the predictive probabilities. Empirically, we observe the best performance with averaging probabilities. While there are interactions with the cold posterior effect, neither averaging logits or averaging probabilities eliminates it.
Priors in Bayesian Deep Learning: A Review
May 26, 2021Abstract:While the choice of prior is one of the most critical parts of the Bayesian inference workflow, recent Bayesian deep learning models have often fallen back on vague priors, such as standard Gaussians. In this review, we highlight the importance of prior choices for Bayesian deep learning and present an overview of different priors that have been proposed for (deep) Gaussian processes, variational autoencoders, and Bayesian neural networks. We also outline different methods of learning priors for these models from data. We hope to motivate practitioners in Bayesian deep learning to think more carefully about the prior specification for their models and to provide them with some inspiration in this regard.
BNNpriors: A library for Bayesian neural network inference with different prior distributions
May 14, 2021
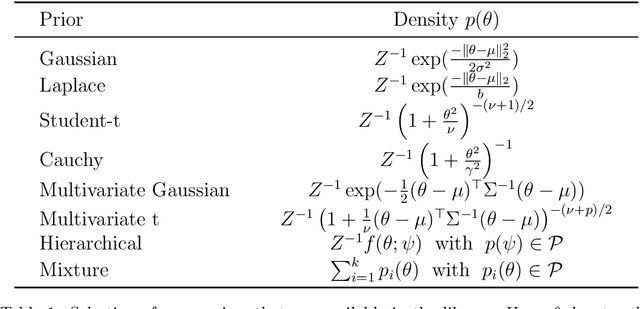

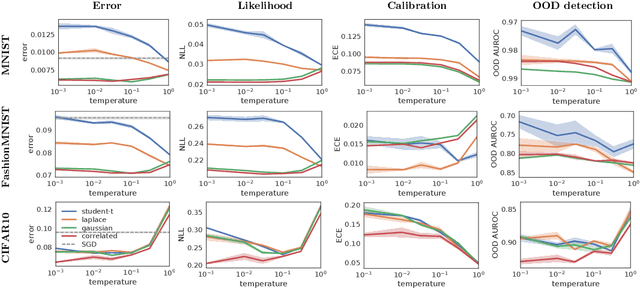
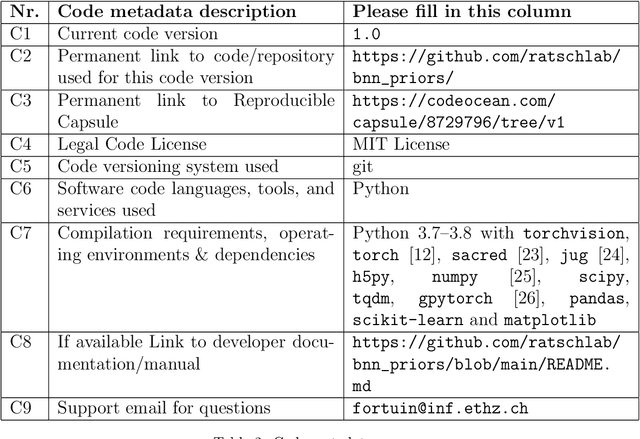
Abstract:Bayesian neural networks have shown great promise in many applications where calibrated uncertainty estimates are crucial and can often also lead to a higher predictive performance. However, it remains challenging to choose a good prior distribution over their weights. While isotropic Gaussian priors are often chosen in practice due to their simplicity, they do not reflect our true prior beliefs well and can lead to suboptimal performance. Our new library, BNNpriors, enables state-of-the-art Markov Chain Monte Carlo inference on Bayesian neural networks with a wide range of predefined priors, including heavy-tailed ones, hierarchical ones, and mixture priors. Moreover, it follows a modular approach that eases the design and implementation of new custom priors. It has facilitated foundational discoveries on the nature of the cold posterior effect in Bayesian neural networks and will hopefully catalyze future research as well as practical applications in this area.
Scalable Marginal Likelihood Estimation for Model Selection in Deep Learning
May 11, 2021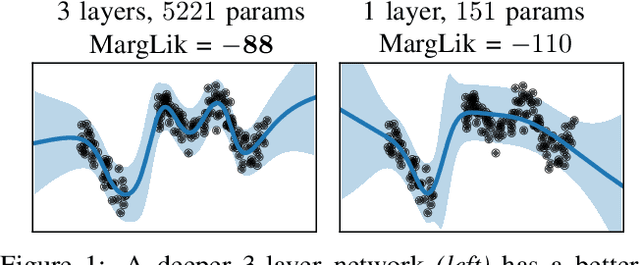
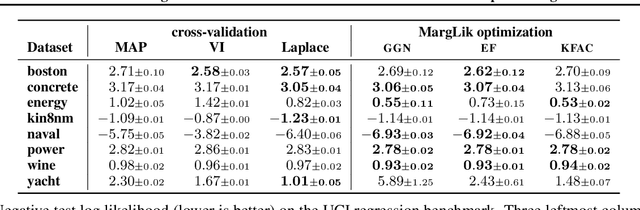
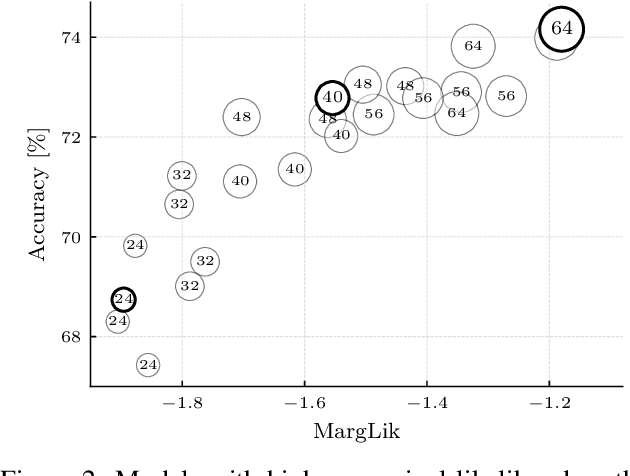
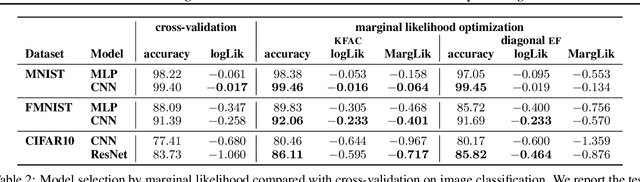
Abstract:Marginal-likelihood based model-selection, even though promising, is rarely used in deep learning due to estimation difficulties. Instead, most approaches rely on validation data, which may not be readily available. In this work, we present a scalable marginal-likelihood estimation method to select both the hyperparameters and network architecture based on the training data alone. Some hyperparameters can be estimated online during training, simplifying the procedure. Our marginal-likelihood estimate is based on Laplace's method and Gauss-Newton approximations to the Hessian, and it outperforms cross-validation and manual-tuning on standard regression and image classification datasets, especially in terms of calibration and out-of-distribution detection. Our work shows that marginal likelihoods can improve generalization and be useful when validation data is unavailable (e.g., in nonstationary settings).
Bayesian Neural Network Priors Revisited
Feb 12, 2021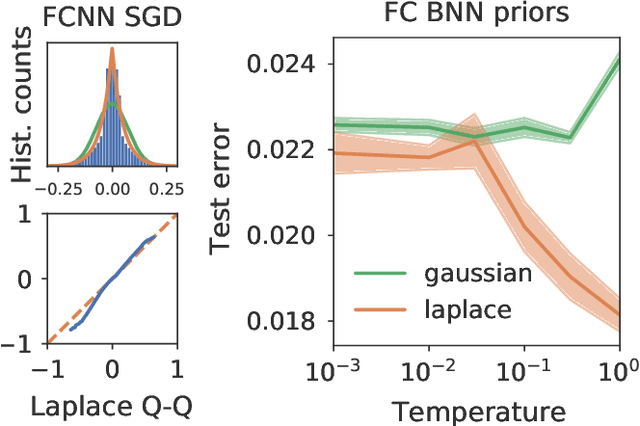
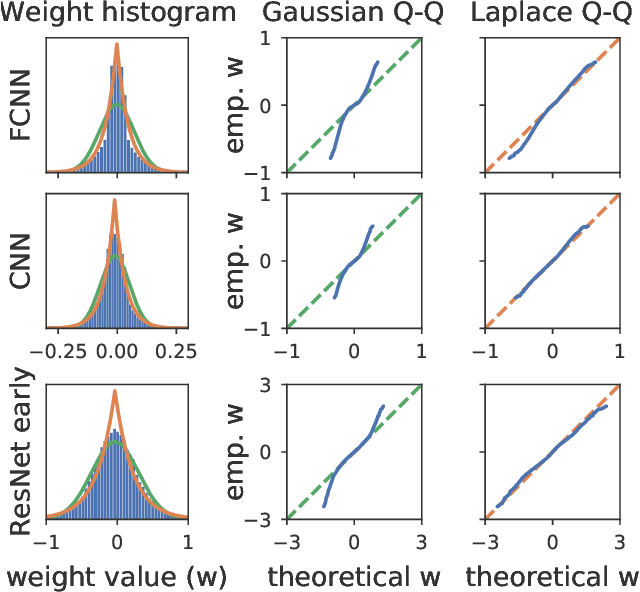
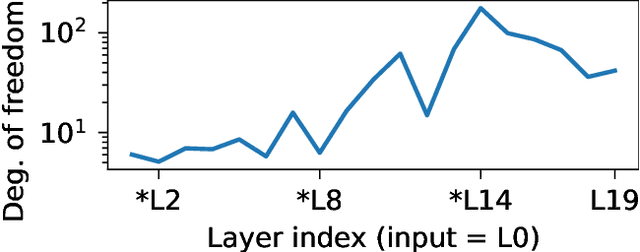
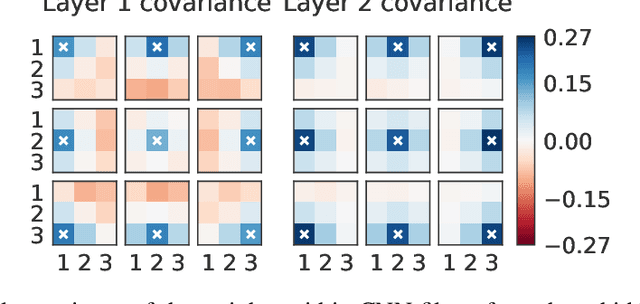
Abstract:Isotropic Gaussian priors are the de facto standard for modern Bayesian neural network inference. However, such simplistic priors are unlikely to either accurately reflect our true beliefs about the weight distributions, or to give optimal performance. We study summary statistics of neural network weights in different networks trained using SGD. We find that fully connected networks (FCNNs) display heavy-tailed weight distributions, while convolutional neural network (CNN) weights display strong spatial correlations. Building these observations into the respective priors leads to improved performance on a variety of image classification datasets. Moreover, we find that these priors also mitigate the cold posterior effect in FCNNs, while in CNNs we see strong improvements at all temperatures, and hence no reduction in the cold posterior effect.
On Disentanglement in Gaussian Process Variational Autoencoders
Feb 10, 2021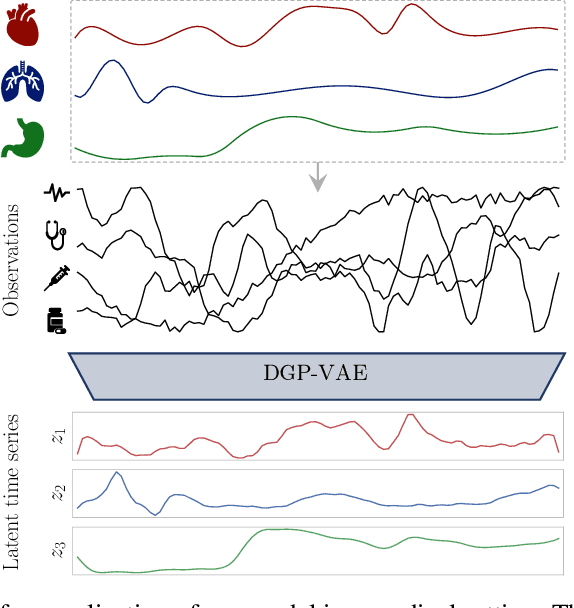


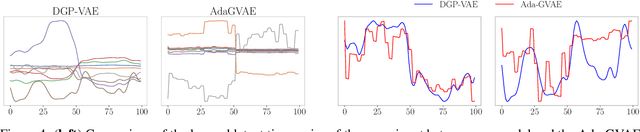
Abstract:Complex multivariate time series arise in many fields, ranging from computer vision to robotics or medicine. Often we are interested in the independent underlying factors that give rise to the high-dimensional data we are observing. While many models have been introduced to learn such disentangled representations, only few attempt to explicitly exploit the structure of sequential data. We investigate the disentanglement properties of Gaussian process variational autoencoders, a class of models recently introduced that have been successful in different tasks on time series data. Our model exploits the temporal structure of the data by modeling each latent channel with a GP prior and employing a structured variational distribution that can capture dependencies in time. We demonstrate the competitiveness of our approach against state-of-the-art unsupervised and weakly-supervised disentanglement methods on a benchmark task. Moreover, we provide evidence that we can learn meaningful disentangled representations on real-world medical time series data.
Annealed Stein Variational Gradient Descent
Feb 08, 2021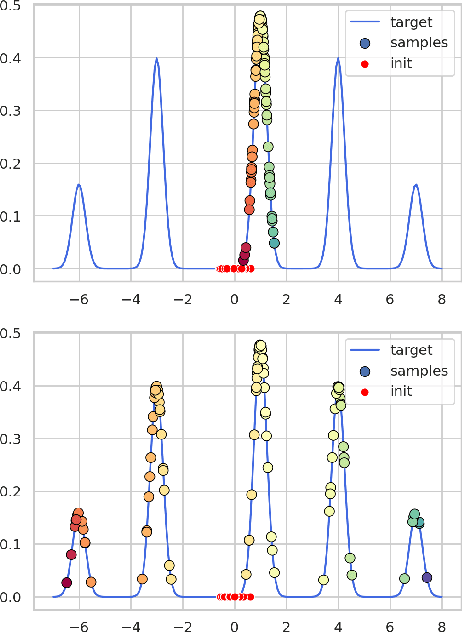
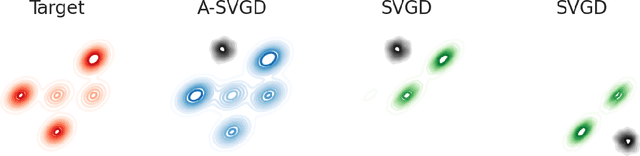
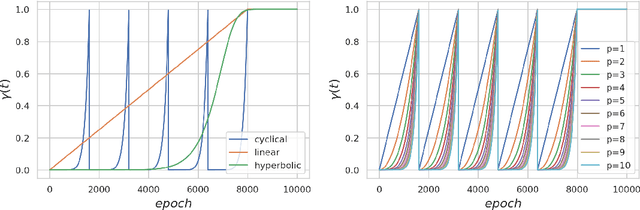
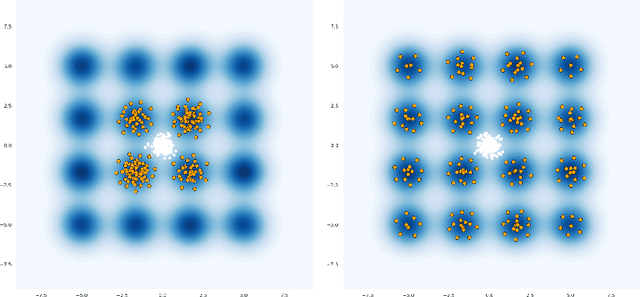
Abstract:Particle based optimization algorithms have recently been developed as sampling methods that iteratively update a set of particles to approximate a target distribution. In particular Stein variational gradient descent has gained attention in the approximate inference literature for its flexibility and accuracy. We empirically explore the ability of this method to sample from multi-modal distributions and focus on two important issues: (i) the inability of the particles to escape from local modes and (ii) the inefficacy in reproducing the density of the different regions. We propose an annealing schedule to solve these issues and show, through various experiments, how this simple solution leads to significant improvements in mode coverage, without invalidating any theoretical properties of the original algorithm.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge