Vikas Garg
The Logical Expressiveness of Topological Neural Networks
Apr 21, 2026Abstract:Graph neural networks (GNNs) are the standard for learning on graphs, yet they have limited expressive power, often expressed in terms of the Weisfeiler-Leman (WL) hierarchy or within the framework of first-order logic. In this context, topological neural networks (TNNs) have recently emerged as a promising alternative for graph representation learning. By incorporating higher-order relational structures into message-passing schemes, TNNs offer higher representational power than traditional GNNs. However, a fundamental question remains open: what is the logical expressiveness of TNNs? Answering this allows us to characterize precisely which binary classifiers TNNs can represent. In this paper, we address this question by analyzing isomorphism tests derived from the underlying mechanisms of general TNNs. We introduce and investigate the power of higher-order variants of WL-based tests for combinatorial complexes, called $k$-CCWL test. In addition, we introduce the topological counting logic (TC$_k$), an extension of standard counting logic featuring a novel pairwise counting quantifier $ \exists^{N}(x_i,x_j)\, \varphi(x_i,x_j), $ which explicitly quantifies pairs $(x_i, x_j)$ satisfying property $\varphi$. We rigorously prove the exact equivalence: $ \text{k-CCWL} \equiv \text{TC}_{k{+}2} \equiv \text{Topological }(k{+}2)\text{-pebble game}.$ These results establish a logical expressiveness theory for TNNs.
* 39 pages, Published at the 14th International Conference on Learning Representations (ICLR 2026)
Contraction and Hourglass Persistence for Learning on Graphs, Simplices, and Cells
Apr 19, 2026Abstract:Persistent homology (PH) encodes global information, such as cycles, and is thus increasingly integrated into graph neural networks (GNNs). PH methods in GNNs typically traverse an increasing sequence of subgraphs. In this work, we first expose limitations of this inclusion procedure. To remedy these shortcomings, we analyze contractions as a principled topological operation, in particular, for graph representation learning. We study the persistence of contraction sequences, which we call Contraction Homology (CH). We establish that forward PH and CH differ in expressivity. We then introduce Hourglass Persistence, a class of topological descriptors that interleave a sequence of inclusions and contractions to boost expressivity, learnability, and stability. We also study related families parametrized by two paradigms. We also discuss how our framework extends to simplicial and cellular networks. We further design efficient algorithms that are pluggable into end-to-end differentiable GNN pipelines, enabling consistent empirical improvements over many PH methods across standard real-world graph datasets. Code is available at \href{https://github.com/Aalto-QuML/Hourglass}{this https URL}.
Canonicalizing Multimodal Contrastive Representation Learning
Feb 19, 2026Abstract:As models and data scale, independently trained networks often induce analogous notions of similarity. But, matching similarities is weaker than establishing an explicit correspondence between the representation spaces, especially for multimodal models, where consistency must hold not only within each modality, but also for the learned image-text coupling. We therefore ask: given two independently trained multimodal contrastive models (with encoders $(f, g)$ and $(\widetilde{f},\widetilde{g})$) -- trained on different distributions and with different architectures -- does a systematic geometric relationship exist between their embedding spaces? If so, what form does it take, and does it hold uniformly across modalities? In this work, we show that across model families such as CLIP, SigLIP, and FLAVA, this geometric relationship is well approximated by an orthogonal map (up to a global mean shift), i.e., there exists an orthogonal map $Q$ where $Q^\top Q = I$ such that $\widetilde{f}(x)\approx Q f(x)$ for paired images $x$. Strikingly, the same $Q$ simultaneously aligns the text encoders i.e., $\widetilde{g}(y)\approx Q g(y)$ for texts $y$. Theoretically, we prove that if the multimodal kernel agrees across models on a small anchor set i.e. $\langle f(x), g(y)\rangle \approx \langle \widetilde{f}(x), \widetilde{g}(y)\rangle$, then the two models must be related by a single orthogonal map $Q$ and the same $Q$ maps images and text across models. More broadly, this finding enables backward-compatible model upgrades, avoiding costly re-embedding, and has implications for the privacy of learned representations. Our project page: https://canonical-multimodal.github.io/
On topological descriptors for graph products
Nov 12, 2025Abstract:Topological descriptors have been increasingly utilized for capturing multiscale structural information in relational data. In this work, we consider various filtrations on the (box) product of graphs and the effect on their outputs on the topological descriptors - the Euler characteristic (EC) and persistent homology (PH). In particular, we establish a complete characterization of the expressive power of EC on general color-based filtrations. We also show that the PH descriptors of (virtual) graph products contain strictly more information than the computation on individual graphs, whereas EC does not. Additionally, we provide algorithms to compute the PH diagrams of the product of vertex- and edge-level filtrations on the graph product. We also substantiate our theoretical analysis with empirical investigations on runtime analysis, expressivity, and graph classification performance. Overall, this work paves way for powerful graph persistent descriptors via product filtrations. Code is available at https://github.com/Aalto-QuML/tda_graph_product.
Heat Kernel Goes Topological
Jul 16, 2025Abstract:Topological neural networks have emerged as powerful successors of graph neural networks. However, they typically involve higher-order message passing, which incurs significant computational expense. We circumvent this issue with a novel topological framework that introduces a Laplacian operator on combinatorial complexes (CCs), enabling efficient computation of heat kernels that serve as node descriptors. Our approach captures multiscale information and enables permutation-equivariant representations, allowing easy integration into modern transformer-based architectures. Theoretically, the proposed method is maximally expressive because it can distinguish arbitrary non-isomorphic CCs. Empirically, it significantly outperforms existing topological methods in terms of computational efficiency. Besides demonstrating competitive performance with the state-of-the-art descriptors on standard molecular datasets, it exhibits superior capability in distinguishing complex topological structures and avoiding blind spots on topological benchmarks. Overall, this work advances topological deep learning by providing expressive yet scalable representations, thereby opening up exciting avenues for molecular classification and property prediction tasks.
Positional Encoding meets Persistent Homology on Graphs
Jun 06, 2025Abstract:The local inductive bias of message-passing graph neural networks (GNNs) hampers their ability to exploit key structural information (e.g., connectivity and cycles). Positional encoding (PE) and Persistent Homology (PH) have emerged as two promising approaches to mitigate this issue. PE schemes endow GNNs with location-aware features, while PH methods enhance GNNs with multiresolution topological features. However, a rigorous theoretical characterization of the relative merits and shortcomings of PE and PH has remained elusive. We bridge this gap by establishing that neither paradigm is more expressive than the other, providing novel constructions where one approach fails but the other succeeds. Our insights inform the design of a novel learnable method, PiPE (Persistence-informed Positional Encoding), which is provably more expressive than both PH and PE. PiPE demonstrates strong performance across a variety of tasks (e.g., molecule property prediction, graph classification, and out-of-distribution generalization), thereby advancing the frontiers of graph representation learning. Code is available at https://github.com/Aalto-QuML/PIPE.
Graph Persistence goes Spectral
Jun 06, 2025Abstract:Including intricate topological information (e.g., cycles) provably enhances the expressivity of message-passing graph neural networks (GNNs) beyond the Weisfeiler-Leman (WL) hierarchy. Consequently, Persistent Homology (PH) methods are increasingly employed for graph representation learning. In this context, recent works have proposed decorating classical PH diagrams with vertex and edge features for improved expressivity. However, due to their dependence on features, these methods still fail to capture basic graph structural information. In this paper, we propose SpectRe -- a new topological descriptor for graphs that integrates spectral information into PH diagrams. Notably, SpectRe is strictly more expressive than existing descriptors on graphs. We also introduce notions of global and local stability to analyze existing descriptors and establish that SpectRe is locally stable. Finally, experiments on synthetic and real-world datasets demonstrate the effectiveness of SpectRe and its potential to enhance the capabilities of graph models in relevant learning tasks.
Spacetime Geometry of Denoising in Diffusion Models
May 23, 2025Abstract:We present a novel perspective on diffusion models using the framework of information geometry. We show that the set of noisy samples, taken across all noise levels simultaneously, forms a statistical manifold -- a family of denoising probability distributions. Interpreting the noise level as a temporal parameter, we refer to this manifold as spacetime. This manifold naturally carries a Fisher-Rao metric, which defines geodesics -- shortest paths between noisy points. Notably, this family of distributions is exponential, enabling efficient geodesic computation even in high-dimensional settings without retraining or fine-tuning. We demonstrate the practical value of this geometric viewpoint in transition path sampling, where spacetime geodesics define smooth sequences of Boltzmann distributions, enabling the generation of continuous trajectories between low-energy metastable states. Code is available at: https://github.com/Aalto-QuML/diffusion-spacetime-geometry.
Heterophily-informed Message Passing
Apr 28, 2025



Abstract:Graph neural networks (GNNs) are known to be vulnerable to oversmoothing due to their implicit homophily assumption. We mitigate this problem with a novel scheme that regulates the aggregation of messages, modulating the type and extent of message passing locally thereby preserving both the low and high-frequency components of information. Our approach relies solely on learnt embeddings, obviating the need for auxiliary labels, thus extending the benefits of heterophily-aware embeddings to broader applications, e.g., generative modelling. Our experiments, conducted across various data sets and GNN architectures, demonstrate performance enhancements and reveal heterophily patterns across standard classification benchmarks. Furthermore, application to molecular generation showcases notable performance improvements on chemoinformatics benchmarks.
Robust Simulation-Based Inference under Missing Data via Neural Processes
Mar 03, 2025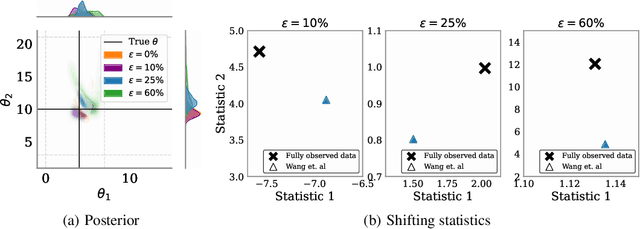

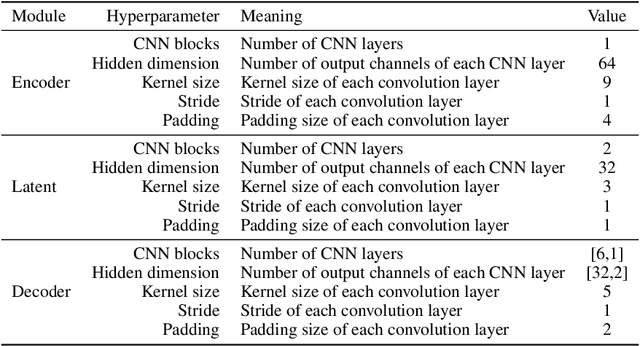
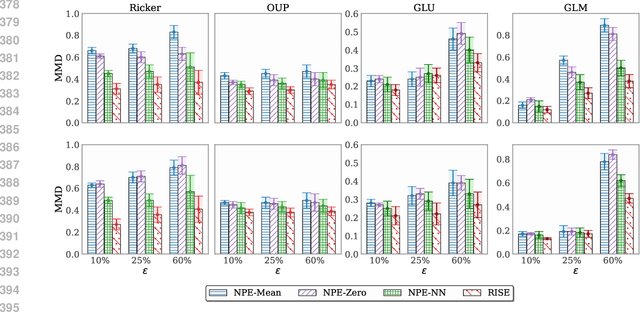
Abstract:Simulation-based inference (SBI) methods typically require fully observed data to infer parameters of models with intractable likelihood functions. However, datasets often contain missing values due to incomplete observations, data corruptions (common in astrophysics), or instrument limitations (e.g., in high-energy physics applications). In such scenarios, missing data must be imputed before applying any SBI method. We formalize the problem of missing data in SBI and demonstrate that naive imputation methods can introduce bias in the estimation of SBI posterior. We also introduce a novel amortized method that addresses this issue by jointly learning the imputation model and the inference network within a neural posterior estimation (NPE) framework. Extensive empirical results on SBI benchmarks show that our approach provides robust inference outcomes compared to standard baselines for varying levels of missing data. Moreover, we demonstrate the merits of our imputation model on two real-world bioactivity datasets (Adrenergic and Kinase assays). Code is available at https://github.com/Aalto-QuML/RISE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge