Veronika Cheplygina
Crowd disagreement about medical images is informative
Aug 17, 2018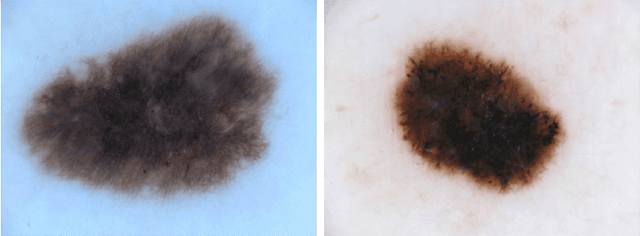
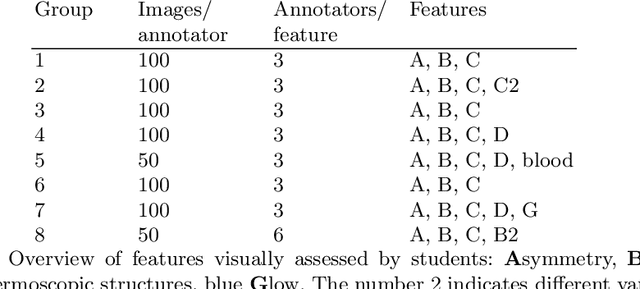
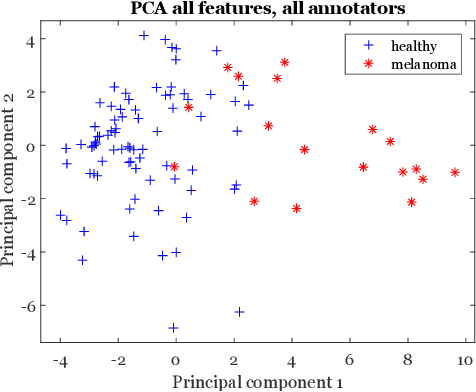
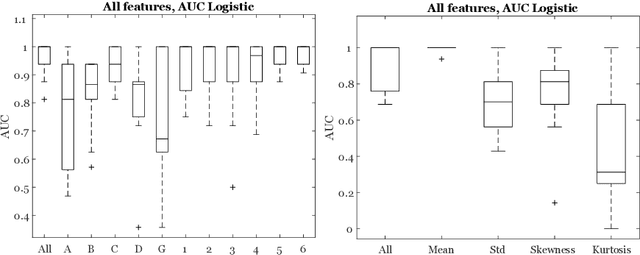
Abstract:Classifiers for medical image analysis are often trained with a single consensus label, based on combining labels given by experts or crowds. However, disagreement between annotators may be informative, and thus removing it may not be the best strategy. As a proof of concept, we predict whether a skin lesion from the ISIC 2017 dataset is a melanoma or not, based on crowd annotations of visual characteristics of that lesion. We compare using the mean annotations, illustrating consensus, to standard deviations and other distribution moments, illustrating disagreement. We show that the mean annotations perform best, but that the disagreement measures are still informative. We also make the crowd annotations used in this paper available at \url{https://figshare.com/s/5cbbce14647b66286544}.
Characterizing multiple instance datasets
Jun 21, 2018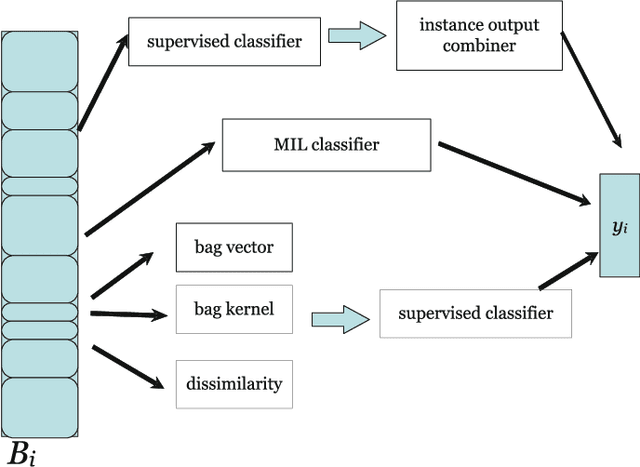
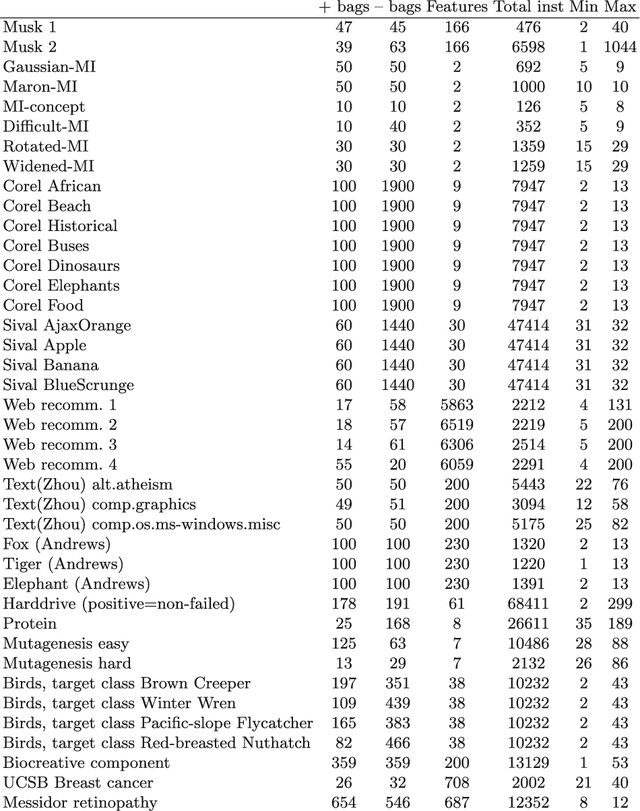
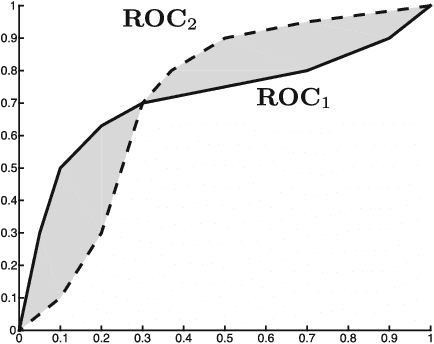
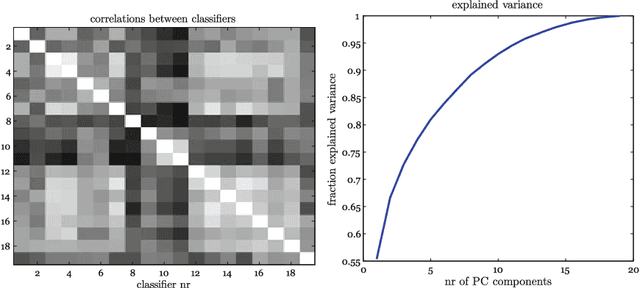
Abstract:In many pattern recognition problems, a single feature vector is not sufficient to describe an object. In multiple instance learning (MIL), objects are represented by sets (\emph{bags}) of feature vectors (\emph{instances}). This requires an adaptation of standard supervised classifiers in order to train and evaluate on these bags of instances. Like for supervised classification, several benchmark datasets and numerous classifiers are available for MIL. When performing a comparison of different MIL classifiers, it is important to understand the differences of the datasets, used in the comparison. Seemingly different (based on factors such as dimensionality) datasets may elicit very similar behaviour in classifiers, and vice versa. This has implications for what kind of conclusions may be drawn from the comparison results. We aim to give an overview of the variability of available benchmark datasets and some popular MIL classifiers. We use a dataset dissimilarity measure, based on the differences between the ROC-curves obtained by different classifiers, and embed this dataset dissimilarity matrix into a low-dimensional space. Our results show that conceptually similar datasets can behave very differently. We therefore recommend examining such dataset characteristics when making comparisons between existing and new MIL classifiers. The datasets are available via Figshare at \url{https://bit.ly/2K9iTja}.
Feature learning based on visual similarity triplets in medical image analysis: A case study of emphysema in chest CT scans
Jun 19, 2018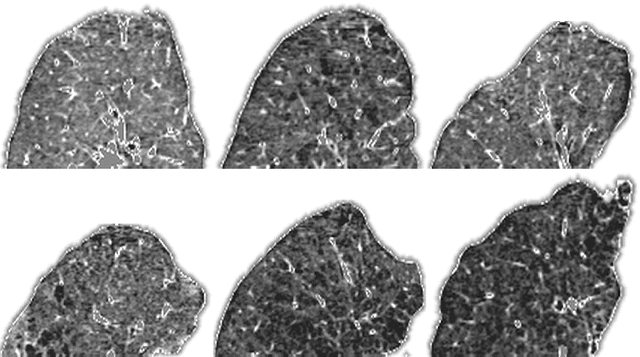
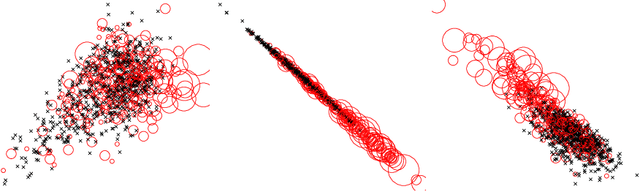



Abstract:Supervised feature learning using convolutional neural networks (CNNs) can provide concise and disease relevant representations of medical images. However, training CNNs requires annotated image data. Annotating medical images can be a time-consuming task and even expert annotations are subject to substantial inter- and intra-rater variability. Assessing visual similarity of images instead of indicating specific pathologies or estimating disease severity could allow non-experts to participate, help uncover new patterns, and possibly reduce rater variability. We consider the task of assessing emphysema extent in chest CT scans. We derive visual similarity triplets from visually assessed emphysema extent and learn a low dimensional embedding using CNNs. We evaluate the networks on 973 images, and show that the CNNs can learn disease relevant feature representations from derived similarity triplets. To our knowledge this is the first medical image application where similarity triplets has been used to learn a feature representation that can be used for embedding unseen test images
Transfer learning for multi-center classification of chronic obstructive pulmonary disease
Nov 23, 2017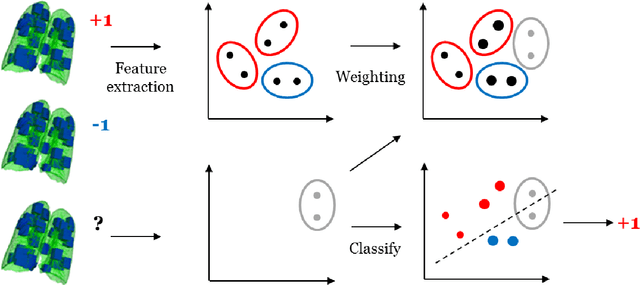
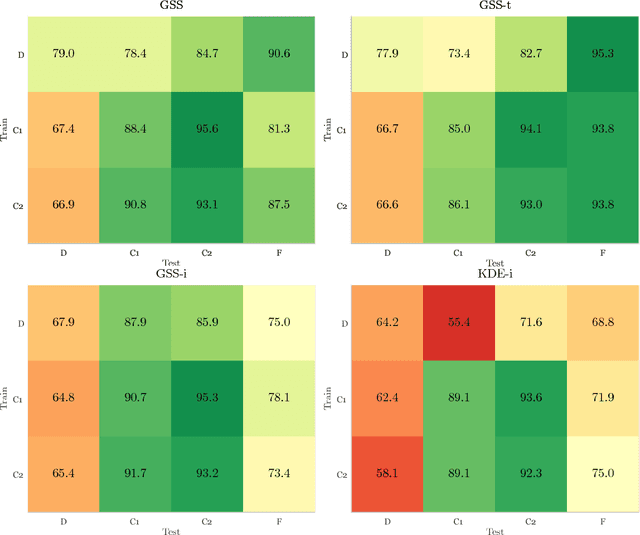
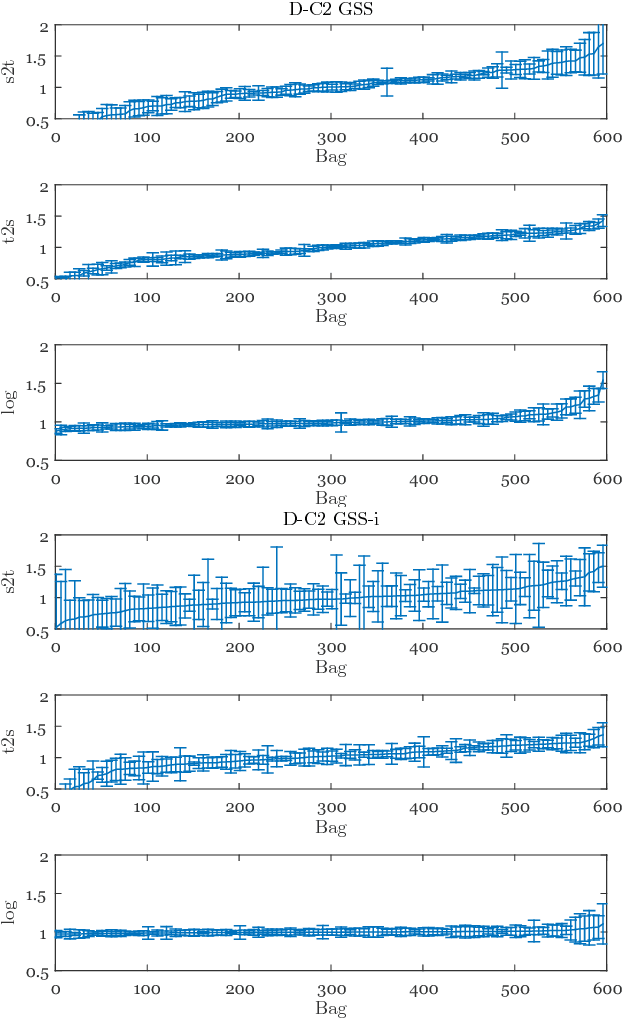
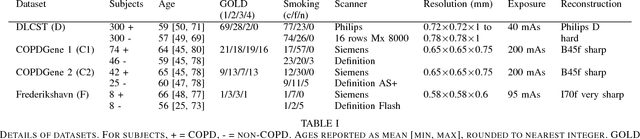
Abstract:Chronic obstructive pulmonary disease (COPD) is a lung disease which can be quantified using chest computed tomography (CT) scans. Recent studies have shown that COPD can be automatically diagnosed using weakly supervised learning of intensity and texture distributions. However, up till now such classifiers have only been evaluated on scans from a single domain, and it is unclear whether they would generalize across domains, such as different scanners or scanning protocols. To address this problem, we investigate classification of COPD in a multi-center dataset with a total of 803 scans from three different centers, four different scanners, with heterogenous subject distributions. Our method is based on Gaussian texture features, and a weighted logistic classifier, which increases the weights of samples similar to the test data. We show that Gaussian texture features outperform intensity features previously used in multi-center classification tasks. We also show that a weighting strategy based on a classifier that is trained to discriminate between scans from different domains, can further improve the results. To encourage further research into transfer learning methods for classification of COPD, upon acceptance of the paper we will release two feature datasets used in this study on http://bigr.nl/research/projects/copd
Exploring the similarity of medical imaging classification problems
Jun 12, 2017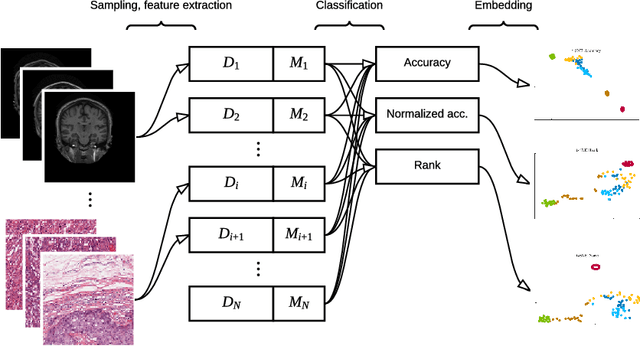
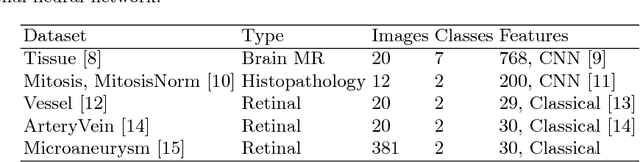
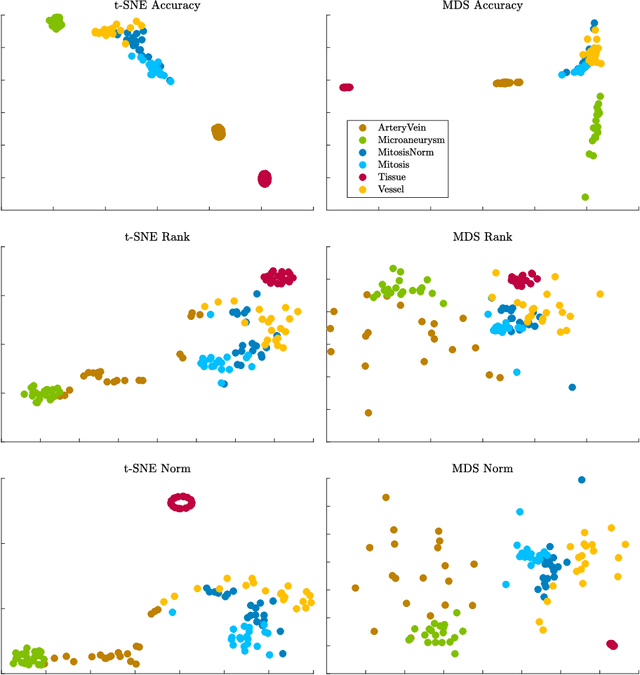
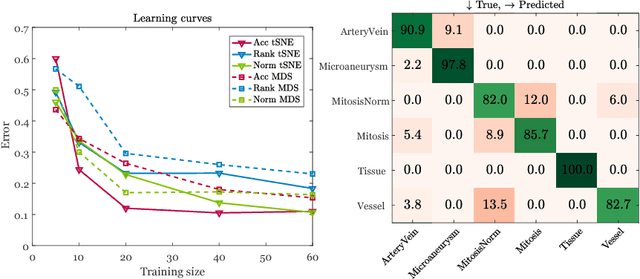
Abstract:Supervised learning is ubiquitous in medical image analysis. In this paper we consider the problem of meta-learning -- predicting which methods will perform well in an unseen classification problem, given previous experience with other classification problems. We investigate the first step of such an approach: how to quantify the similarity of different classification problems. We characterize datasets sampled from six classification problems by performance ranks of simple classifiers, and define the similarity by the inverse of Euclidean distance in this meta-feature space. We visualize the similarities in a 2D space, where meaningful clusters start to emerge, and show that the proposed representation can be used to classify datasets according to their origin with 89.3\% accuracy. These findings, together with the observations of recent trends in machine learning, suggest that meta-learning could be a valuable tool for the medical imaging community.
Early Experiences with Crowdsourcing Airway Annotations in Chest CT
Jun 07, 2017
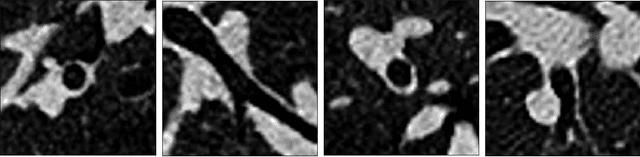
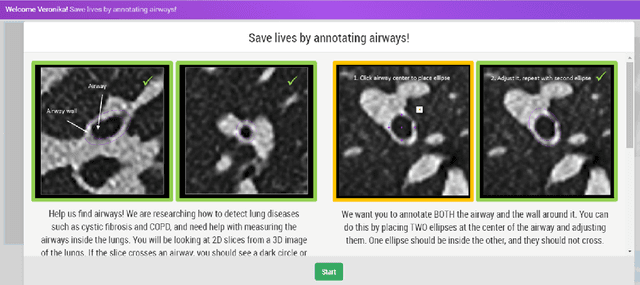
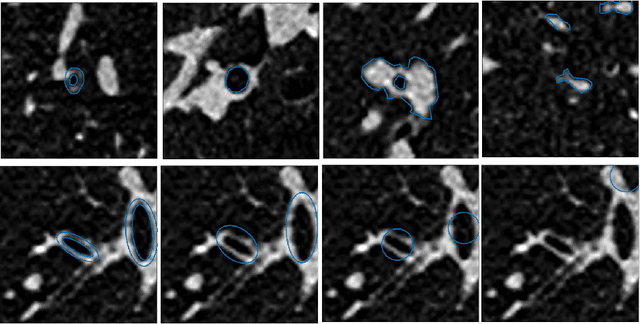
Abstract:Measuring airways in chest computed tomography (CT) images is important for characterizing diseases such as cystic fibrosis, yet very time-consuming to perform manually. Machine learning algorithms offer an alternative, but need large sets of annotated data to perform well. We investigate whether crowdsourcing can be used to gather airway annotations which can serve directly for measuring the airways, or as training data for the algorithms. We generate image slices at known locations of airways and request untrained crowd workers to outline the airway lumen and airway wall. Our results show that the workers are able to interpret the images, but that the instructions are too complex, leading to many unusable annotations. After excluding unusable annotations, quantitative results show medium to high correlations with expert measurements of the airways. Based on this positive experience, we describe a number of further research directions and provide insight into the challenges of crowdsourcing in medical images from the perspective of first-time users.
Label Stability in Multiple Instance Learning
Mar 15, 2017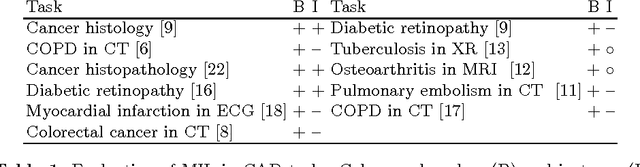

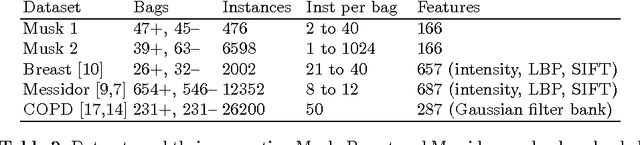
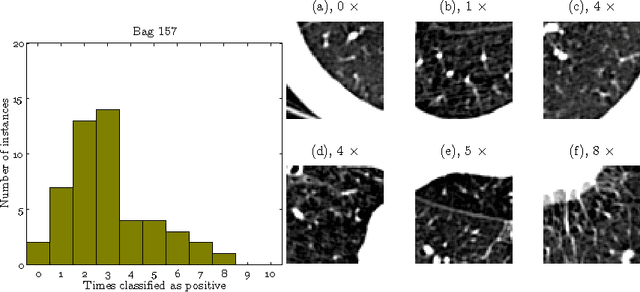
Abstract:We address the problem of \emph{instance label stability} in multiple instance learning (MIL) classifiers. These classifiers are trained only on globally annotated images (bags), but often can provide fine-grained annotations for image pixels or patches (instances). This is interesting for computer aided diagnosis (CAD) and other medical image analysis tasks for which only a coarse labeling is provided. Unfortunately, the instance labels may be unstable. This means that a slight change in training data could potentially lead to abnormalities being detected in different parts of the image, which is undesirable from a CAD point of view. Despite MIL gaining popularity in the CAD literature, this issue has not yet been addressed. We investigate the stability of instance labels provided by several MIL classifiers on 5 different datasets, of which 3 are medical image datasets (breast histopathology, diabetic retinopathy and computed tomography lung images). We propose an unsupervised measure to evaluate instance stability, and demonstrate that a performance-stability trade-off can be made when comparing MIL classifiers.
Transfer Learning by Asymmetric Image Weighting for Segmentation across Scanners
Mar 15, 2017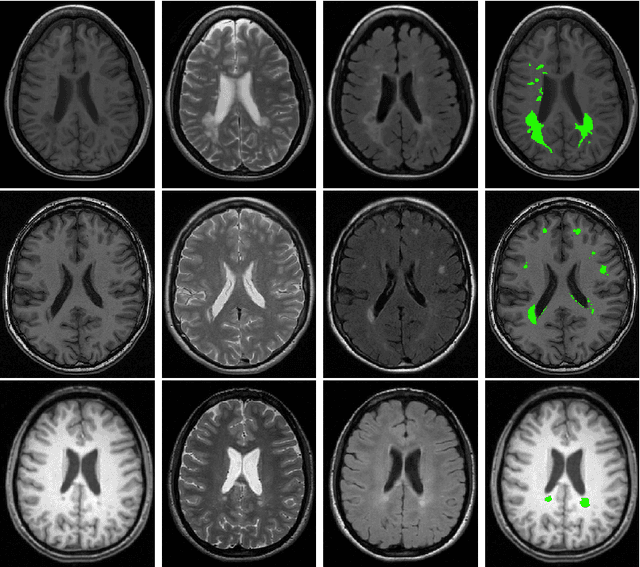
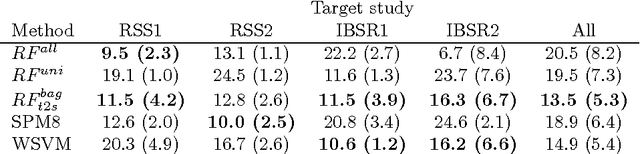
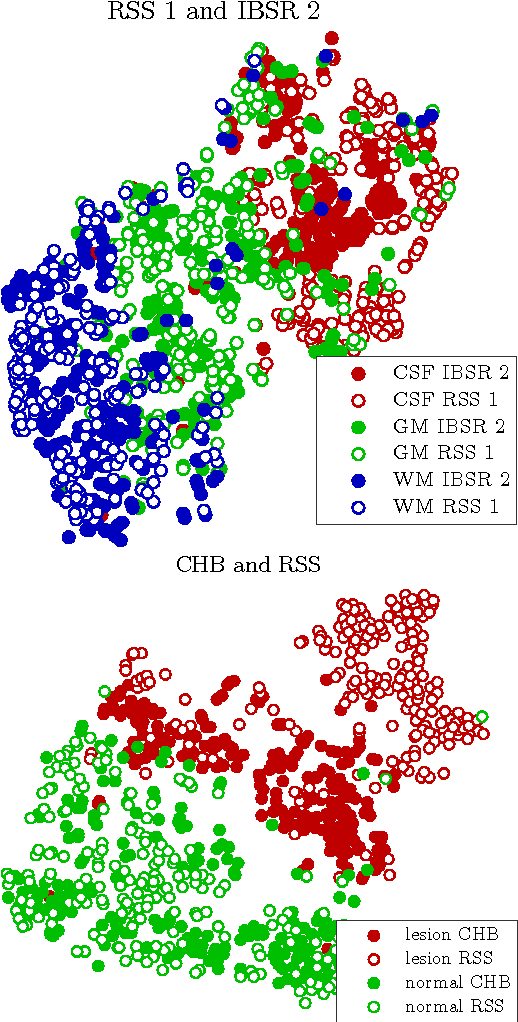
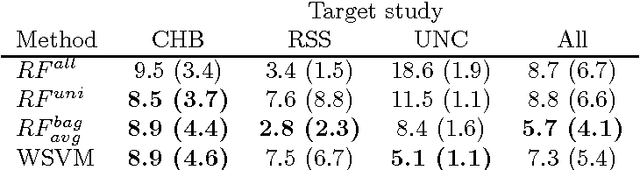
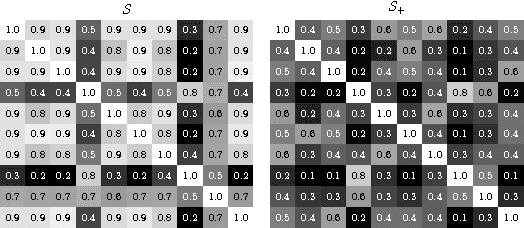
Abstract:Supervised learning has been very successful for automatic segmentation of images from a single scanner. However, several papers report deteriorated performances when using classifiers trained on images from one scanner to segment images from other scanners. We propose a transfer learning classifier that adapts to differences between training and test images. This method uses a weighted ensemble of classifiers trained on individual images. The weight of each classifier is determined by the similarity between its training image and the test image. We examine three unsupervised similarity measures, which can be used in scenarios where no labeled data from a newly introduced scanner or scanning protocol is available. The measures are based on a divergence, a bag distance, and on estimating the labels with a clustering procedure. These measures are asymmetric. We study whether the asymmetry can improve classification. Out of the three similarity measures, the bag similarity measure is the most robust across different studies and achieves excellent results on four brain tissue segmentation datasets and three white matter lesion segmentation datasets, acquired at different centers and with different scanners and scanning protocols. We show that the asymmetry can indeed be informative, and that computing the similarity from the test image to the training images is more appropriate than the opposite direction.
Classification of COPD with Multiple Instance Learning
Mar 15, 2017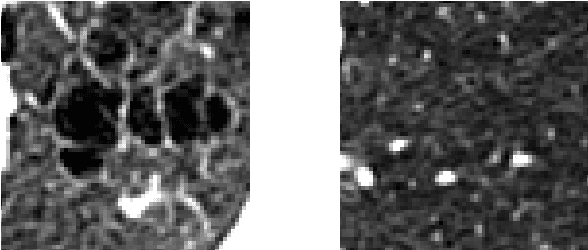
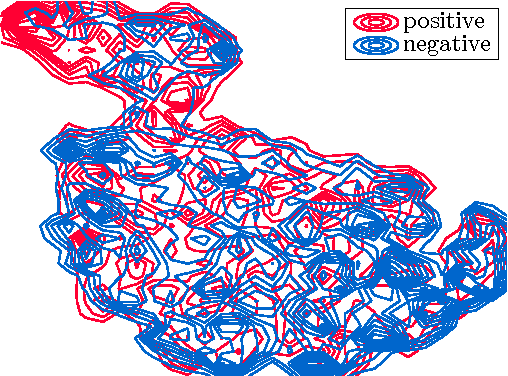

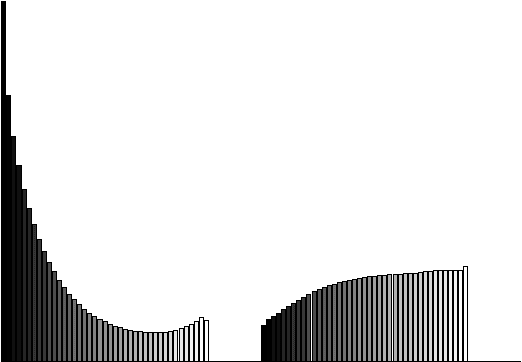
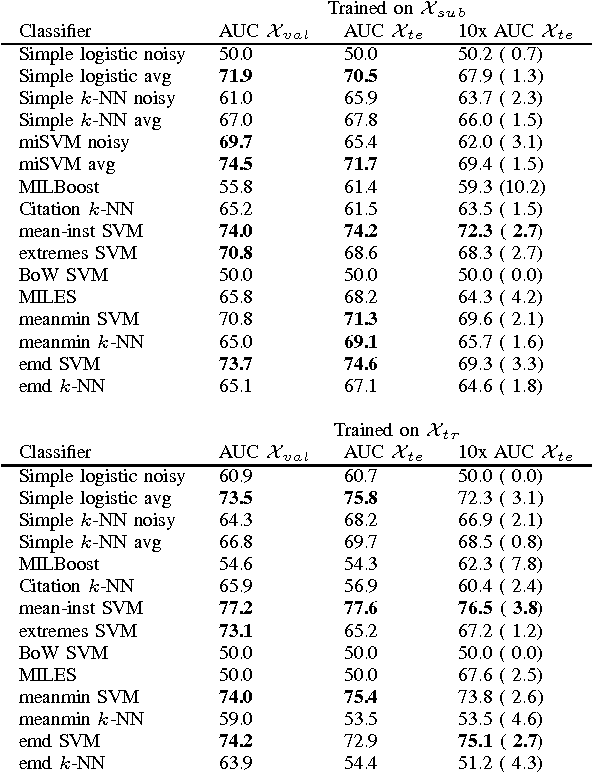
Abstract:Chronic obstructive pulmonary disease (COPD) is a lung disease where early detection benefits the survival rate. COPD can be quantified by classifying patches of computed tomography images, and combining patch labels into an overall diagnosis for the image. As labeled patches are often not available, image labels are propagated to the patches, incorrectly labeling healthy patches in COPD patients as being affected by the disease. We approach quantification of COPD from lung images as a multiple instance learning (MIL) problem, which is more suitable for such weakly labeled data. We investigate various MIL assumptions in the context of COPD and show that although a concept region with COPD-related disease patterns is present, considering the whole distribution of lung tissue patches improves the performance. The best method is based on averaging instances and obtains an AUC of 0.742, which is higher than the previously reported best of 0.713 on the same dataset. Using the full training set further increases performance to 0.776, which is significantly higher (DeLong test) than previous results.
Multiple Instance Learning: A Survey of Problem Characteristics and Applications
Dec 11, 2016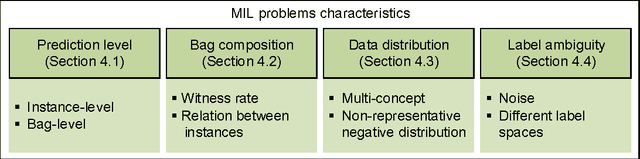
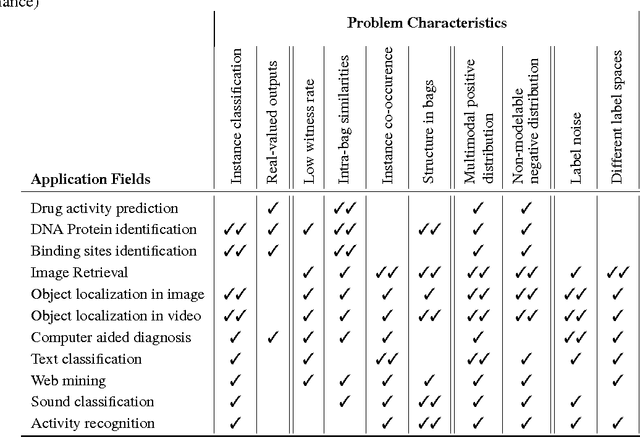
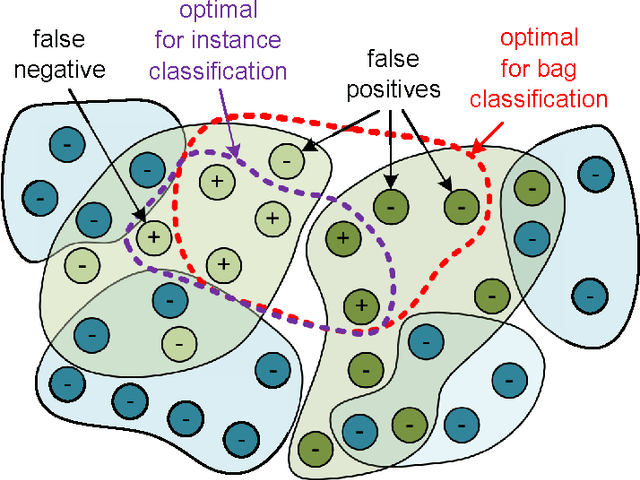
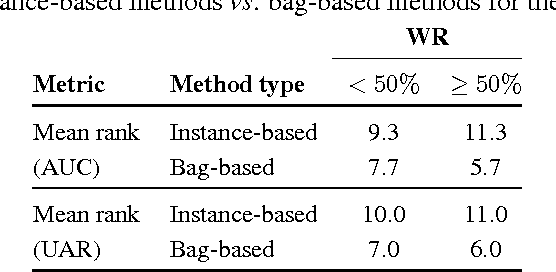
Abstract:Multiple instance learning (MIL) is a form of weakly supervised learning where training instances are arranged in sets, called bags, and a label is provided for the entire bag. This formulation is gaining interest because it naturally fits various problems and allows to leverage weakly labeled data. Consequently, it has been used in diverse application fields such as computer vision and document classification. However, learning from bags raises important challenges that are unique to MIL. This paper provides a comprehensive survey of the characteristics which define and differentiate the types of MIL problems. Until now, these problem characteristics have not been formally identified and described. As a result, the variations in performance of MIL algorithms from one data set to another are difficult to explain. In this paper, MIL problem characteristics are grouped into four broad categories: the composition of the bags, the types of data distribution, the ambiguity of instance labels, and the task to be performed. Methods specialized to address each category are reviewed. Then, the extent to which these characteristics manifest themselves in key MIL application areas are described. Finally, experiments are conducted to compare the performance of 16 state-of-the-art MIL methods on selected problem characteristics. This paper provides insight on how the problem characteristics affect MIL algorithms, recommendations for future benchmarking and promising avenues for research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge